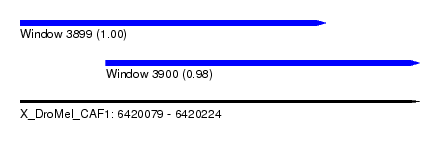

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 6,420,079 – 6,420,224 |
| Length | 145 |
| Max. P | 0.995560 |

| Location | 6,420,079 – 6,420,190 |
|---|---|
| Length | 111 |
| Sequences | 5 |
| Columns | 111 |
| Reading direction | forward |
| Mean pairwise identity | 82.83 |
| Mean single sequence MFE | -33.54 |
| Consensus MFE | -25.78 |
| Energy contribution | -26.78 |
| Covariance contribution | 1.00 |
| Combinations/Pair | 1.13 |
| Mean z-score | -3.01 |
| Structure conservation index | 0.77 |
| SVM decision value | 2.59 |
| SVM RNA-class probability | 0.995560 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
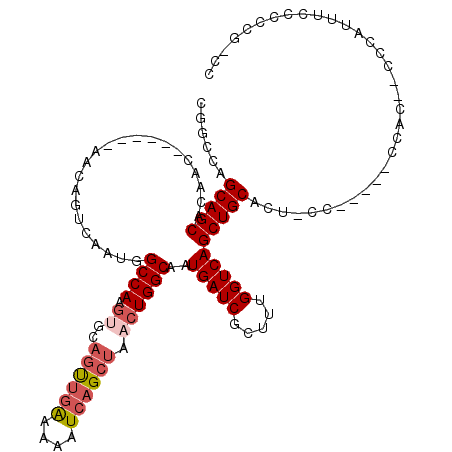
>X_DroMel_CAF1 6420079 111 + 22224390 ACCGAAGGCUGAGAGAAAAACGACUGUUGGCCGGCCAGCAGCACAACAAAAUGGUCAGUCAAUGGCCAAGUGCAGUUGAAAAAUCAGCUAACUGGCAAUGAUCGCUUUGGU (((((((((..(.((........)).)..))).(((((..((((.......((((((.....)))))).))))((((((....))))))..))))).........)))))) ( -38.90) >DroSec_CAF1 37047 105 + 1 ACCGAAGACUAAAAGAAAAACGACUGUUGGCCGGCCAGCAGCACAAC------AACAGUCAAUGGCCAAGUGCAGUGUGCAAAUCAGCUAACUGGCAAUGAUCGCUUUGGU (((((((..............((((((((....((.....))....)------)))))))...(..((..((((((..((......))..))).))).))..).))))))) ( -30.10) >DroSim_CAF1 36135 111 + 1 ACCGAAGGCUAAAAGAAAAACGACUGUUGGCCGGCCAGCAGCACAACAACAACAACAGUCAAUGGCCAAGUGCAGUUGAAAAAUCAGCUAACUGGCAAUGAUCGCUUUGGU (((((((..............(.(((((((....))))))))...............((((...((((.((..((((((....)))))).))))))..))))..))))))) ( -34.20) >DroEre_CAF1 37086 96 + 1 AUCACAGGCAGCGAGAUGAACGACUGUUGGCC----AGCAGC-----------AGCAGUCAAUGGCCAAGUGCAGCUGAAAAAUCAGCUAACUGGCAAUGAUCGCUUUGGU ....(((..(((((.......((((((((...----.....)-----------)))))))....((((.((..((((((....)))))).)))))).....)))))))).. ( -34.50) >DroYak_CAF1 39943 96 + 1 AUCAAUGGCAGAGAGAAAAACGACUGUUGACC----AGCAGC-----------AACAGUCAAUGGCCAAGUGCAGUUGAAAAAUCAGCUACCUGGCAAUGAUCGCUUUGGU ........(((((.((.....((((((((...----.....)-----------)))))))....((((.((..((((((....)))))))).)))).....)).))))).. ( -30.00) >consensus ACCGAAGGCUGAGAGAAAAACGACUGUUGGCCGGCCAGCAGCACAAC______AACAGUCAAUGGCCAAGUGCAGUUGAAAAAUCAGCUAACUGGCAAUGAUCGCUUUGGU (((((((..............(.(((((((....))))))))...............((((...((((.((..((((((....)))))).))))))..))))..))))))) (-25.78 = -26.78 + 1.00)


