
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 5,897,916 – 5,898,046 |
| Length | 130 |
| Max. P | 0.938579 |

| Location | 5,897,916 – 5,898,006 |
|---|---|
| Length | 90 |
| Sequences | 4 |
| Columns | 90 |
| Reading direction | reverse |
| Mean pairwise identity | 97.78 |
| Mean single sequence MFE | -27.33 |
| Consensus MFE | -26.00 |
| Energy contribution | -26.25 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.10 |
| Structure conservation index | 0.95 |
| SVM decision value | 0.89 |
| SVM RNA-class probability | 0.875520 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 5897916 90 - 22224390 UUGUGUAUCUAUGAGAUACGCGCAACGUGAAUUGGCAUUUUCCAUUUGGCAAUUUGCAUGUGCAAUGCAAUGCGGGCUCCAAGAUCUGCA (((((((((.....)))))(((((..(..(((((.((.........)).)))))..).)))))...))))(((((((.....).)))))) ( -27.50) >DroSec_CAF1 14659 90 - 1 UUGUGUAUCUAUGAGAUACGCGCAACGUGAAUUGGCAUUUUCCAUUUGGCAAUUUGCAUGUGCAAUGUAAUGCGGGCUCCAAGAUCUGCA ..(((((((.....)))))))(((..(..(((((.((.........)).)))))..)...))).......(((((((.....).)))))) ( -27.20) >DroEre_CAF1 14755 90 - 1 UUGUGUAUCUAUGAGAUACGCGCAACGUGAAUUGGCAUUUUCCAUUUGGCAAUUUGCAUGUGCAAUGCAAUGCGGGCUCCAAGAUCUGCA (((((((((.....)))))(((((..(..(((((.((.........)).)))))..).)))))...))))(((((((.....).)))))) ( -27.50) >DroYak_CAF1 14860 90 - 1 UUGUGUAUCUAUGAGAUACGCGCAACGUGAAUUGGCAUUUUCCAUUUGGCAAUUUGCAUGUGCAAUGCAAUGCGGGCAGCAAGAUAUGCA .(((((((((.........(((((..(..(((((.((.........)).)))))..).))))).......(((.....)))))))))))) ( -27.10) >consensus UUGUGUAUCUAUGAGAUACGCGCAACGUGAAUUGGCAUUUUCCAUUUGGCAAUUUGCAUGUGCAAUGCAAUGCGGGCUCCAAGAUCUGCA ..(((((((.....)))))))(((..(..(((((.((.........)).)))))..)...))).......(((((((.....).)))))) (-26.00 = -26.25 + 0.25)


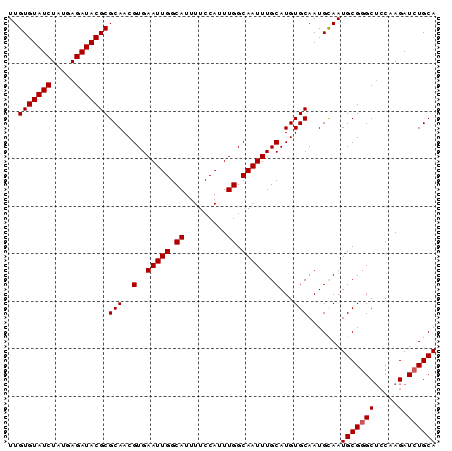
| Location | 5,897,948 – 5,898,046 |
|---|---|
| Length | 98 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 84.27 |
| Mean single sequence MFE | -22.44 |
| Consensus MFE | -14.14 |
| Energy contribution | -15.74 |
| Covariance contribution | 1.60 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.85 |
| Structure conservation index | 0.63 |
| SVM decision value | 1.27 |
| SVM RNA-class probability | 0.938579 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 5897948 98 + 22224390 UGCA---------------------AAUUGCCAAAUGGAAAAUGCCAAUUCACGUUGCGCGUAUCUCAUAGAUACACAA-AAUUGAAAUUAUAAUAAAAAGCAAUGAGGCAAAAAACGGU ....---------------------..(((((...(((......))).....((((((..(((((.....))))).((.-...))...............)))))).)))))........ ( -21.10) >DroSec_CAF1 14691 98 + 1 UGCA---------------------AAUUGCCAAAUGGAAAAUGCCAAUUCACGUUGCGCGUAUCUCAUAGAUACACAA-AAUUGAAAUUAUAAUAAAAAGCAAUGAGGCAAAAAACGGU ....---------------------..(((((...(((......))).....((((((..(((((.....))))).((.-...))...............)))))).)))))........ ( -21.10) >DroEre_CAF1 14787 97 + 1 UGCA---------------------AAUUGCCAAAUGGAAAAUGCCAAUUCACGUUGCGCGUAUCUCAUAGAUACACAA-AAUUGAAAUUAUAAUAAAAAGCAAUGAGGCAA-AAACGGU ....---------------------..(((((...(((......))).....((((((..(((((.....))))).((.-...))...............)))))).)))))-....... ( -21.10) >DroWil_CAF1 14757 118 + 1 UGCAAAUUGCAAAUUACGAUUGGCGAAUUGCAAAAUGCAAAAUGCCAAUUCACCUAGCGCGUAUCUCAUAGAUACACAAAAAUUGAAAUUAUAAGACUGCCCCCUGAGGCUCAUAG-GC- (((.....)))......((((((((..((((.....))))..))))))))..((((..(.(((((.....))))).).....................(((......)))...)))-).- ( -27.80) >DroYak_CAF1 14892 98 + 1 UGCA---------------------AAUUGCCAAAUGGAAAAUGCCAAUUCACGUUGCGCGUAUCUCAUAGAUACACAA-AAUUGAAAUUAUAAUAAAAAGCAAUGAGGCAAAAAACGGU ....---------------------..(((((...(((......))).....((((((..(((((.....))))).((.-...))...............)))))).)))))........ ( -21.10) >consensus UGCA_____________________AAUUGCCAAAUGGAAAAUGCCAAUUCACGUUGCGCGUAUCUCAUAGAUACACAA_AAUUGAAAUUAUAAUAAAAAGCAAUGAGGCAAAAAACGGU ...........................(((((...(((......))).....((((((..(((((.....))))).((.....))...............)))))).)))))........ (-14.14 = -15.74 + 1.60)



