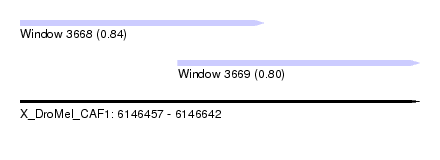
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 6,146,457 – 6,146,642 |
| Length | 185 |
| Max. P | 0.844625 |

| Location | 6,146,457 – 6,146,570 |
|---|---|
| Length | 113 |
| Sequences | 4 |
| Columns | 113 |
| Reading direction | forward |
| Mean pairwise identity | 89.40 |
| Mean single sequence MFE | -25.35 |
| Consensus MFE | -24.18 |
| Energy contribution | -24.25 |
| Covariance contribution | 0.06 |
| Combinations/Pair | 1.15 |
| Mean z-score | -1.54 |
| Structure conservation index | 0.95 |
| SVM decision value | 0.76 |
| SVM RNA-class probability | 0.844625 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
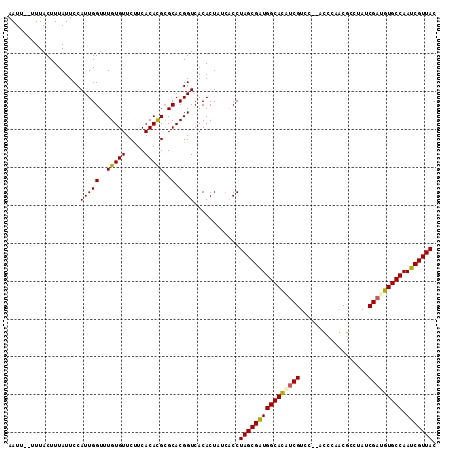
>X_DroMel_CAF1 6146457 113 + 22224390 AAUUUUUUUACUUUAUUCCAUUGGUUUGUGUUCUUCACACGGGCACGGUCACACUUUCACCUAGCGAUGGCAUUUCGUCCCAAUUCAACGCCUAUCGAUGUGCCAAUCGUUAC ...................((((((((((((.....)))))))).))))............((((((((((((.(((..................))).))))).))))))). ( -24.97) >DroSec_CAF1 1891 111 + 1 AAUUU-UUUACUUUAUUCCAUUGGUUUGUGUUCUUCACACGCGCACGGUCACACUUUCACCUAGCGAUGGCACCUCGUCCC-AUUCAACGCCUAUCGAUGUGCCAAUCGUUAC .....-.............((((((.(((((.....))))).)).))))............((((((((((((.(((....-.............))).))))).))))))). ( -23.73) >DroEre_CAF1 1875 109 + 1 AAUU--UUUACUUUAUUCCAUUGGUUUGUGUUCUUCACAUGCGCACGGUCACACUAUCCCCUAGCGGUGGCACGCCGUGC--ACCCAGCGCCUAUCGAUGUGCCAAUCGUUAC ....--.............((((((.((((.....)))).))(((((((..((((..........))))....)))))))--.....((((........))))))))...... ( -24.40) >DroYak_CAF1 1905 107 + 1 AAUU--UUUACUUUAUUCCAUUGGUUUGUGUUCCUCACACGCGCACGGUCACACUA--CCCUAGCGGUGGCACAUCGUCU--ACCCAGCGCCUAUCGAUGUGCCAAUCGUUAC ....--................(((.((((..((............)).))))..)--)).((((((((((((((((...--.............))))))))).))))))). ( -28.29) >consensus AAUU__UUUACUUUAUUCCAUUGGUUUGUGUUCUUCACACGCGCACGGUCACACUAUCACCUAGCGAUGGCACAUCGUCC__ACCCAACGCCUAUCGAUGUGCCAAUCGUUAC ...................(((((..(((((.....)))))..).))))............((((((((((((((((..................))))))))).))))))). (-24.18 = -24.25 + 0.06)


| Location | 6,146,530 – 6,146,642 |
|---|---|
| Length | 112 |
| Sequences | 4 |
| Columns | 116 |
| Reading direction | forward |
| Mean pairwise identity | 77.70 |
| Mean single sequence MFE | -29.52 |
| Consensus MFE | -19.97 |
| Energy contribution | -21.16 |
| Covariance contribution | 1.19 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.78 |
| Structure conservation index | 0.68 |
| SVM decision value | 0.61 |
| SVM RNA-class probability | 0.798737 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 6146530 112 + 22224390 UUCGUCCCAAUUCAACGCCUAUCGAUGUGCCAAUCGUUACGUGACGUU----GGCGCAGGUCGUACUUAACGUUACGUCACGUAACGGAAAAGCGUUAAUUCAAAUUGCACUUAUU .............(((((...(((((......)))((((((((((((.----((((.(((.....)))..)))))))))))))))).))...)))))................... ( -30.70) >DroSec_CAF1 1963 107 + 1 CUCGUCCC-AUUCAACGCCUAUCGAUGUGCCAAUCGUUACGUGACGUU----GGCGCAGGUCGUAUUUAACGUUACGUUACGUAACGGAAAAGCGUUAAUUCAAA----AUUUUUG ........-....(((((....(((((((((((.((((....))))))----))))))..)))..(((..(((((((...)))))))..))))))))........----....... ( -30.40) >DroEre_CAF1 1946 103 + 1 GCCGUGC--ACCCAGCGCCUAUCGAUGUGCCAAUCGUUACGUGACGUUGUAUGACGCUGGUCAUAUUUAA----------CGUAACGGAAAAGCGUUACUUCAAAUUA-AAUUAAU ...((((--.....)))).....(((((.....(((((((((......(((((((....)))))))...)----------))))))))....)))))...........-....... ( -26.30) >DroYak_CAF1 1974 102 + 1 AUCGUCU--ACCCAGCGCCUAUCGAUGUGCCAAUCGUUACGUGACGUU----GACGCAGGUCUUAUUUAACGUUACGUAACGUAACGGAUAAGCGUUAUUUUAAAUGG-------- .......--....(((((.(((((((......)))(((((((((((((----((............)))))))))))))))......)))).)))))...........-------- ( -30.70) >consensus AUCGUCC__ACCCAACGCCUAUCGAUGUGCCAAUCGUUACGUGACGUU____GACGCAGGUCGUAUUUAACGUUACGU_ACGUAACGGAAAAGCGUUAAUUCAAAUUG_A_UUAUU .............(((((...(((((......)))((((((((((((.....(((....)))............)))))))))))).))...)))))................... (-19.97 = -21.16 + 1.19)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:30:08 2006