| Location | 6,420,110 – 6,420,224 |
|---|---|
| Length | 114 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 75.06 |
| Mean single sequence MFE | -29.08 |
| Consensus MFE | -20.95 |
| Energy contribution | -22.07 |
| Covariance contribution | 1.11 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.89 |
| Structure conservation index | 0.72 |
| SVM decision value | 1.79 |
| SVM RNA-class probability | 0.977368 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 6420110 114 + 22224390 CGGCCAGCAGCACAACAAAAUGGUCAGUCAAUGGCCAAGUGCAGUUGAAAAAUCAGCUAACUGGCAAUGAUCGCUUUGGUCAGCUGCACUGCC-----CCACUCCCCAUUUCCCCCG-CC .(((..(((((....((((.((((((.......((((.((..((((((....)))))).))))))..)))))).))))....)))))...)))-----...................-.. ( -34.50) >DroSec_CAF1 37078 107 + 1 CGGCCAGCAGCACAAC------AACAGUCAAUGGCCAAGUGCAGUGUGCAAAUCAGCUAACUGGCAAUGAUCGCUUUGGUCAGCUGCACU-CC-----CCACCACCCAUUUCCCCCG-CC .(((((...((.....------....))...))))).((((((((..........(((....)))..(((((.....)))))))))))))-..-----...................-.. ( -27.50) >DroSim_CAF1 36166 113 + 1 CGGCCAGCAGCACAACAACAACAACAGUCAAUGGCCAAGUGCAGUUGAAAAAUCAGCUAACUGGCAAUGAUCGCUUUGGUCAGCUGCACU-CC-----CCACCACCCAUUUCCCCCG-CC .(((((...((...............))...))))).(((((((((((..((.(((....)))((.......)).))..)))))))))))-..-----...................-.. ( -27.76) >DroEre_CAF1 37117 91 + 1 C----AGCAGC-----------AGCAGUCAAUGGCCAAGUGCAGCUGAAAAAUCAGCUAACUGGCAAUGAUCGCUUUGGUCAGCUGCACU-CC------------CCAUUUCCCCCG-CC .----.(((((-----------(((.((((...((((.((..((((((....)))))).))))))..)))).))).......)))))...-..------------............-.. ( -30.21) >DroYak_CAF1 39974 90 + 1 C----AGCAGC-----------AACAGUCAAUGGCCAAGUGCAGUUGAAAAAUCAGCUACCUGGCAAUGAUCGCUUUGGUCAGCUGCACU-CC------------CCAUUUCCCCC--CC .----.(((((-----------...........((((.((..((((((....)))))))).))))..(((((.....))))))))))...-..------------...........--.. ( -25.90) >DroAna_CAF1 49059 102 + 1 AGAAGAGCAGCAAACCAUAAAUAACAGUCAGUGGCCAAGUGCAG-----------------UGGCAAUGAUCGCUUUGGUCAGCUGCACU-CUAGCUGCCACGCCCCCUCCGCCCUGAUC ..........................(((((.(((..((.((.(-----------------(((((.(((((.....)))))((((....-.))))))))))))...))..)))))))). ( -28.60) >consensus CGGCCAGCAGCACAAC______AACAGUCAAUGGCCAAGUGCAGUUGAAAAAUCAGCUAACUGGCAAUGAUCGCUUUGGUCAGCUGCACU_CC_____CCAC__CCCAUUUCCCCCG_CC ......(((((......................((((.((..((((((....)))))).))))))..(((((.....))))))))))................................. (-20.95 = -22.07 + 1.11)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:33:31 2006