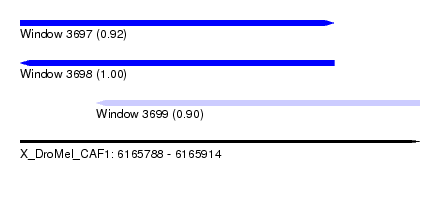

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 6,165,788 – 6,165,914 |
| Length | 126 |
| Max. P | 0.999674 |

| Location | 6,165,788 – 6,165,887 |
|---|---|
| Length | 99 |
| Sequences | 5 |
| Columns | 100 |
| Reading direction | forward |
| Mean pairwise identity | 86.70 |
| Mean single sequence MFE | -26.54 |
| Consensus MFE | -22.74 |
| Energy contribution | -23.14 |
| Covariance contribution | 0.40 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.85 |
| Structure conservation index | 0.86 |
| SVM decision value | 1.12 |
| SVM RNA-class probability | 0.917724 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
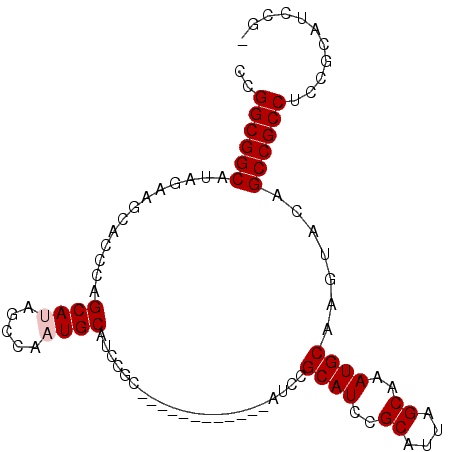
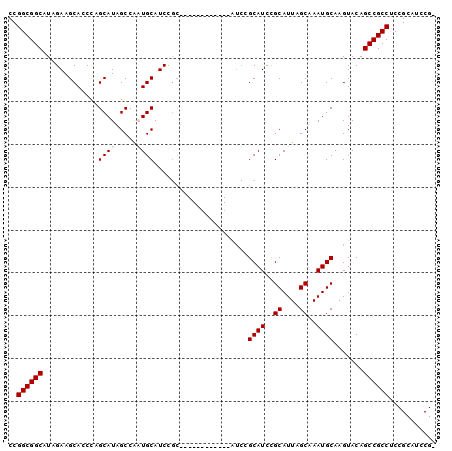
>X_DroMel_CAF1 6165788 99 + 22224390 CCGGCGGCAUAGAAGCACCCAGCAUAGCCAAUGCAUCCGCAACCUCAACUGCAUCCGCAUCCGCAUUAGCAAAUGCAAGUACAGCCGCCUCCGCAUCCG- ..((((((...((.((.....((((.....))))....(((........)))....)).)).(((((....))))).......))))))..........- ( -27.90) >DroSec_CAF1 4399 87 + 1 CCGGCGGCAUAGAAGCACCCAGCAUAGCCAAUGCAUCCGC------------AUCUGCAUCCGCAUUAGCAAAUGCAAGUACAGCCGCCUCCGCAUCCG- ..((((((..(((.((.....((((.....))))....))------------.)))((((..((....))..)))).......))))))..........- ( -26.10) >DroSim_CAF1 6715 87 + 1 CCGGCGGCAUAGAAGCACCCAGCAUAGCCAAUGCAUCCGC------------AUCUGCAUCCGCAUUAGCAAAUGCAAGUACAGCCGCCUCCGCAUCCG- ..((((((..(((.((.....((((.....))))....))------------.)))((((..((....))..)))).......))))))..........- ( -26.10) >DroEre_CAF1 4263 81 + 1 CCGGCGGCAUUGAAGCACCUAGCAUAGCCGCUGCAUCC------------------GCAUCCGCAUUGGCAAAUGCAAGUACAGCCGCCUCCGCAUCCG- ..((((((......((.....))...((((.(((....------------------......))).)))).............))))))..........- ( -24.60) >DroYak_CAF1 10055 94 + 1 CCGGCGGCAUUGAAGCACCCAGCAUAGCCACUGCAUCUGCAUCC------GCAUCCGCAUCUGCAUUAGCAAAUGCAAGUACAGCCGCCUCCGCAUCCGC ..((((((......((.....))...((...(((....)))...------))....((((.(((....))).)))).......))))))........... ( -28.00) >consensus CCGGCGGCAUAGAAGCACCCAGCAUAGCCAAUGCAUCCGC____________AUCCGCAUCCGCAUUAGCAAAUGCAAGUACAGCCGCCUCCGCAUCCG_ ..((((((.............((((.....))))......................((((..((....))..)))).......))))))........... (-22.74 = -23.14 + 0.40)

