
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 5,899,973 – 5,900,104 |
| Length | 131 |
| Max. P | 0.800100 |

| Location | 5,899,973 – 5,900,079 |
|---|---|
| Length | 106 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 92.18 |
| Mean single sequence MFE | -30.26 |
| Consensus MFE | -25.36 |
| Energy contribution | -25.60 |
| Covariance contribution | 0.24 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.43 |
| Structure conservation index | 0.84 |
| SVM decision value | 0.45 |
| SVM RNA-class probability | 0.741705 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
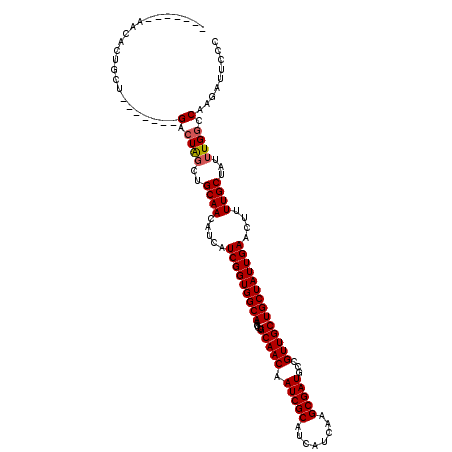
>X_DroMel_CAF1 5899973 106 - 22224390 -------AACACUGCU-------GACUAGCUGCAACAUCAUCGGUGGCAAGCUGCAACAAUCGCAUCAUCAAGCGAUGCCGUUGCUGCUAUUGAACUUUUGCUAUUUGGCCAAGAUUCCC -------......((.-------.(.((((..........(((((((((....(((((.(((((........)))))...))))))))))))))......)))).)..)).......... ( -29.49) >DroSec_CAF1 16655 106 - 1 -------AACACUGCU-------GACUAGCUGCAACAUCAUCGGUGGCAAGCUGCAACAAUCGCAUCAUCAAGCGAUGCCGUUGCUGCUAUUGAACUUUUGCUAUGUGGCCAAGAUUCCC -------..(((.(((-------....))).((((.....(((((((((....(((((.(((((........)))))...))))))))))))))....))))...)))............ ( -29.00) >DroSim_CAF1 14006 106 - 1 -------AACACUGCU-------GACUAGCUGCAACAUCAUCGGUGGCAAGCUGCAACAAUCGCAUCAUCAAGCGAUGCCGUUGCUGCUAUUGAACUUUUGCUAUUUGGCCAAGAUUCCC -------......((.-------.(.((((..........(((((((((....(((((.(((((........)))))...))))))))))))))......)))).)..)).......... ( -29.49) >DroEre_CAF1 16751 113 - 1 -------AACACCGCAGACGGACGACUGGCUGCAACAUCAUCGGUGGCAAGCUGCAACAAUCGCAUCAUCAAGCGAUGCCGUUGCUGCUAUUGAACUUUUGCUAUUUGCCCAAGAUUCCC -------......((((.(((....))).)))).......(((((((((....(((((.(((((........)))))...)))))))))))))).(((..((.....))..)))...... ( -32.80) >DroYak_CAF1 16905 116 - 1 CUACAAAAACACUGCA----GACGACUAGCUGCAACAUCAUCGGUGGCAAGCUGCAACAAUCGCAUCAUCAGGCGAUGCCGUUGCUGCUAUUGAACUUUUGCUAUUUGGCCAAGAUUCCC ............((((----(.(.....))))))......(((((((((....(((((.(((((........)))))...)))))))))))))).(((..(((....))).)))...... ( -30.50) >consensus _______AACACUGCU_______GACUAGCUGCAACAUCAUCGGUGGCAAGCUGCAACAAUCGCAUCAUCAAGCGAUGCCGUUGCUGCUAUUGAACUUUUGCUAUUUGGCCAAGAUUCCC .......................(.((((..((((.....(((((((((....(((((.(((((........)))))...))))))))))))))....))))...)))).)......... (-25.36 = -25.60 + 0.24)


