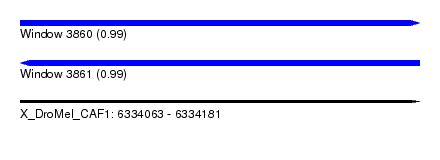

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 6,334,063 – 6,334,181 |
| Length | 118 |
| Max. P | 0.993065 |

| Location | 6,334,063 – 6,334,181 |
|---|---|
| Length | 118 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 84.93 |
| Mean single sequence MFE | -24.21 |
| Consensus MFE | -19.16 |
| Energy contribution | -20.23 |
| Covariance contribution | 1.06 |
| Combinations/Pair | 1.04 |
| Mean z-score | -3.00 |
| Structure conservation index | 0.79 |
| SVM decision value | 2.37 |
| SVM RNA-class probability | 0.993065 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
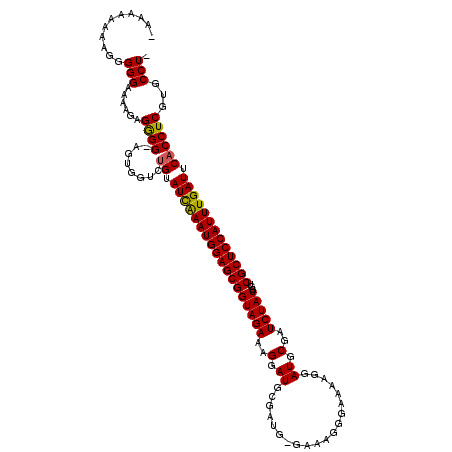
>X_DroMel_CAF1 6334063 118 + 22224390 -AGGCACGAGGUGAAUCAAAUGGAGCGACCAUAGAUCGCAUCCUUUUCCCUUUCCCAUCGCAUCCUUUCUACCGCUCCAUUCGAUACAGACCACU-CCCCUUGUUUCCCCCCUUUUUUUC -.((.((((((.(((((.(((((((((......(((.((....................)))))........))))))))).))).........)-).))))))..))............ ( -23.39) >DroSec_CAF1 18637 116 + 1 -AGGCACGAGGUGAAUCAAAUGGAGCGACCAUAGAUCGCAUCCUUUUCCCUUUC-CAUCGCAUCCUUUCUACCGCUCCAUUUGAUACAGACCACU-CCCCGCUUUUCCCCCUUUUUUUU- -((((..(((.((.(((((((((((((......(((.((...............-....)))))........))))))))))))).)).....))-)...))))...............- ( -25.15) >DroEre_CAF1 16222 117 + 1 AAGGCACGAGGCGAAUCAAAUGGAGCGACCAUAGAGAGCAUCUUCCCACCUUUC-CAUCGCAUCCUUUCUACCGCUCCAUUUAAUACUAA-CACUUCCCCUCUUCUCCCAUUUUUUUUU- ..((...((((.(((..((((((((((....(((((((................-.........))))))).))))))))))........-...))).)))).....))..........- ( -22.93) >DroYak_CAF1 15003 109 + 1 -AGGCACGAGGUGAAUCAAAUGGAGCGACCAUAGAUCGCAUC------CCUUAC-CAUCGCAUCCUUUCUACCGCUCCAUUUGAUACAAACCACUUCCUCUCUUUUCCCCAUUUUGC--- -......((((((.(((((((((((((......(((.((...------......-....)))))........)))))))))))))......))))))....................--- ( -25.36) >consensus _AGGCACGAGGUGAAUCAAAUGGAGCGACCAUAGAUCGCAUCCUUUUCCCUUUC_CAUCGCAUCCUUUCUACCGCUCCAUUUGAUACAAACCACU_CCCCUCUUUUCCCCCUUUUUUUU_ ..((...((((((.(((((((((((((....((((..((....................))......)))).)))))))))))))......)))))).)).................... (-19.16 = -20.23 + 1.06)


| Location | 6,334,063 – 6,334,181 |
|---|---|
| Length | 118 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 84.93 |
| Mean single sequence MFE | -34.21 |
| Consensus MFE | -27.39 |
| Energy contribution | -27.08 |
| Covariance contribution | -0.31 |
| Combinations/Pair | 1.10 |
| Mean z-score | -2.68 |
| Structure conservation index | 0.80 |
| SVM decision value | 2.03 |
| SVM RNA-class probability | 0.986211 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 6334063 118 - 22224390 GAAAAAAAGGGGGGAAACAAGGGG-AGUGGUCUGUAUCGAAUGGAGCGGUAGAAAGGAUGCGAUGGGAAAGGGAAAAGGAUGCGAUCUAUGGUCGCUCCAUUUGAUUCACCUCGUGCCU- .........((((....)...(((-.((((.....((((((((((((((((((..(.((.(................).)).)..)))))...))))))))))))))))))))...)))- ( -33.99) >DroSec_CAF1 18637 116 - 1 -AAAAAAAAGGGGGAAAAGCGGGG-AGUGGUCUGUAUCAAAUGGAGCGGUAGAAAGGAUGCGAUG-GAAAGGGAAAAGGAUGCGAUCUAUGGUCGCUCCAUUUGAUUCACCUCGUGCCU- -..........(((....((((((-((....))(.((((((((((((((((((..(.((.(....-...........).)).)..)))))...))))))))))))).).)))))).)))- ( -36.56) >DroEre_CAF1 16222 117 - 1 -AAAAAAAAAUGGGAGAAGAGGGGAAGUG-UUAGUAUUAAAUGGAGCGGUAGAAAGGAUGCGAUG-GAAAGGUGGGAAGAUGCUCUCUAUGGUCGCUCCAUUUGAUUCGCCUCGUGCCUU -..........(((....((((.((....-.....(((((((((((((...........((.(((-((..((((......)))).))))).))))))))))))))))).))))...))). ( -33.40) >DroYak_CAF1 15003 109 - 1 ---GCAAAAUGGGGAAAAGAGAGGAAGUGGUUUGUAUCAAAUGGAGCGGUAGAAAGGAUGCGAUG-GUAAGG------GAUGCGAUCUAUGGUCGCUCCAUUUGAUUCACCUCGUGCCU- ---..................(((..(.(((....((((((((((((((((.......)))(((.-(((...------..))).)))......)))))))))))))..))).)...)))- ( -32.90) >consensus _AAAAAAAAGGGGGAAAAGAGGGG_AGUGGUCUGUAUCAAAUGGAGCGGUAGAAAGGAUGCGAUG_GAAAGGGAAAAGGAUGCGAUCUAUGGUCGCUCCAUUUGAUUCACCUCGUGCCU_ ...........(((......((((........((.((((((((((((((((((..(.((....................)).)..)))))...))))))))))))).))))))...))). (-27.39 = -27.08 + -0.31)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:32:57 2006