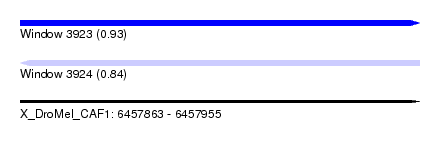

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 6,457,863 – 6,457,955 |
| Length | 92 |
| Max. P | 0.927223 |

| Location | 6,457,863 – 6,457,955 |
|---|---|
| Length | 92 |
| Sequences | 4 |
| Columns | 96 |
| Reading direction | forward |
| Mean pairwise identity | 80.78 |
| Mean single sequence MFE | -23.05 |
| Consensus MFE | -15.18 |
| Energy contribution | -16.43 |
| Covariance contribution | 1.25 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.21 |
| Structure conservation index | 0.66 |
| SVM decision value | 1.18 |
| SVM RNA-class probability | 0.927223 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
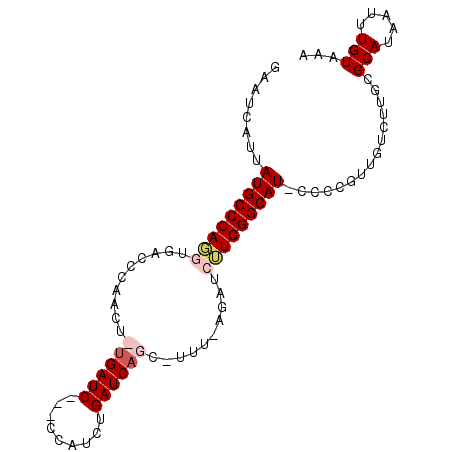
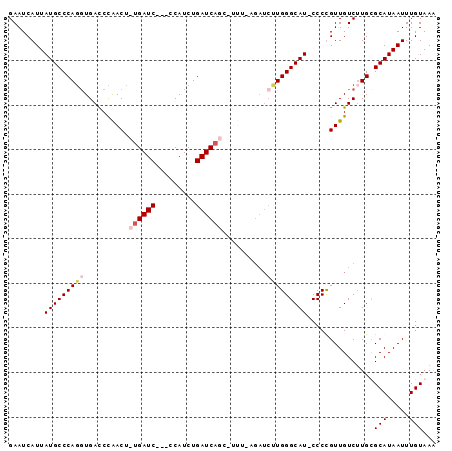
>X_DroMel_CAF1 6457863 92 + 22224390 GAAUCAUUAUGCCCAGGUGACCCGACUCUGAUC---CCAUCUGAUCAGU-UUUGAGAUCUUGGGCAUUCCCUGUUGUCUUGCGCAUAAUUUGUAAA ((..((..(((((((((...(.(((..((((((---......)))))).-.))).)..)))))))))....))...))....(((.....)))... ( -24.90) >DroSec_CAF1 12303 95 + 1 GAAUCAUUAUGCCCAGGUGACCCGACUCUGAUCAUUCCAUCUGAUCAGG-UUUAAGAUCUUGGGCAUUCCCUGUUGUCAUGCGCAUAAUUUGUAAA ((..((..(((((((((......(((.(((((((.......))))))))-))......)))))))))....))...))....(((.....)))... ( -26.80) >DroEre_CAF1 11539 91 + 1 GAAUCAUUAUGCCCAUCUGAUCUAACU-UGAUC---CCAUCUGAUCCUCACCCCCGAUCGUGGGCAU-CCCCGUUGUCUUGUGCAUAAUUUGUAAC .....(((((((.((...(((..(((.-.((((---((((..((((.........))))))))).))-)...)))))).)).)))))))....... ( -19.50) >DroYak_CAF1 12490 80 + 1 GAAUCAUUAUGCCCAGGUGAUCCAACU-UGAUC---CCAUCUGAUCAAC-----------UGGGCAU-CCCCGUUGUCUUGUGCAUAAUUUGUAAC .....(((((((.((((.((((.....-.))))---))....((.((((-----------.(((...-))).)))))).)).)))))))....... ( -21.00) >consensus GAAUCAUUAUGCCCAGGUGACCCAACU_UGAUC___CCAUCUGAUCAGC_UUU_AGAUCUUGGGCAU_CCCCGUUGUCUUGCGCAUAAUUUGUAAA ........(((((((((..........((((((.........))))))..........)))))))))...............(((.....)))... (-15.18 = -16.43 + 1.25)

