
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 5,677,315 – 5,677,474 |
| Length | 159 |
| Max. P | 0.992286 |

| Location | 5,677,315 – 5,677,434 |
|---|---|
| Length | 119 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 85.75 |
| Mean single sequence MFE | -39.20 |
| Consensus MFE | -30.91 |
| Energy contribution | -30.03 |
| Covariance contribution | -0.88 |
| Combinations/Pair | 1.19 |
| Mean z-score | -2.06 |
| Structure conservation index | 0.79 |
| SVM decision value | 0.41 |
| SVM RNA-class probability | 0.726663 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 5677315 119 - 22224390 CAGCGCAUACAGUUAGUAUGCGAACUUCCAAU-CGGGAGGUAGUGCCGCCGGUCCGGGUUCGAAUCCCGGUGAAGUUAUCAAAAGUAGCAAUGGAAAAUAUAUGAUGAUGGUGCACUUCC ...(((((((.....))))))).((((((...-..))))))(((((.(((...(((((.......)))))....(((((((...(((...........))).)))))))))))))))... ( -38.20) >DroSec_CAF1 16218 118 - 1 CAGCGCAUACAGCUAGUAUGCGAACUCCCGAU-UCGGGUGUGGUGCCGCCGGUCCGGGUUCGAAUCCCGGUGAAGUUAAUAAAAGUAGCAACGGAAAAUGU-UACUAAUGUUAAACUUUA ...(((((((.....))))))).(((...(((-(((..(.(((.(((...)))))).)..))))))..)))((((((((((..(((((((........)))-))))..)))).)))))). ( -38.00) >DroSim_CAF1 17224 118 - 1 CAGCGCAUACAGCUAGUAUGCGAACUCCCGAUCUCGGGUGUGGUGCCGCCGGUCCGGGUUCGAAUCCCGGUGAAGUUAAUAAAAGUAGCAACGGAAAUUGU-UACAAACGUUA-ACUUUA ...(((((((.....))))))).((.((((....)))).))((.....))...(((((.......)))))((((((((((....(((((((......))))-)))....))))-)))))) ( -41.40) >consensus CAGCGCAUACAGCUAGUAUGCGAACUCCCGAU_UCGGGUGUGGUGCCGCCGGUCCGGGUUCGAAUCCCGGUGAAGUUAAUAAAAGUAGCAACGGAAAAUGU_UACUAAUGUUA_ACUUUA ..((((((((.....))))))..((.((((....)))).))......))....(((((.......))))).((((((((((..((((...(((.....))).))))..))))).))))). (-30.91 = -30.03 + -0.88)


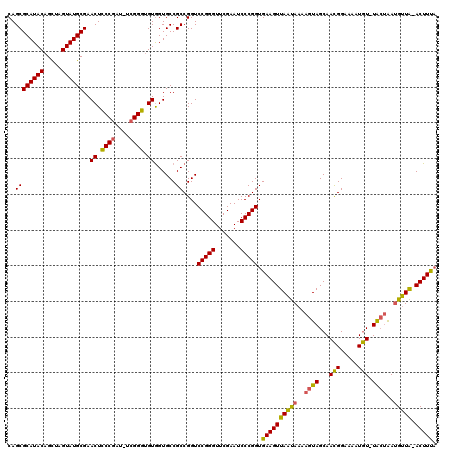
| Location | 5,677,355 – 5,677,474 |
|---|---|
| Length | 119 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 89.42 |
| Mean single sequence MFE | -54.77 |
| Consensus MFE | -51.48 |
| Energy contribution | -51.93 |
| Covariance contribution | 0.45 |
| Combinations/Pair | 1.09 |
| Mean z-score | -3.48 |
| Structure conservation index | 0.94 |
| SVM decision value | 2.32 |
| SVM RNA-class probability | 0.992286 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 5677355 119 + 22224390 GAUAACUUCACCGGGAUUCGAACCCGGACCGGCGGCACUACCUCCCG-AUUGGAAGUUCGCAUACUAACUGUAUGCGCUGCCGCCGGCGGCGAGGGGUCAGCUCCCAGGGGAUAAACUAG (((..((((.(((((.......))))).(((((((((((.((.....-...)).))).(((((((.....)))))))..))))))))....)))).)))...(((....)))........ ( -49.50) >DroSec_CAF1 16257 119 + 1 AUUAACUUCACCGGGAUUCGAACCCGGACCGGCGGCACCACACCCGA-AUCGGGAGUUCGCAUACUAGCUGUAUGCGCUGCCGCCGGCGGCGAGGGGACUCGCUCCAAAGGAUAAACUAG ..........(((((.......))))).(((((((((..((.((((.-..)))).)).(((((((.....))))))).)))))))((.((((((....))))))))...))......... ( -56.30) >DroSim_CAF1 17262 120 + 1 AUUAACUUCACCGGGAUUCGAACCCGGACCGGCGGCACCACACCCGAGAUCGGGAGUUCGCAUACUAGCUGUAUGCGCUGCCGCCGGCGGCGAGGGGACUCGCUCCAAAGGAUAAACUAG ..........(((((.......))))).(((((((((..((.((((....)))).)).(((((((.....))))))).)))))))((.((((((....))))))))...))......... ( -58.50) >consensus AUUAACUUCACCGGGAUUCGAACCCGGACCGGCGGCACCACACCCGA_AUCGGGAGUUCGCAUACUAGCUGUAUGCGCUGCCGCCGGCGGCGAGGGGACUCGCUCCAAAGGAUAAACUAG ..........(((((.......))))).(((((((((..((.((((....)))).)).(((((((.....))))))).)))))))((.((((((....))))))))...))......... (-51.48 = -51.93 + 0.45)