| Location | 5,897,948 – 5,898,046 |
|---|---|
| Length | 98 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 84.27 |
| Mean single sequence MFE | -23.32 |
| Consensus MFE | -15.34 |
| Energy contribution | -16.14 |
| Covariance contribution | 0.80 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.09 |
| Structure conservation index | 0.66 |
| SVM decision value | 0.47 |
| SVM RNA-class probability | 0.748424 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 5897948 98 - 22224390 ACCGUUUUUUGCCUCAUUGCUUUUUAUUAUAAUUUCAAUU-UUGUGUAUCUAUGAGAUACGCGCAACGUGAAUUGGCAUUUUCCAUUUGGCAAUU---------------------UGCA ...((...(((((....................((((...-.((((((((.....)))))))).....)))).(((......)))...)))))..---------------------.)). ( -20.50) >DroSec_CAF1 14691 98 - 1 ACCGUUUUUUGCCUCAUUGCUUUUUAUUAUAAUUUCAAUU-UUGUGUAUCUAUGAGAUACGCGCAACGUGAAUUGGCAUUUUCCAUUUGGCAAUU---------------------UGCA ...((...(((((....................((((...-.((((((((.....)))))))).....)))).(((......)))...)))))..---------------------.)). ( -20.50) >DroEre_CAF1 14787 97 - 1 ACCGUUU-UUGCCUCAUUGCUUUUUAUUAUAAUUUCAAUU-UUGUGUAUCUAUGAGAUACGCGCAACGUGAAUUGGCAUUUUCCAUUUGGCAAUU---------------------UGCA ...((..-(((((....................((((...-.((((((((.....)))))))).....)))).(((......)))...)))))..---------------------.)). ( -20.20) >DroWil_CAF1 14757 118 - 1 -GC-CUAUGAGCCUCAGGGGGCAGUCUUAUAAUUUCAAUUUUUGUGUAUCUAUGAGAUACGCGCUAGGUGAAUUGGCAUUUUGCAUUUUGCAAUUCGCCAAUCGUAAUUUGCAAUUUGCA -((-(.....((((....))))...........((((.((..((((((((.....))))))))..)).))))..)))....((((..((((((.(..(.....)..).))))))..)))) ( -34.90) >DroYak_CAF1 14892 98 - 1 ACCGUUUUUUGCCUCAUUGCUUUUUAUUAUAAUUUCAAUU-UUGUGUAUCUAUGAGAUACGCGCAACGUGAAUUGGCAUUUUCCAUUUGGCAAUU---------------------UGCA ...((...(((((....................((((...-.((((((((.....)))))))).....)))).(((......)))...)))))..---------------------.)). ( -20.50) >consensus ACCGUUUUUUGCCUCAUUGCUUUUUAUUAUAAUUUCAAUU_UUGUGUAUCUAUGAGAUACGCGCAACGUGAAUUGGCAUUUUCCAUUUGGCAAUU_____________________UGCA ........(((((....................((((.....((((((((.....)))))))).....)))).(((......)))...)))))........................... (-15.34 = -16.14 + 0.80)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:27:54 2006