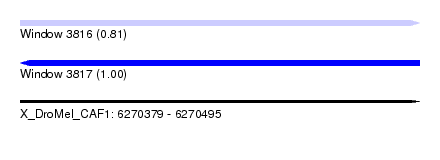

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 6,270,379 – 6,270,495 |
| Length | 116 |
| Max. P | 0.995334 |

| Location | 6,270,379 – 6,270,495 |
|---|---|
| Length | 116 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 78.34 |
| Mean single sequence MFE | -21.23 |
| Consensus MFE | -14.29 |
| Energy contribution | -14.60 |
| Covariance contribution | 0.31 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.83 |
| Structure conservation index | 0.67 |
| SVM decision value | 0.66 |
| SVM RNA-class probability | 0.814813 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 6270379 116 + 22224390 ----UGAACUACGCAUUACGUUAAACCGGUUUCGAAUGAAACCGAAAUGCAAAUCAGUUCUGCAUUAUUUACUAAGUAGUUCUUAUUUAACUGCGACGCAAUUCAACAACGCAAUUCAAC ----.(((((((..............(((((((....))))))).((((((.........)))))).........))))))).........((((..(........)..))))....... ( -26.00) >DroSec_CAF1 12603 103 + 1 ----UGAACUACGCAUUACGUUAAAUCAGUUUCGAAUGAAACCGAAAUGCAAAACAGUUCUGCAUUAUUUACCAAGUAGUUCUUACUUAACUGCGACGCAAUU-------------CAAC ----.(((((((.............((.(((((....))))).))((((((.........)))))).........))))))).........(((...)))...-------------.... ( -18.50) >DroEre_CAF1 1648 104 + 1 UUCUUGAACUACGCAUUACGUUAAGCCA-UUUCGAAUGAAACCGAAAUGCAAAACAGUUCUGCAUUGUUUACCAAGUAGUUCCUAUGUAACUACGAC--AAUU-------------CUUC .....((((((((((..(((((..((.(-(((((........))))))))..))).))..))).(((.....))))))))))...............--....-------------.... ( -21.30) >DroYak_CAF1 9821 104 + 1 UUUAUGAAUUACGCAUUACUGUAAACCA-UUUCGAAUGAAAAGGAAAUGCAAAAUAGGAUUGCAUUACUUACUAAGUAGUUCGCAUGUAACUACGAC--AAUU-------------CGUC ...(((((((..(((...((((....((-((((..........))))))....))))...)))............((((((.......))))))...--))))-------------))). ( -19.10) >consensus ____UGAACUACGCAUUACGUUAAACCA_UUUCGAAUGAAACCGAAAUGCAAAACAGUUCUGCAUUAUUUACCAAGUAGUUCUUAUGUAACUACGAC__AAUU_____________CAAC .....(((((((..............(.(((((....))))).).((((((.........)))))).........)))))))...................................... (-14.29 = -14.60 + 0.31)

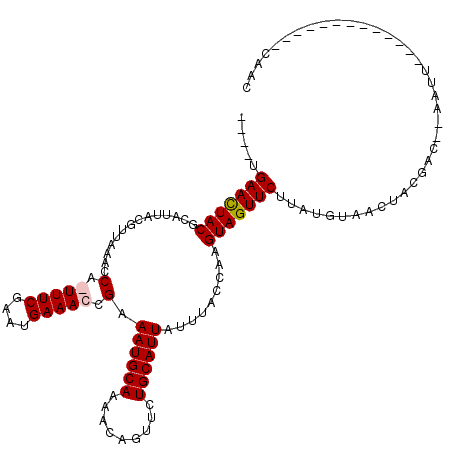
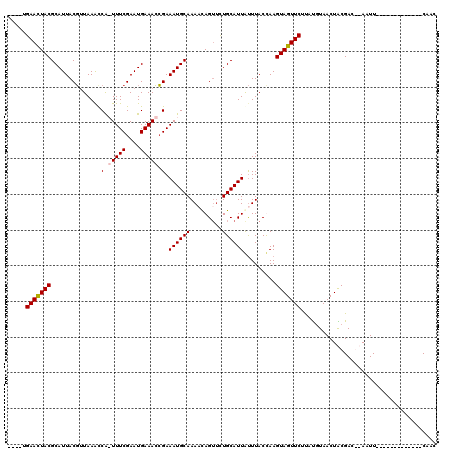
| Location | 6,270,379 – 6,270,495 |
|---|---|
| Length | 116 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 78.34 |
| Mean single sequence MFE | -28.55 |
| Consensus MFE | -19.60 |
| Energy contribution | -20.35 |
| Covariance contribution | 0.75 |
| Combinations/Pair | 1.13 |
| Mean z-score | -2.57 |
| Structure conservation index | 0.69 |
| SVM decision value | 2.57 |
| SVM RNA-class probability | 0.995334 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 6270379 116 - 22224390 GUUGAAUUGCGUUGUUGAAUUGCGUCGCAGUUAAAUAAGAACUACUUAGUAAAUAAUGCAGAACUGAUUUGCAUUUCGGUUUCAUUCGAAACCGGUUUAACGUAAUGCGUAGUUCA---- (((.(((((((.(((......))).))))))).)))..(((((((.........((((((((.....))))))))((((((((....)))))))).............))))))).---- ( -35.50) >DroSec_CAF1 12603 103 - 1 GUUG-------------AAUUGCGUCGCAGUUAAGUAAGAACUACUUGGUAAAUAAUGCAGAACUGUUUUGCAUUUCGGUUUCAUUCGAAACUGAUUUAACGUAAUGCGUAGUUCA---- ..((-------------((((((((....(((((((..................((((((((.....)))))))).(((((((....)))))))))))))).....))))))))))---- ( -29.30) >DroEre_CAF1 1648 104 - 1 GAAG-------------AAUU--GUCGUAGUUACAUAGGAACUACUUGGUAAACAAUGCAGAACUGUUUUGCAUUUCGGUUUCAUUCGAAA-UGGCUUAACGUAAUGCGUAGUUCAAGAA ....-------------....--...............((((((((((.....))).(((..((.(((..(((((((((......))))))-).))..)))))..))))))))))..... ( -25.60) >DroYak_CAF1 9821 104 - 1 GACG-------------AAUU--GUCGUAGUUACAUGCGAACUACUUAGUAAGUAAUGCAAUCCUAUUUUGCAUUUCCUUUUCAUUCGAAA-UGGUUUACAGUAAUGCGUAAUUCAUAAA ...(-------------((((--(.((((.((((.((..(((((........(.(((((((.......))))))).)..((((....))))-)))))..))))))))))))))))..... ( -23.80) >consensus GAUG_____________AAUU__GUCGCAGUUAAAUAAGAACUACUUAGUAAAUAAUGCAGAACUGUUUUGCAUUUCGGUUUCAUUCGAAA_UGGUUUAACGUAAUGCGUAGUUCA____ ......................................(((((((.........((((((((.....))))))))((((((((....)))))))).............)))))))..... (-19.60 = -20.35 + 0.75)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:32:18 2006