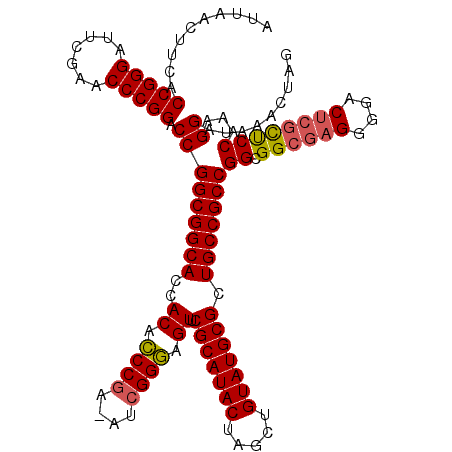

| Location | 5,677,355 – 5,677,474 |
|---|---|
| Length | 119 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 89.42 |
| Mean single sequence MFE | -55.63 |
| Consensus MFE | -50.60 |
| Energy contribution | -50.27 |
| Covariance contribution | -0.33 |
| Combinations/Pair | 1.06 |
| Mean z-score | -3.03 |
| Structure conservation index | 0.91 |
| SVM decision value | 1.90 |
| SVM RNA-class probability | 0.982044 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 5677355 119 - 22224390 CUAGUUUAUCCCCUGGGAGCUGACCCCUCGCCGCCGGCGGCAGCGCAUACAGUUAGUAUGCGAACUUCCAAU-CGGGAGGUAGUGCCGCCGGUCCGGGUUCGAAUCCCGGUGAAGUUAUC ...(((..(((....)))...)))...(((((((((((((((.(((((((.....))))))).((((((...-..))))))..))))))))))..(((.......))))))))....... ( -55.70) >DroSec_CAF1 16257 119 - 1 CUAGUUUAUCCUUUGGAGCGAGUCCCCUCGCCGCCGGCGGCAGCGCAUACAGCUAGUAUGCGAACUCCCGAU-UCGGGUGUGGUGCCGCCGGUCCGGGUUCGAAUCCCGGUGAAGUUAAU ..........((((.(.(((((....))))))((((((((((.(((((((.....))))))..((.((((..-.)))).))).))))))))))(((((.......))))).))))..... ( -54.50) >DroSim_CAF1 17262 120 - 1 CUAGUUUAUCCUUUGGAGCGAGUCCCCUCGCCGCCGGCGGCAGCGCAUACAGCUAGUAUGCGAACUCCCGAUCUCGGGUGUGGUGCCGCCGGUCCGGGUUCGAAUCCCGGUGAAGUUAAU ..........((((.(.(((((....))))))((((((((((.(((((((.....))))))..((.((((....)))).))).))))))))))(((((.......))))).))))..... ( -56.70) >consensus CUAGUUUAUCCUUUGGAGCGAGUCCCCUCGCCGCCGGCGGCAGCGCAUACAGCUAGUAUGCGAACUCCCGAU_UCGGGUGUGGUGCCGCCGGUCCGGGUUCGAAUCCCGGUGAAGUUAAU ((((........))))...........(((((((((((((((.(((((((.....))))))).((.((((....)))).))..))))))))))..(((.......))))))))....... (-50.60 = -50.27 + -0.33)

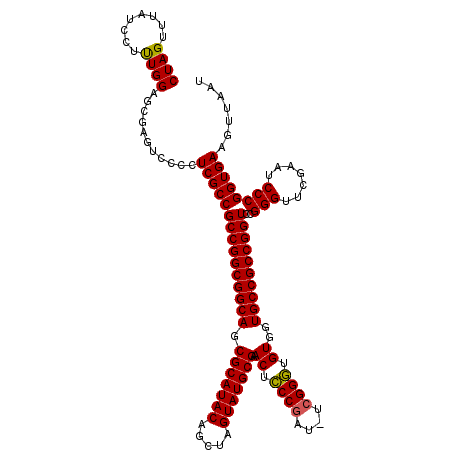
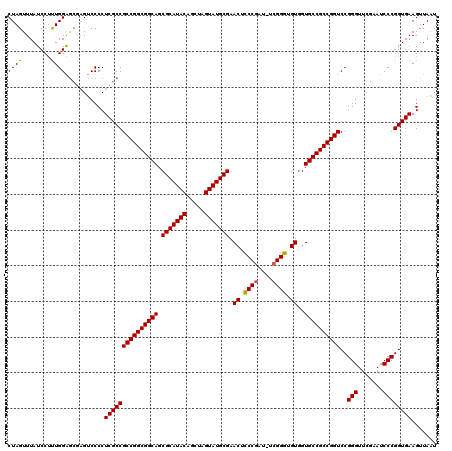
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:25:50 2006