
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 6,248,021 – 6,248,136 |
| Length | 115 |
| Max. P | 0.979267 |

| Location | 6,248,021 – 6,248,136 |
|---|---|
| Length | 115 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 84.17 |
| Mean single sequence MFE | -42.65 |
| Consensus MFE | -32.99 |
| Energy contribution | -35.05 |
| Covariance contribution | 2.06 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.56 |
| Structure conservation index | 0.77 |
| SVM decision value | 1.83 |
| SVM RNA-class probability | 0.979267 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
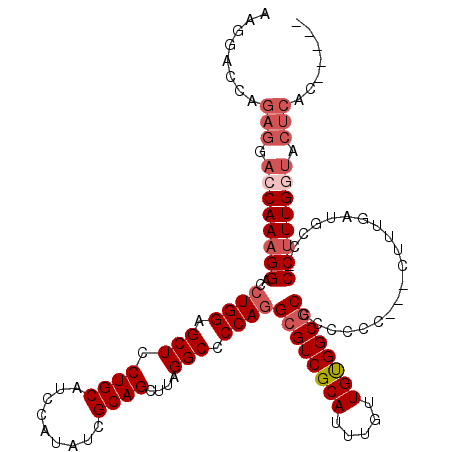
>X_DroMel_CAF1 6248021 115 + 22224390 AAGGACCAGAGGACCAAAGGACCUGGAGCUUCUGCAUCCAUAUCGCAGUUUUGGCCCCAGGCGUCGCAUUUGUUGUGGCCCCCCCCCCUCUUUUGAUGCCUCCUUUGGUACUCAU----- .((.(((((((((.((((((((((((.(((.((((.........))))....))).))))).((((((.....)))))).........))))))).....))))))))).))...----- ( -41.60) >DroSec_CAF1 14846 112 + 1 AAGGACCAGAGGACCAAAGGACCUGGAGCUCCUGCAUCCAUAUCGCAGCUUUGGCCCCAGGCGUCGCAUUUGUUGUGGCGCCCCCCC---AUUUGAUGCCUCCUUUGGUACUCAC----- ........(((.(((((((((..((((((....)).))))....(((....(((.....(((((((((.....)))))))))...))---).....))).))))))))).)))..----- ( -44.90) >DroEre_CAF1 9549 100 + 1 AAGGGGCAGAGGACCAAAGGACCUGGAGCUCCUGCAUCCAUAUCGCAGCUUAGGCCCCAGGCGUCGCAUUUGUUGCGGCGCC---CC---CUUUGAUGCC-CCUUUG------------- ((((((((......((((((...(((.(((.((((.........))))....))).)))(((((((((.....)))))))))---.)---))))).))))-))))..------------- ( -47.00) >DroYak_CAF1 21941 116 + 1 GAGAAGCAGAGGACCAAAGGACCUGGAGCUCCUGCAUCCAUAUCGCAGUUUAGGCCCCAAGCGUCGCAUUUGUUGUGGCGCCCCCCC---CUUUGAUGCC-CCCUUGAUGCUCCCCUCGA (((.((((((((..((((((...(((.(((.((((.........))))....))).))).((((((((.....)))))))).....)---))))).....-.))))..))))...))).. ( -37.10) >consensus AAGGACCAGAGGACCAAAGGACCUGGAGCUCCUGCAUCCAUAUCGCAGCUUAGGCCCCAGGCGUCGCAUUUGUUGUGGCGCCCCCCC___CUUUGAUGCC_CCUUUGGUACUCAC_____ ........(((.((((((((..((((.(((.((((.........))))....))).))))((((((((.....))))))))....................)))))))).)))....... (-32.99 = -35.05 + 2.06)


| Location | 6,248,021 – 6,248,136 |
|---|---|
| Length | 115 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 84.17 |
| Mean single sequence MFE | -42.54 |
| Consensus MFE | -33.58 |
| Energy contribution | -34.33 |
| Covariance contribution | 0.75 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.40 |
| Structure conservation index | 0.79 |
| SVM decision value | -0.07 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 6248021 115 - 22224390 -----AUGAGUACCAAAGGAGGCAUCAAAAGAGGGGGGGGGGCCACAACAAAUGCGACGCCUGGGGCCAAAACUGCGAUAUGGAUGCAGAAGCUCCAGGUCCUUUGGUCCUCUGGUCCUU -----..(((.(((((((((.((((........((.......)).......))))....((((((((.....(((((.......)))))..))))))))))))))))).)))........ ( -39.56) >DroSec_CAF1 14846 112 - 1 -----GUGAGUACCAAAGGAGGCAUCAAAU---GGGGGGGCGCCACAACAAAUGCGACGCCUGGGGCCAAAGCUGCGAUAUGGAUGCAGGAGCUCCAGGUCCUUUGGUCCUCUGGUCCUU -----..(((.(((((((((.((((....(---(((......)).))....))))....((((((((.....(((((.......)))))..))))))))))))))))).)))........ ( -41.00) >DroEre_CAF1 9549 100 - 1 -------------CAAAGG-GGCAUCAAAG---GG---GGCGCCGCAACAAAUGCGACGCCUGGGGCCUAAGCUGCGAUAUGGAUGCAGGAGCUCCAGGUCCUUUGGUCCUCUGCCCCUU -------------..((((-((((......---.(---((((.((((.....)))).)))))((((((.(((....(((.((((.((....)))))).)))))).)))))).)))))))) ( -46.10) >DroYak_CAF1 21941 116 - 1 UCGAGGGGAGCAUCAAGGG-GGCAUCAAAG---GGGGGGGCGCCACAACAAAUGCGACGCUUGGGGCCUAAACUGCGAUAUGGAUGCAGGAGCUCCAGGUCCUUUGGUCCUCUGCUUCUC ....((((((((....(((-(((..(((((---(......(((..........)))..(((((((((.....(((((.......)))))..))))))))))))))))))))))))))))) ( -43.50) >consensus _____GUGAGUACCAAAGG_GGCAUCAAAG___GGGGGGGCGCCACAACAAAUGCGACGCCUGGGGCCAAAACUGCGAUAUGGAUGCAGGAGCUCCAGGUCCUUUGGUCCUCUGCUCCUU .......(((.((((((((.(((..(............)..)))..............(((((((((.....(((((.......)))))..))))))))))))))))).)))........ (-33.58 = -34.33 + 0.75)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:31:18 2006