
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 747,835 – 747,925 |
| Length | 90 |
| Max. P | 0.717126 |

| Location | 747,835 – 747,925 |
|---|---|
| Length | 90 |
| Sequences | 3 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 74.93 |
| Mean single sequence MFE | -31.41 |
| Consensus MFE | -17.42 |
| Energy contribution | -18.59 |
| Covariance contribution | 1.17 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.93 |
| Structure conservation index | 0.55 |
| SVM decision value | 0.39 |
| SVM RNA-class probability | 0.717126 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
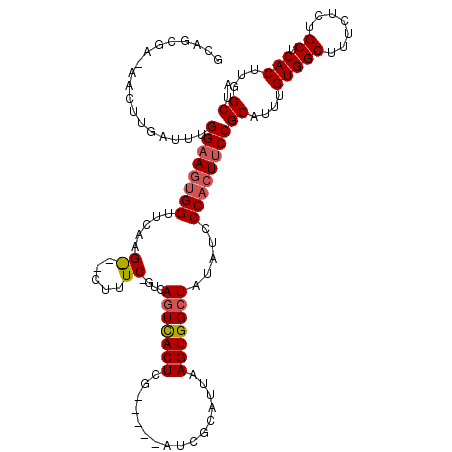
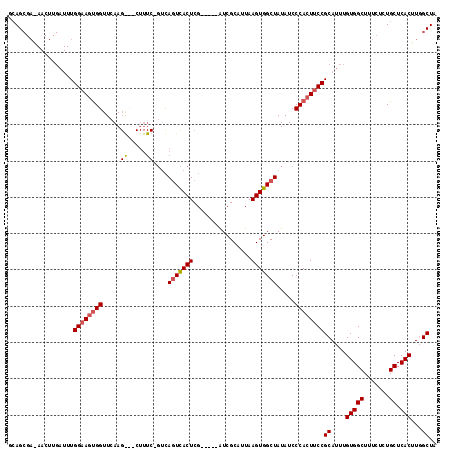
>X_DroMel_CAF1 747835 90 - 22224390 ACAG-------------UGGAAGUGGUUCAA----------GUCAGUCACUCG-----AUCGCAUUAAGUGGCUAUAUCCCACUUCCGCAUUUGUGGCUUUCUCUGCUCACUUGGCUA ...(-------------(((((((((.....----------...(((((((..-----((...))..))))))).....))))))))))....(((((.......)).)))....... ( -29.52) >DroSec_CAF1 62888 112 - 1 GCAGCGAGAACUUGAUUUGGAAGUGGUUCAAGGC-CUUUCAGUCAGUCACUCG-----AUCGCAUUAAGUGGCUAUAUCCCACUUCCGCAUUUGUGGCUUUCUCUGCUCACUUUGCUA ((((.((((.((.(((.(((((((((.....(((-......)))(((((((..-----((...))..))))))).....))))))))).)))...)).)))).))))........... ( -36.00) >DroYak_CAF1 65790 115 - 1 GAAGCGAAAACUUGAUUUGGAAGCGGUUCAAGAUCCUUUC---UACUUACUCCGUCGCGUCGCUUUAAGUGGCUAUAUCCCAGUGCCGCAUUUGUGGCUUUCUCUGCUCACUUGGCUA ((.((((...........(((((.((((...)))))))))---...........)))).))(((..(((((((...........(((((....))))).......)).)))))))).. ( -28.72) >consensus GCAGCGA_AACUUGAUUUGGAAGUGGUUCAAG___CUUUC_GUCAGUCACUCG_____AUCGCAUUAAGUGGCUAUAUCCCACUUCCGCAUUUGUGGCUUUCUCUGCUCACUUGGCUA ..................((((((((.....((.....))....(((((((................))))))).....))))))))((....(((((.......)).)))...)).. (-17.42 = -18.59 + 1.17)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:39:03 2006