
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 6,288,946 – 6,289,062 |
| Length | 116 |
| Max. P | 0.990537 |

| Location | 6,288,946 – 6,289,062 |
|---|---|
| Length | 116 |
| Sequences | 3 |
| Columns | 116 |
| Reading direction | forward |
| Mean pairwise identity | 79.75 |
| Mean single sequence MFE | -19.56 |
| Consensus MFE | -16.47 |
| Energy contribution | -16.81 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.33 |
| Structure conservation index | 0.84 |
| SVM decision value | 2.22 |
| SVM RNA-class probability | 0.990537 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
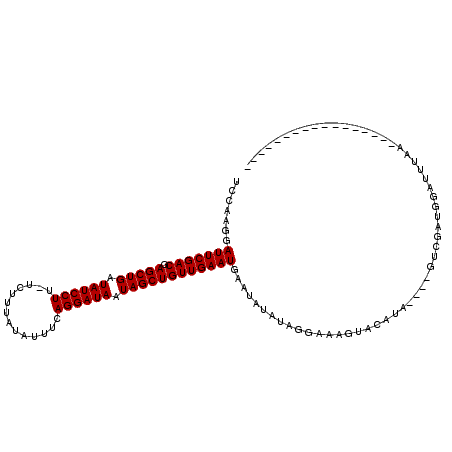
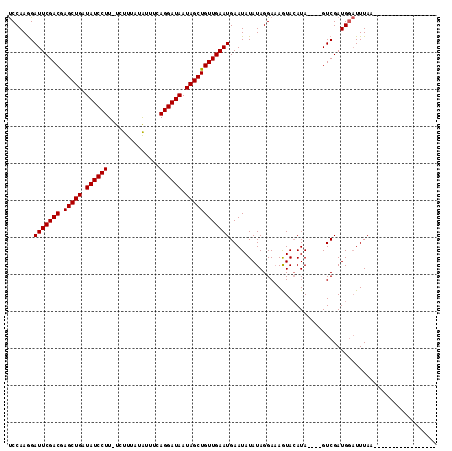
>X_DroMel_CAF1 6288946 116 + 22224390 UGAAUUUAAAAAUGUGUUUAAAAGCAUCGACUAACUAUGUAUUGUCCUAUAUAUUCAUUCAACAGCUAUUAUCCUGAAACAAAAAAACAAGGAUAUCAGCUCGUCGAAUCCUUGGG ((..((((((......))))))..))(((((....((((((......))))))..........((((..((((((..............))))))..)))).)))))......... ( -19.94) >DroSec_CAF1 26105 94 + 1 -----------------UUAAAUCCAUCGAC----UAUUUACUUUCCUAUAUAUUCAUUCAACAGCUAUUAUCCUGAAAUAUAAAGA-AAGGAUAUCAGCUCGUCGAAUCUUUGGA -----------------.....((((..((.----(((.((......)).))).))((((.((((((..((((((............-.))))))..)))).)).))))...)))) ( -17.42) >DroSim_CAF1 24056 94 + 1 -----------------UUAAAUCCAUCGAC----UAUGUACUUUCCUAUAUAUUCAUUCAACAGCUAUUAUCCUGAAAUAUAAAGA-AAGGAUAUCAGCUUGUCGAAUCCUUGGA -----------------.....((((..((.----((((((......)))))).))((((.((((((..((((((............-.))))))..))).))).))))...)))) ( -21.32) >consensus _________________UUAAAUCCAUCGAC____UAUGUACUUUCCUAUAUAUUCAUUCAACAGCUAUUAUCCUGAAAUAUAAAGA_AAGGAUAUCAGCUCGUCGAAUCCUUGGA ..........................(((((....((((((......))))))..........((((..((((((..............))))))..)))).)))))......... (-16.47 = -16.81 + 0.33)



| Location | 6,288,946 – 6,289,062 |
|---|---|
| Length | 116 |
| Sequences | 3 |
| Columns | 116 |
| Reading direction | reverse |
| Mean pairwise identity | 79.75 |
| Mean single sequence MFE | -21.76 |
| Consensus MFE | -18.54 |
| Energy contribution | -18.54 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.59 |
| Structure conservation index | 0.85 |
| SVM decision value | 1.04 |
| SVM RNA-class probability | 0.905430 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 6288946 116 - 22224390 CCCAAGGAUUCGACGAGCUGAUAUCCUUGUUUUUUUGUUUCAGGAUAAUAGCUGUUGAAUGAAUAUAUAGGACAAUACAUAGUUAGUCGAUGCUUUUAAACACAUUUUUAAAUUCA .......(((((((.(((((.((((((..............)))))).))))))))))))((((......((((((.....))).))).......(((((......))))))))). ( -21.64) >DroSec_CAF1 26105 94 - 1 UCCAAAGAUUCGACGAGCUGAUAUCCUU-UCUUUAUAUUUCAGGAUAAUAGCUGUUGAAUGAAUAUAUAGGAAAGUAAAUA----GUCGAUGGAUUUAA----------------- ((((...(((((((.(((((.((((((.-............)))))).))))))))))))((.(((.((......)).)))----.))..)))).....----------------- ( -21.72) >DroSim_CAF1 24056 94 - 1 UCCAAGGAUUCGACAAGCUGAUAUCCUU-UCUUUAUAUUUCAGGAUAAUAGCUGUUGAAUGAAUAUAUAGGAAAGUACAUA----GUCGAUGGAUUUAA----------------- ((((...((((((((.((((.((((((.-............)))))).))))))))))))((...(((.(.......))))----.))..)))).....----------------- ( -21.92) >consensus UCCAAGGAUUCGACGAGCUGAUAUCCUU_UCUUUAUAUUUCAGGAUAAUAGCUGUUGAAUGAAUAUAUAGGAAAGUACAUA____GUCGAUGGAUUUAA_________________ .......(((((((.(((((.((((((..............)))))).))))))))))))........................................................ (-18.54 = -18.54 + -0.00)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:32:40 2006