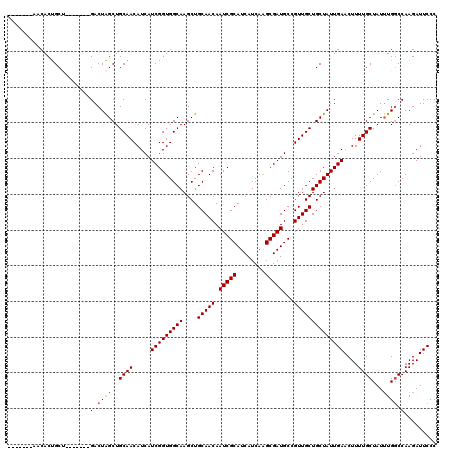
| Location | 5,900,013 – 5,900,104 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 84.85 |
| Mean single sequence MFE | -26.10 |
| Consensus MFE | -16.92 |
| Energy contribution | -16.60 |
| Covariance contribution | -0.32 |
| Combinations/Pair | 1.14 |
| Mean z-score | -2.32 |
| Structure conservation index | 0.65 |
| SVM decision value | 0.61 |
| SVM RNA-class probability | 0.800100 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 5900013 91 - 22224390 UAGC------AGCAAUACUAGAAACACUGGA-------------AACACUGCU-------GACUAGCUGCAACAUCAUCGGUGGCAAGCUGCAACAAUCGCAUCAUCAAGCGAUGCC ..((------(((....((((.....)))).-------------..(((((.(-------((............))).)))))....)))))....(((((........)))))... ( -24.60) >DroSec_CAF1 16695 91 - 1 UAGC------AGCAACACUAGAAACACUGGA-------------AACACUGCU-------GACUAGCUGCAACAUCAUCGGUGGCAAGCUGCAACAAUCGCAUCAUCAAGCGAUGCC ..((------(((....((((.....)))).-------------..(((((.(-------((............))).)))))....)))))....(((((........)))))... ( -24.60) >DroSim_CAF1 14046 91 - 1 UAGC------AGCAACACUAGAAACACUGGA-------------AACACUGCU-------GACUAGCUGCAACAUCAUCGGUGGCAAGCUGCAACAAUCGCAUCAUCAAGCGAUGCC ..((------(((....((((.....)))).-------------..(((((.(-------((............))).)))))....)))))....(((((........)))))... ( -24.60) >DroEre_CAF1 16791 98 - 1 CAGC------AGCAACACUGGAAACUCUAGA-------------AACACCGCAGACGGACGACUGGCUGCAACAUCAUCGGUGGCAAGCUGCAACAAUCGCAUCAUCAAGCGAUGCC ..((------(((....((((.....)))).-------------..((((((((.(((....))).)))).........))))....)))))....(((((........)))))... ( -30.00) >DroYak_CAF1 16945 113 - 1 CAGCAGCAGCAGCAACACUUGAAACACUAGACACACACUACAAAAACACUGCA----GACGACUAGCUGCAACAUCAUCGGUGGCAAGCUGCAACAAUCGCAUCAUCAGGCGAUGCC ..((.(((((.((.((...(((.....(((.......))).........((((----(.(.....))))))...)))...)).))..)))))....(((((........))))))). ( -26.70) >consensus UAGC______AGCAACACUAGAAACACUGGA_____________AACACUGCU_______GACUAGCUGCAACAUCAUCGGUGGCAAGCUGCAACAAUCGCAUCAUCAAGCGAUGCC ..((.......((....((((.....))))..................................((((((..(((.....))))).))))))....(((((........))))))). (-16.92 = -16.60 + -0.32)

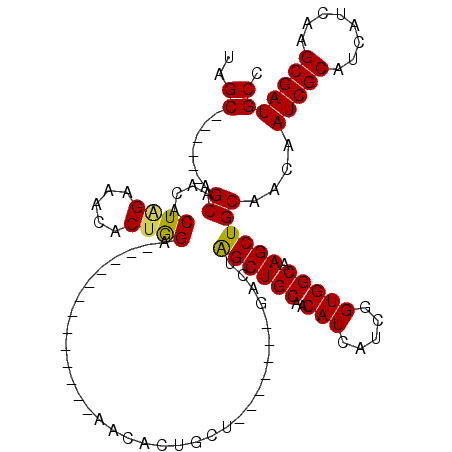

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:28:00 2006