
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 5,779,192 – 5,779,325 |
| Length | 133 |
| Max. P | 0.962818 |

| Location | 5,779,192 – 5,779,300 |
|---|---|
| Length | 108 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 87.68 |
| Mean single sequence MFE | -22.99 |
| Consensus MFE | -18.07 |
| Energy contribution | -18.82 |
| Covariance contribution | 0.75 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.52 |
| Structure conservation index | 0.79 |
| SVM decision value | 0.18 |
| SVM RNA-class probability | 0.620365 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 5779192 108 - 22224390 AUAAUCCCGCGACAUAAAACGAUUUCGGAAUUAACUCAAACGAAAUUGGUUUUCAUACGCGACACGAAAUGGAGCGGAAAAAGGAGUGGGGCUGAUAUAAAA------------AGUGUA .((..(((((..(......((((((((.............)))))))).(((((...(((..((.....))..)))))))).)..)))))..))........------------...... ( -26.22) >DroSec_CAF1 46758 108 - 1 AUAAUCCCGCGACAUAAACCGAUUUCGGAAUUAACUCAAACGAAAUUGGUUUUCAUACGCGACACGAAAUGAAACGGAAAAAGGAGUGGGUCUGAUAUAAAA------------AGUAUA .....(((((..(...(((((((((((.............)))))))))))(((((...((...))..))))).........)..)))))............------------...... ( -24.32) >DroEre_CAF1 47783 106 - 1 AUAAUCCCGCGGCAUAAACCGAUUUCGGAAUUAACUCAAACGAAAUUGGUUUUCAUACGCGACACGAAACGGAACGGAAAAAGGAGUGGCGCUGAUA--AAA------------AAUAUA .......((((....((((((((((((.............)))))))))))).....)))).((((..((....(.......)..))..)).))...--...------------...... ( -23.62) >DroYak_CAF1 51075 118 - 1 AUAAUCCCGCGGCAUAAACCGAUUUCGGAAUUAACUCAAACGAAAUUGCUUUUCAUACUCGACACGAAAUGGAACGGAAAAAGGAGUGGUGCUGAUA--AUAAAAAAGAAAGAGAAUAUA .........((((((...(((((((((.............))))))).(.((((((..(((...))).)))))).)......))....))))))...--..................... ( -17.82) >consensus AUAAUCCCGCGACAUAAACCGAUUUCGGAAUUAACUCAAACGAAAUUGGUUUUCAUACGCGACACGAAAUGGAACGGAAAAAGGAGUGGGGCUGAUA__AAA____________AAUAUA ....(((((((....((((((((((((.............)))))))))))).....))))...((........))......)))................................... (-18.07 = -18.82 + 0.75)

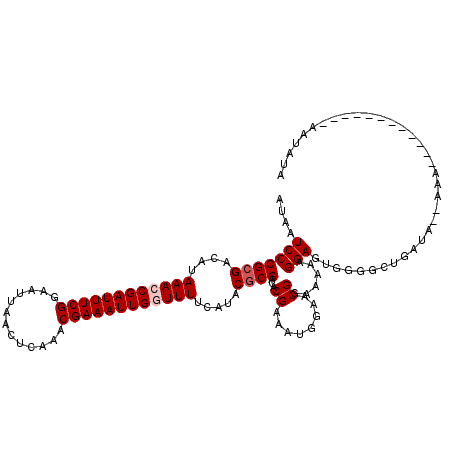

| Location | 5,779,220 – 5,779,325 |
|---|---|
| Length | 105 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 85.51 |
| Mean single sequence MFE | -26.02 |
| Consensus MFE | -23.39 |
| Energy contribution | -23.72 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.35 |
| Structure conservation index | 0.90 |
| SVM decision value | 1.54 |
| SVM RNA-class probability | 0.962818 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 5779220 105 - 22224390 GACCUCACCUCAGCG---------------ACAAAAAGGGAUAAUCCCGCGACAUAAAACGAUUUCGGAAUUAACUCAAACGAAAUUGGUUUUCAUACGCGACACGAAAUGGAGCGGAAA ..((.(.((....((---------------.......(((.....)))(((....((((((((((((.............))))))).)))))....)))....))....)).).))... ( -23.62) >DroSec_CAF1 46786 105 - 1 GACCUCACCGCGGCG---------------ACAAAAAGGGAUAAUCCCGCGACAUAAACCGAUUUCGGAAUUAACUCAAACGAAAUUGGUUUUCAUACGCGACACGAAAUGAAACGGAAA .......(((...((---------------.......(((.....)))(((....((((((((((((.............)))))))))))).....))).........))...)))... ( -27.02) >DroEre_CAF1 47809 120 - 1 GACCUCACCUUGGCAGAAAAAAAAAGAAAAAAAAAAAGGGAUAAUCCCGCGGCAUAAACCGAUUUCGGAAUUAACUCAAACGAAAUUGGUUUUCAUACGCGACACGAAACGGAACGGAAA ..((...((............................(((.....)))(((....((((((((((((.............)))))))))))).....)))..........))...))... ( -27.42) >consensus GACCUCACCUCGGCG_______________ACAAAAAGGGAUAAUCCCGCGACAUAAACCGAUUUCGGAAUUAACUCAAACGAAAUUGGUUUUCAUACGCGACACGAAAUGGAACGGAAA .......((............................(((.....)))(((....((((((((((((.............)))))))))))).....)))...............))... (-23.39 = -23.72 + 0.33)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:26:57 2006