
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 6,438,694 – 6,438,789 |
| Length | 95 |
| Max. P | 0.934742 |

| Location | 6,438,694 – 6,438,789 |
|---|---|
| Length | 95 |
| Sequences | 6 |
| Columns | 96 |
| Reading direction | forward |
| Mean pairwise identity | 73.94 |
| Mean single sequence MFE | -35.05 |
| Consensus MFE | -17.02 |
| Energy contribution | -17.16 |
| Covariance contribution | 0.14 |
| Combinations/Pair | 1.20 |
| Mean z-score | -1.65 |
| Structure conservation index | 0.49 |
| SVM decision value | -0.07 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 6438694 95 + 22224390 AUUGCUGCGCAUAAUGGCCUCACUAAUGGCCAGCCAAAAGAUGCUGGCCAAAAGACUCAGCGAGAACUAUGUGUCGCGCUAUGAGGUGGUGGCCG- ..(((...)))....((((((((...((((((((........)))))))).....(((((((.((.(.....))).)))..)))))))).)))).- ( -36.20) >DroVir_CAF1 2559 93 + 1 GCUGCUCCGCAUUGUGGCCGCCUUGAUGGCCAGCAACAAGACGCUGCCGCAUCGCCUCAGCGAAGCCUAUGUGGGCCGCUAUGCUGUGU---CCCA (..((.(.((((.((((((........(((.(((........))))))((.((((....)))).)).......)))))).)))).).))---..). ( -34.10) >DroEre_CAF1 1572 92 + 1 GUUGCUGCGCAUAAUGGCCUCACUAAUGGCCAGCAAGAAGAUGCUGGCCAGGAGACUCAGCGAGAACUAUGUAUCGCGCUAUGAGGUUG---CCG- ......(.(((.....((((((((..(((((((((......)))))))))..))....((((.((........)).)))).))))))))---)).- ( -35.30) >DroWil_CAF1 1489 95 + 1 ACUCUUACGCAUAUUGGCUUCCCUAAUGGCCAGCAAGAAAAUGCUGGCGAAUCGCCUGAGCGAGGUCUAUGUCGGUCGCUAUGGAGUCGCGUCUG- (((((...((..((((((..........(((((((......)))))))((.((((....))))..))...)))))).))...)))))........- ( -29.30) >DroYak_CAF1 1825 92 + 1 GUUGCUGCACAUAAUGGCCUCACUAAUGGCCAGCAAGAAGAUGCUGGCCAAAAGACUCAGCGAGAACUAUGUAUCGCGCUAUGAGGUUG---CCG- ...............(((((((((..(((((((((......)))))))))..))....((((.((........)).)))).))))))).---...- ( -32.20) >DroMoj_CAF1 1914 93 + 1 GCUGCUGCGCAUUGUGGCAGCUUUGAUGGCCAGCAACAAGGCGCUGCCGCGGCGCCUCAGCGAUGCCUAUGUGGGCCGCUUUGCGCUGC---CCGA (((((..(.....)..)))))......(((.(((........))))))(((((((...((((..((((....))))))))..)))))))---.... ( -43.20) >consensus GCUGCUGCGCAUAAUGGCCUCACUAAUGGCCAGCAAGAAGAUGCUGGCCAAAAGACUCAGCGAGAACUAUGUGGCGCGCUAUGAGGUGG___CCG_ ................(((((.......((((((........))))))..........((((..............))))..)))))......... (-17.02 = -17.16 + 0.14)

