
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 6,160,715 – 6,160,891 |
| Length | 176 |
| Max. P | 0.724153 |

| Location | 6,160,715 – 6,160,833 |
|---|---|
| Length | 118 |
| Sequences | 3 |
| Columns | 118 |
| Reading direction | forward |
| Mean pairwise identity | 80.23 |
| Mean single sequence MFE | -23.62 |
| Consensus MFE | -19.82 |
| Energy contribution | -20.48 |
| Covariance contribution | 0.67 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.33 |
| Structure conservation index | 0.84 |
| SVM decision value | 0.41 |
| SVM RNA-class probability | 0.724153 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
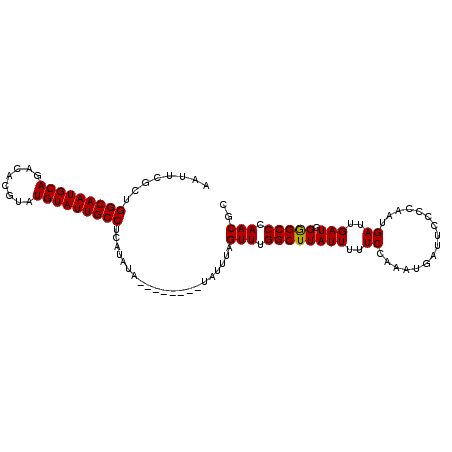
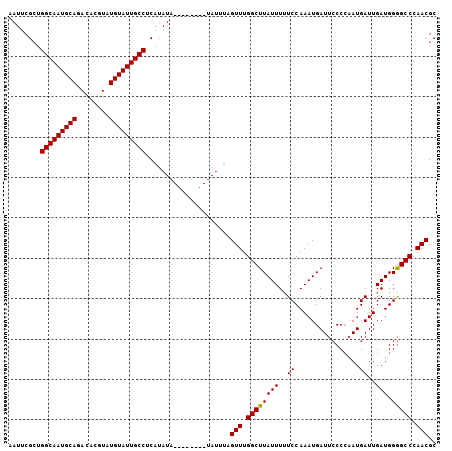
>X_DroMel_CAF1 6160715 118 + 22224390 UACCCCCUUUUCUCUGCUCUCUUCUCCUGCCAUCUCACUCUGUCGCCGCAUUUGUCUCGCUCGCUUGCAGUGGCGAAUAAUUCGAUGGCAAUGCAGACACGUAUGUAUUGCCUCAUAU ............(((((..........(((((((........(((((((....(...)((......)).))))))).......)))))))..)))))...(((((........))))) ( -28.06) >DroSim_CAF1 1695 118 + 1 UACCCCCUUUUCUCUGCUCUCUUCUCCUGCCGUCUCCCUCUGUCGCCGCAUCCGUCUCGCUCGCUCCCAGUGGCGAAUAAUUCGCUGGCAAUGCAGACACGUAUGUAUUGCCUCAUAU ...............((..........(((((.(.......).))..)))......((((.(((.....))))))).......)).(((((((((........)))))))))...... ( -24.80) >DroYak_CAF1 5155 99 + 1 UGUCCCCUUU-CUCU------------------CCCUUUCUCUCACCGCAUCCGUCUCGCUCGCUCGCAGUGGCAAAUUAUUCGCUGGCAAUGCAGACACGUAUGUAUUGCCUCAAAU ..........-....------------------.........((((.((...((.......))...)).)))).............(((((((((........)))))))))...... ( -18.00) >consensus UACCCCCUUUUCUCUGCUCUCUUCUCCUGCC_UCUCACUCUGUCGCCGCAUCCGUCUCGCUCGCUCGCAGUGGCGAAUAAUUCGCUGGCAAUGCAGACACGUAUGUAUUGCCUCAUAU ..........................................(((((((....................)))))))..........(((((((((........)))))))))...... (-19.82 = -20.48 + 0.67)



| Location | 6,160,793 – 6,160,891 |
|---|---|
| Length | 98 |
| Sequences | 3 |
| Columns | 106 |
| Reading direction | forward |
| Mean pairwise identity | 90.32 |
| Mean single sequence MFE | -25.53 |
| Consensus MFE | -21.51 |
| Energy contribution | -21.29 |
| Covariance contribution | -0.22 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.82 |
| Structure conservation index | 0.84 |
| SVM decision value | 0.02 |
| SVM RNA-class probability | 0.542810 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 6160793 98 + 22224390 AAUUCGAUGGCAAUGCAGACACGUAUGUAUUGCCUCAUAUA--------UAUUUAGUUUGGCUUAUUUUUCCAAAUGAUUCCCGAAUGAUUGAUGGGGCCCAACGC .....((.(((((((((........))))))))))).....--------......(((.(((((((((....))))))..((((.........))))))).))).. ( -24.50) >DroSim_CAF1 1773 98 + 1 AAUUCGCUGGCAAUGCAGACACGUAUGUAUUGCCUCAUAUA--------UAUUUAGUUUGGCUUAUUUUUCCAAAUGAUUCCCCAAUGAUUGAUGGAGCCCAACGC .....((.(((((((((........))))))))).......--------......(((.((((((((..((....((......))..))..))).))))).))))) ( -23.40) >DroYak_CAF1 5214 106 + 1 UAUUCGCUGGCAAUGCAGACACGUAUGUAUUGCCUCAAAUAUAUGUAUGUAUUUAGUUUGGCUUAUUUCUCCAAAUGAUUCCCCCAUGAUUGAUGGGGCCCAACGC .....((.(((((((((........)))))))))..(((((((....))))))).(((.(((((((((....))))))....(((((.....)))))))).))))) ( -28.70) >consensus AAUUCGCUGGCAAUGCAGACACGUAUGUAUUGCCUCAUAUA________UAUUUAGUUUGGCUUAUUUUUCCAAAUGAUUCCCCAAUGAUUGAUGGGGCCCAACGC ........(((((((((........))))))))).....................(((.((((((((..((................))..))).))))).))).. (-21.51 = -21.29 + -0.22)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:30:25 2006