
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 6,236,415 – 6,236,526 |
| Length | 111 |
| Max. P | 0.949293 |

| Location | 6,236,415 – 6,236,526 |
|---|---|
| Length | 111 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 84.06 |
| Mean single sequence MFE | -24.70 |
| Consensus MFE | -22.66 |
| Energy contribution | -22.98 |
| Covariance contribution | 0.31 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.96 |
| Structure conservation index | 0.92 |
| SVM decision value | 1.39 |
| SVM RNA-class probability | 0.949293 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 6236415 111 + 22224390 ---------UCUGAUUCUGAUUCUGAUUCCUAUUCCGGAUUCGGAAUCCGCUUCUUAUUCCGGCUGCCUUGAUUACGUGGCCAAAUCGAUUUCUUUAUCUGCCACCCCCCGCCCCCCUCG ---------...((..(.((((((((.(((......))).)))))))).)..)).......(((.(((.((....)).)))......(((......))).)))................. ( -24.60) >DroSec_CAF1 9407 101 + 1 -----------CGAUUCCGAUUCUGAUUCCUAUUCCGGAUUCGGAAUCCGAUUCCUAUUCCGGCUGCCUUGAUUACGUGGCCAAAUCGAUUUCUUUAUCUGCC--------CCCCGCUCG -----------.((.((.((((((((.(((......))).)))))))).)).)).......(((.(((.((....)).)))......(((......))).)))--------......... ( -25.60) >DroEre_CAF1 4080 109 + 1 CAUGCCGAUUCCGAUUCCGAUUCAGAUUCCUAUUCCGGAUUCCGAAUCCGAUUCUUAUUCCGGCUGCCUUGAUUACGUGGCCAAAUCGAUUUCUUUAUCUGCC--------CC---CUCG .....(((((..((.((.(((((.((.(((......))).)).))))).)).)).......(((..(.........)..))).)))))...............--------..---.... ( -25.10) >DroYak_CAF1 17076 108 + 1 CAUUCACAUUGCGAUUCCGAUUCAGAUUCCUAUUCCGGAUUCGGAAUCCGAUUCUUAUUCCGGCUGCCUUGAUUACGUGGCCAAAUCGAUUUCUUUAUCUGCC--------CU---C-CG .......((((.(((((((((((.((.......)).))).)))))))))))).........(((.(((.((....)).)))......(((......))).)))--------..---.-.. ( -23.50) >consensus _________UCCGAUUCCGAUUCAGAUUCCUAUUCCGGAUUCGGAAUCCGAUUCUUAUUCCGGCUGCCUUGAUUACGUGGCCAAAUCGAUUUCUUUAUCUGCC________CC___CUCG ...........(((((..((((((((.(((......))).)))))))).............(((..(.........)..))).)))))................................ (-22.66 = -22.98 + 0.31)



| Location | 6,236,415 – 6,236,526 |
|---|---|
| Length | 111 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 84.06 |
| Mean single sequence MFE | -30.20 |
| Consensus MFE | -24.25 |
| Energy contribution | -24.50 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.01 |
| Structure conservation index | 0.80 |
| SVM decision value | 1.03 |
| SVM RNA-class probability | 0.902669 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 6236415 111 - 22224390 CGAGGGGGGCGGGGGGUGGCAGAUAAAGAAAUCGAUUUGGCCACGUAAUCAAGGCAGCCGGAAUAAGAAGCGGAUUCCGAAUCCGGAAUAGGAAUCAGAAUCAGAAUCAGA--------- .......(((.....(((((((((..(....)..)))).)))))((.......)).))).......((..(.(((((.((.(((......))).)).))))).)..))...--------- ( -27.90) >DroSec_CAF1 9407 101 - 1 CGAGCGGGG--------GGCAGAUAAAGAAAUCGAUUUGGCCACGUAAUCAAGGCAGCCGGAAUAGGAAUCGGAUUCCGAAUCCGGAAUAGGAAUCAGAAUCGGAAUCG----------- ((((((..(--------(.(((((..(....)..))))).)).)))...........((......))..)))((((((((.((.((........)).)).)))))))).----------- ( -27.80) >DroEre_CAF1 4080 109 - 1 CGAG---GG--------GGCAGAUAAAGAAAUCGAUUUGGCCACGUAAUCAAGGCAGCCGGAAUAAGAAUCGGAUUCGGAAUCCGGAAUAGGAAUCUGAAUCGGAAUCGGAAUCGGCAUG ....---.(--------(.(((((..(....)..))))).))..............(((((.....((.((.((((((((.(((......))).)))))))).)).))....)))))... ( -36.40) >DroYak_CAF1 17076 108 - 1 CG-G---AG--------GGCAGAUAAAGAAAUCGAUUUGGCCACGUAAUCAAGGCAGCCGGAAUAAGAAUCGGAUUCCGAAUCCGGAAUAGGAAUCUGAAUCGGAAUCGCAAUGUGAAUG ((-.---.(--------(.(((((..(....)..))))).)).))...(((..((..((((....(((.((..((((((....))))))..)).)))...))))....))....)))... ( -28.70) >consensus CGAG___GG________GGCAGAUAAAGAAAUCGAUUUGGCCACGUAAUCAAGGCAGCCGGAAUAAGAAUCGGAUUCCGAAUCCGGAAUAGGAAUCAGAAUCGGAAUCGGA_________ .................(((.(((......)))......(((..........))).))).......((.((.(((((.((.(((......))).)).))))).)).))............ (-24.25 = -24.50 + 0.25)

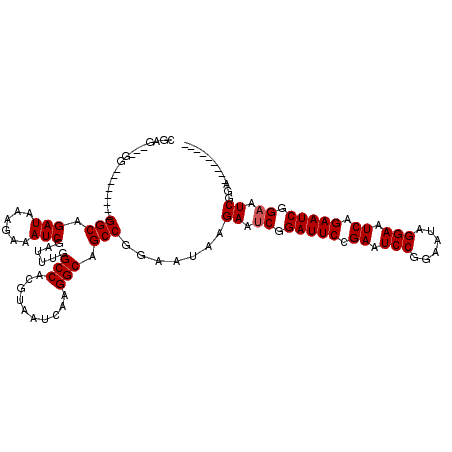
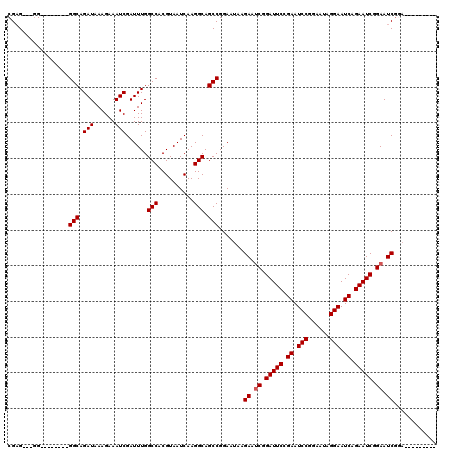
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:31:07 2006