| Location | 6,457,863 – 6,457,955 |
|---|---|
| Length | 92 |
| Sequences | 4 |
| Columns | 96 |
| Reading direction | reverse |
| Mean pairwise identity | 80.78 |
| Mean single sequence MFE | -24.95 |
| Consensus MFE | -16.92 |
| Energy contribution | -17.93 |
| Covariance contribution | 1.00 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.94 |
| Structure conservation index | 0.68 |
| SVM decision value | 0.73 |
| SVM RNA-class probability | 0.835908 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
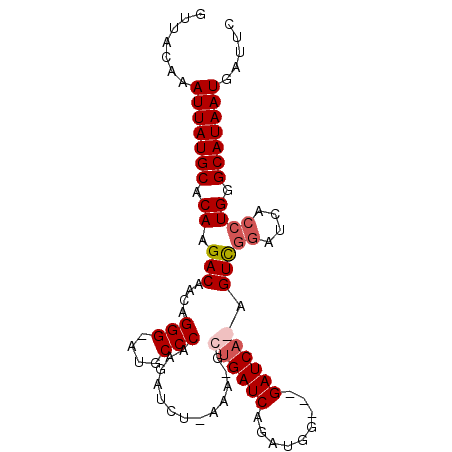
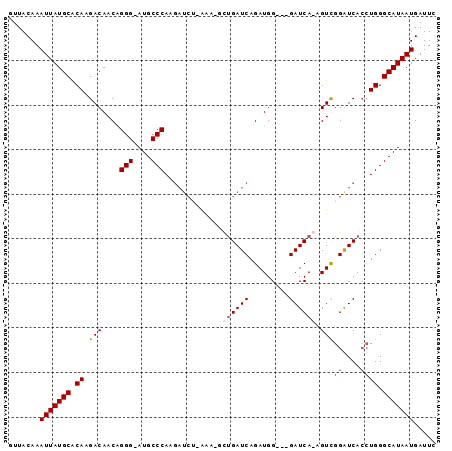
>X_DroMel_CAF1 6457863 92 - 22224390 UUUACAAAUUAUGCGCAAGACAACAGGGAAUGCCCAAGAUCUCAAA-ACUGAUCAGAUGG---GAUCAGAGUCGGGUCACCUGGGCAUAAUGAUUC .......(((((((.((.(((....(((....)))..(((((((..-.((....)).)))---))))...)))((....)))).)))))))..... ( -27.40) >DroSec_CAF1 12303 95 - 1 UUUACAAAUUAUGCGCAUGACAACAGGGAAUGCCCAAGAUCUUAAA-CCUGAUCAGAUGGAAUGAUCAGAGUCGGGUCACCUGGGCAUAAUGAUUC .......(((((((.((.(......(((....)))..(((((...(-((((((((.......))))))).)).))))).).)).)))))))..... ( -28.60) >DroEre_CAF1 11539 91 - 1 GUUACAAAUUAUGCACAAGACAACGGGG-AUGCCCACGAUCGGGGGUGAGGAUCAGAUGG---GAUCA-AGUUAGAUCAGAUGGGCAUAAUGAUUC .......(((((((.((.......(((.-...)))..((((..((.(...((((......---)))).-).)).))))...)).)))))))..... ( -20.80) >DroYak_CAF1 12490 80 - 1 GUUACAAAUUAUGCACAAGACAACGGGG-AUGCCCA-----------GUUGAUCAGAUGG---GAUCA-AGUUGGAUCACCUGGGCAUAAUGAUUC .......(((((((.........(((((-((.((.(-----------.((((((......---)))))-).).))))).)))).)))))))..... ( -23.00) >consensus GUUACAAAUUAUGCACAAGACAACAGGG_AUGCCCAAGAUCU_AAA_GCUGAUCAGAUGG___GAUCA_AGUCGGAUCACCUGGGCAUAAUGAUUC .......(((((((.((.(((....(((....))).............((((((.........)))))).)))((....)))).)))))))..... (-16.92 = -17.93 + 1.00)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:33:53 2006