| Location | 6,165,788 – 6,165,887 |
|---|---|
| Length | 99 |
| Sequences | 5 |
| Columns | 100 |
| Reading direction | reverse |
| Mean pairwise identity | 86.70 |
| Mean single sequence MFE | -40.42 |
| Consensus MFE | -36.95 |
| Energy contribution | -37.31 |
| Covariance contribution | 0.36 |
| Combinations/Pair | 1.14 |
| Mean z-score | -2.96 |
| Structure conservation index | 0.91 |
| SVM decision value | 3.87 |
| SVM RNA-class probability | 0.999674 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
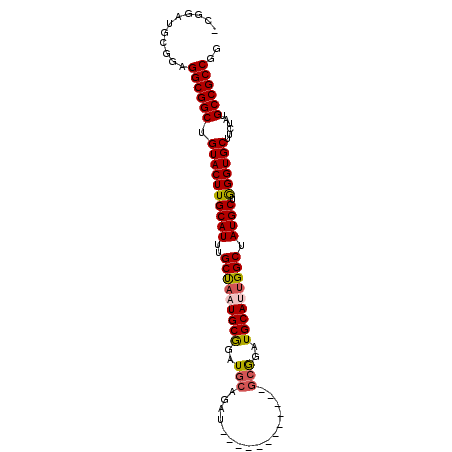
>X_DroMel_CAF1 6165788 99 - 22224390 -CGGAUGCGGAGGCGGCUGUACUUGCAUUUGCUAAUGCGGAUGCGGAUGCAGUUGAGGUUGCGGAUGCAUUGGCUAUGCUGGGUGCUUCUAUGCCGCCGG -..........((((((.((((((((((..(((((((((..(((((...(....)...)))))..))))))))).)))).))))))......)))))).. ( -42.40) >DroSec_CAF1 4399 87 - 1 -CGGAUGCGGAGGCGGCUGUACUUGCAUUUGCUAAUGCGGAUGCAGAU------------GCGGAUGCAUUGGCUAUGCUGGGUGCUUCUAUGCCGCCGG -..........((((((.((((((((((..(((((((((..(((....------------)))..))))))))).)))).))))))......)))))).. ( -41.00) >DroSim_CAF1 6715 87 - 1 -CGGAUGCGGAGGCGGCUGUACUUGCAUUUGCUAAUGCGGAUGCAGAU------------GCGGAUGCAUUGGCUAUGCUGGGUGCUUCUAUGCCGCCGG -..........((((((.((((((((((..(((((((((..(((....------------)))..))))))))).)))).))))))......)))))).. ( -41.00) >DroEre_CAF1 4263 81 - 1 -CGGAUGCGGAGGCGGCUGUACUUGCAUUUGCCAAUGCGGAUGC------------------GGAUGCAGCGGCUAUGCUAGGUGCUUCAAUGCCGCCGG -(((..(((..(((.((((((..(((((((((....))))))))------------------)..)))))).))).)))..(((........))).))). ( -36.70) >DroYak_CAF1 10055 94 - 1 GCGGAUGCGGAGGCGGCUGUACUUGCAUUUGCUAAUGCAGAUGCGGAUGC------GGAUGCAGAUGCAGUGGCUAUGCUGGGUGCUUCAAUGCCGCCGG .(((..(((..(((.((((((.((((((((((...(((....)))...))------)))))))).)))))).))).)))..(((........))).))). ( -41.00) >consensus _CGGAUGCGGAGGCGGCUGUACUUGCAUUUGCUAAUGCGGAUGCAGAU____________GCGGAUGCAUUGGCUAUGCUGGGUGCUUCUAUGCCGCCGG ...........((((((.((((((((((..(((((((((..(((................)))..))))))))).)))).))))))......)))))).. (-36.95 = -37.31 + 0.36)



| Location | 6,165,812 – 6,165,914 |
|---|---|
| Length | 102 |
| Sequences | 5 |