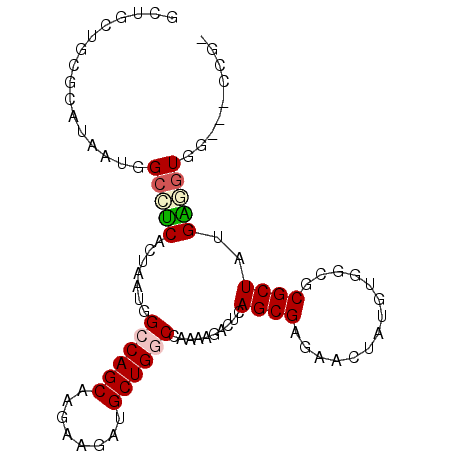

| Location | 6,438,694 – 6,438,789 |
|---|---|
| Length | 95 |
| Sequences | 6 |
| Columns | 96 |
| Reading direction | reverse |
| Mean pairwise identity | 73.94 |
| Mean single sequence MFE | -35.37 |
| Consensus MFE | -15.67 |
| Energy contribution | -16.33 |
| Covariance contribution | 0.67 |
| Combinations/Pair | 1.28 |
| Mean z-score | -2.23 |
| Structure conservation index | 0.44 |
| SVM decision value | 1.24 |
| SVM RNA-class probability | 0.934742 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 6438694 95 - 22224390 -CGGCCACCACCUCAUAGCGCGACACAUAGUUCUCGCUGAGUCUUUUGGCCAGCAUCUUUUGGCUGGCCAUUAGUGAGGCCAUUAUGCGCAGCAAU -.((((.....((((..(((.((........)).))))))).((..((((((((........))))))))..))...))))....(((...))).. ( -34.20) >DroVir_CAF1 2559 93 - 1 UGGG---ACACAGCAUAGCGGCCCACAUAGGCUUCGCUGAGGCGAUGCGGCAGCGUCUUGUUGCUGGCCAUCAAGGCGGCCACAAUGCGGAGCAGC ((((---.(...((...)).))))).....(((((((((.(((....(((((((.....)))))))(((.....))).))).))..)))))))... ( -36.90) >DroEre_CAF1 1572 92 - 1 -CGG---CAACCUCAUAGCGCGAUACAUAGUUCUCGCUGAGUCUCCUGGCCAGCAUCUUCUUGCUGGCCAUUAGUGAGGCCAUUAUGCGCAGCAAC -.(.---...)......(((((...(((((((((((((((......(((((((((......))))))))))))))))))..))))))))).))... ( -36.00) >DroWil_CAF1 1489 95 - 1 -CAGACGCGACUCCAUAGCGACCGACAUAGACCUCGCUCAGGCGAUUCGCCAGCAUUUUCUUGCUGGCCAUUAGGGAAGCCAAUAUGCGUAAGAGU -...(((((.(.((..(((((............)))))..)).)....(((((((......))))))).....((....))....)))))...... ( -27.70) >DroYak_CAF1 1825 92 - 1 -CGG---CAACCUCAUAGCGCGAUACAUAGUUCUCGCUGAGUCUUUUGGCCAGCAUCUUCUUGCUGGCCAUUAGUGAGGCCAUUAUGUGCAGCAAC -.(.---...)........((..(((((((((((((((((......(((((((((......))))))))))))))))))..))))))))..))... ( -37.30) >DroMoj_CAF1 1914 93 - 1 UCGG---GCAGCGCAAAGCGGCCCACAUAGGCAUCGCUGAGGCGCCGCGGCAGCGCCUUGUUGCUGGCCAUCAAAGCUGCCACAAUGCGCAGCAGC ..((---.((((((..(((((((......)))..))))(((((((.......)))))))).))))).))......((((((.....).)))))... ( -40.10) >consensus _CGG___CCACCUCAUAGCGCCACACAUAGUCCUCGCUGAGGCGAUUCGCCAGCAUCUUCUUGCUGGCCAUUAGUGAGGCCAUUAUGCGCAGCAAC ..........((((.(((((..((.....))...))))).........(((((((......))))))).......))))................. (-15.67 = -16.33 + 0.67)


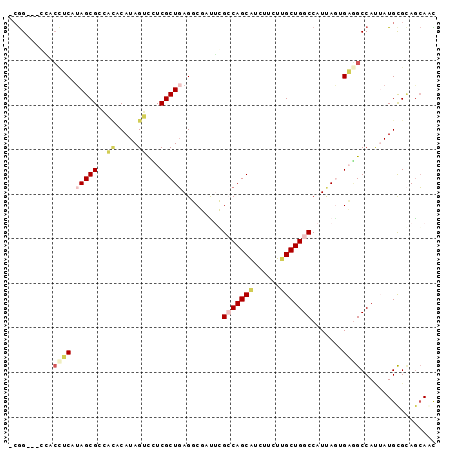
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:33:44 2006