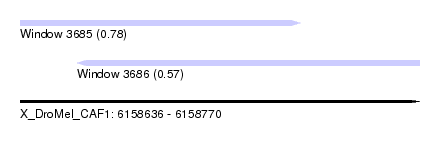

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 6,158,636 – 6,158,770 |
| Length | 134 |
| Max. P | 0.780929 |

| Location | 6,158,636 – 6,158,730 |
|---|---|
| Length | 94 |
| Sequences | 3 |
| Columns | 115 |
| Reading direction | forward |
| Mean pairwise identity | 77.78 |
| Mean single sequence MFE | -18.97 |
| Consensus MFE | -16.24 |
| Energy contribution | -15.48 |
| Covariance contribution | -0.77 |
| Combinations/Pair | 1.21 |
| Mean z-score | -1.07 |
| Structure conservation index | 0.86 |
| SVM decision value | 0.56 |
| SVM RNA-class probability | 0.780929 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 6158636 94 + 22224390 ---------------------AAAGUAAAGUGCAGUAAAUGCAACAAAGUGUUUUGCAUAUGUGCAGCUAAUUGGAACAAAUUCAAUUUAUGUUUAGCUAAAUUAGGCAAAACUU ---------------------.........((((.....)))).......(((((((....((..(((((((((((.....)))))).......)))))..))...))))))).. ( -18.21) >DroSec_CAF1 11436 94 + 1 ---------------------AAAGUAAAGCGCAGUAAAUGCAACAUAGUGUUUUGCAUAUGUGCGGCUAAUUGGAACAAAUUUAAUUUAUGUUUAGUUAAAUUAUGCGAAACUU ---------------------..........(((.....)))........(((((((((.(((....((....)).)))(((((((((.......)))))))))))))))))).. ( -16.80) >DroEre_CAF1 9118 114 + 1 AGAAGACUUUCGGUGAAAAGUAAAGUUAAGUGCAGUCAAUGUAACAAAGUGUUUUGCAUAUGUGCAGCUAAUUGAAACAAAUUCAAUUU-UGUGUAACUUAAUUAUGCGAAACUU ....(((((((........).))))))...((((.....)))).......((((((((((..((((...(((((((.....))))))).-..)))).......)))))))))).. ( -21.90) >consensus _____________________AAAGUAAAGUGCAGUAAAUGCAACAAAGUGUUUUGCAUAUGUGCAGCUAAUUGGAACAAAUUCAAUUUAUGUUUAGCUAAAUUAUGCGAAACUU ...............................(((.....)))........((((((((((((..((...(((((((.....)))))))..))..)).......)))))))))).. (-16.24 = -15.48 + -0.77)

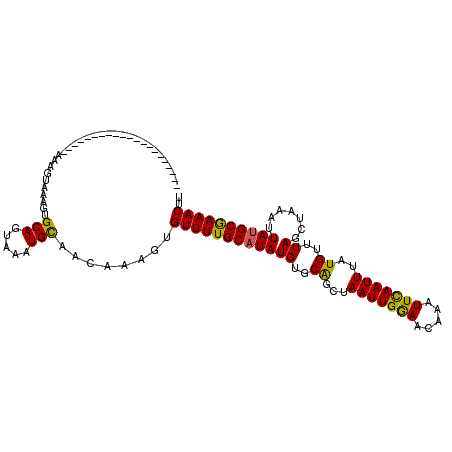
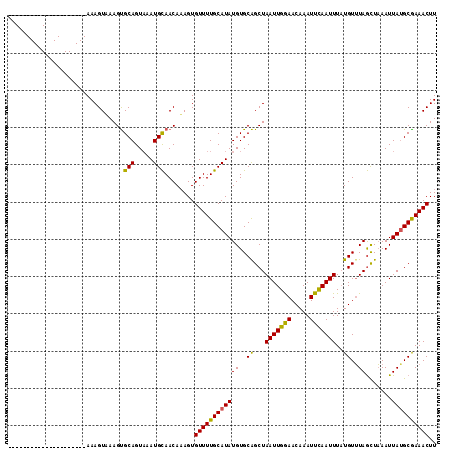
| Location | 6,158,655 – 6,158,770 |
|---|---|
| Length | 115 |
| Sequences | 3 |
| Columns | 115 |
| Reading direction | reverse |
| Mean pairwise identity | 86.67 |
| Mean single sequence MFE | -25.74 |
| Consensus MFE | -16.79 |
| Energy contribution | -17.47 |
| Covariance contribution | 0.68 |
| Combinations/Pair | 1.19 |
| Mean z-score | -2.41 |
| Structure conservation index | 0.65 |
| SVM decision value | 0.08 |
| SVM RNA-class probability | 0.573948 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 6158655 115 - 22224390 UUUCGUGCGUGUUGCGGCUAAAGUUUCUGUUUACGUUACAAAGUUUUGCCUAAUUUAGCUAAACAUAAAUUGAAUUUGUUCCAAUUAGCUGCACAUAUGCAAAACACUUUGUUGC .....(((((((((((((((((((((.((((((.((((...((((......)))))))))))))).)))))(((....)))...))))))))).))))))).............. ( -26.90) >DroSec_CAF1 11455 115 - 1 UUUCGUGCAUGUUGCGGCUUUAUUUUAUGUUUACGUUACAAAGUUUCGCAUAAUUUAACUAAACAUAAAUUAAAUUUGUUCCAAUUAGCCGCACAUAUGCAAAACACUAUGUUGC .....(((((((((((((((((.((((((((((.((((...((((......)))))))))))))))))).))).............))))))).))))))).............. ( -28.61) >DroEre_CAF1 9158 114 - 1 UUUUGUGUAUGUUGUGCUCAAAUUUUCUGUUUACGUUACAAAGUUUCGCAUAAUUAAGUUACACA-AAAUUGAAUUUGUUUCAAUUAGCUGCACAUAUGCAAAACACUUUGUUAC ..(((((((.(((((((..(((((((.((.......)).))))))).))))))).....))))))-)(((((((.....)))))))...((((....)))).............. ( -21.70) >consensus UUUCGUGCAUGUUGCGGCUAAAUUUUCUGUUUACGUUACAAAGUUUCGCAUAAUUUAGCUAAACAUAAAUUGAAUUUGUUCCAAUUAGCUGCACAUAUGCAAAACACUUUGUUGC .....((((((((((((((((.......(....)(..((((((.....).(((((((........)))))))..)))))..)..))))))))).))))))).............. (-16.79 = -17.47 + 0.68)

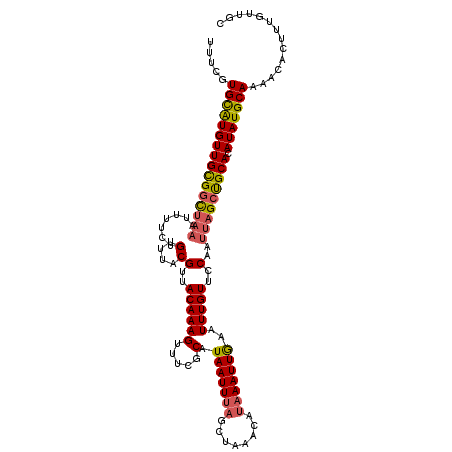

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:30:23 2006