| Columns | 111 |
| Reading direction | reverse |
| Mean pairwise identity | 83.83 |
| Mean single sequence MFE | -35.26 |
| Consensus MFE | -25.30 |
| Energy contribution | -25.62 |
| Covariance contribution | 0.32 |
| Combinations/Pair | 1.12 |
| Mean z-score | -2.20 |
| Structure conservation index | 0.72 |
| SVM decision value | 0.99 |
| SVM RNA-class probability | 0.896385 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
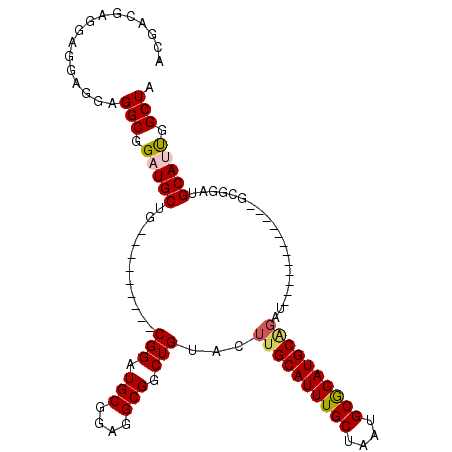
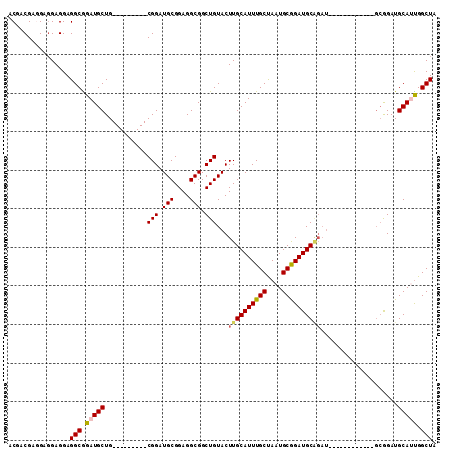
>X_DroMel_CAF1 6165812 102 - 22224390 ACGACGAUGAGGAGGAGGCGGAUGCUG---------CGGAUGCGGAGGCGGCUGUACUUGCAUUUGCUAAUGCGGAUGCGGAUGCAGUUGAGGUUGCGGAUGCAUUGGCUA .(..(.....)..)..(((.(((((..---------....(((((...((((((((..(((((((((....)))))))))..))))))))...)))))...))))).))). ( -40.40) >DroSec_CAF1 4423 90 - 1 ACGACGAGGAGGAGGAGGCGGAUGCUG---------CGGAUGCGGAGGCGGCUGUACUUGCAUUUGCUAAUGCGGAUGCAGAU------------GCGGAUGCAUUGGCUA .(..(.....)..)..(((.(((.(((---------(....))))..(((.(((((.((((((((((....)))))))))).)------------)))).)))))).))). ( -32.10) >DroSim_CAF1 6739 90 - 1 ACGACGAGGAGGAGGAGGCGGAUGCUG---------CGGAUGCGGAGGCGGCUGUACUUGCAUUUGCUAAUGCGGAUGCAGAU------------GCGGAUGCAUUGGCUA .(..(.....)..)..(((.(((.(((---------(....))))..(((.(((((.((((((((((....)))))))))).)------------)))).)))))).))). ( -32.10) >DroEre_CAF1 4287 84 - 1 ACGACGAGGAUGAGGAGGCGGAUGCUG---------CGGAUGCGGAGGCGGCUGUACUUGCAUUUGCCAAUGCGGAUGC------------------GGAUGCAGCGGCUA .(..((....((....(((....))).---------))....))..)((.((((((..(((((((((....))))))))------------------)..)))))).)).. ( -31.50) >DroYak_CAF1 10079 105 - 1 ACGACGACGAGGAGGAGGCAGAUGCUGCGGAUGUUGCGGAUGCGGAGGCGGCUGUACUUGCAUUUGCUAAUGCAGAUGCGGAUGC------GGAUGCAGAUGCAGUGGCUA .(..(.....)..)...(((..(((.((....)).)))..)))...(((.((((((.((((((((((...(((....)))...))------)))))))).)))))).))). ( -40.20) >consensus ACGACGAGGAGGAGGAGGCGGAUGCUG_________CGGAUGCGGAGGCGGCUGUACUUGCAUUUGCUAAUGCGGAUGCAGAU____________GCGGAUGCAUUGGCUA ................(((.(((((...........(((.(((....))).)))...((((((((((....))))))))))....................))))).))). (-25.30 = -25.62 + 0.32)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:30:34 2006