
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 5,676,636 – 5,676,748 |
| Length | 112 |
| Max. P | 0.998414 |

| Location | 5,676,636 – 5,676,748 |
|---|---|
| Length | 112 |
| Sequences | 3 |
| Columns | 112 |
| Reading direction | forward |
| Mean pairwise identity | 80.61 |
| Mean single sequence MFE | -30.03 |
| Consensus MFE | -23.07 |
| Energy contribution | -23.73 |
| Covariance contribution | 0.67 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.51 |
| Structure conservation index | 0.77 |
| SVM decision value | 1.96 |
| SVM RNA-class probability | 0.984125 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 5676636 112 + 22224390 AAUCAACAAUCGGUGGCAAGACAACAUCUGCUGCAGUUGCAGUUGCACUUGCACUUGCUUGAGUGCUGAUGUUGCCAAUAUUUGUUGUAAAGCUGGGUAGAAAAUAUAUUGG ...((((((....(((((.....(((((.(((((....))))).((((((((....))..)))))).))))))))))....))))))......................... ( -32.00) >DroSec_CAF1 15589 86 + 1 AAUCAACAAUCGGUGGCAACACAACAUCUGCAGCAGUUGCAGUUGCACU------UGCAUGAGUGCUGAUGUUGCCAAUAUUUGUAGUAAAG-------------------- .....((((....((((((((((....((((((...))))))..(((((------(....)))))))).))))))))....)))).......-------------------- ( -28.40) >DroSim_CAF1 16166 106 + 1 AAUCAACAAUCGGUGGCAACACAACAUCUGCUGCAGUUGCAGUUGCACU------UGCUUGAGUGCUGAUGUUGCCAAUAUUUGUAGUAAAGUUGGAUAGAAAAUAUAAUAG .....((((....((((((((((......(((((....))))).(((((------(....)))))))).))))))))....))))........................... ( -29.70) >consensus AAUCAACAAUCGGUGGCAACACAACAUCUGCUGCAGUUGCAGUUGCACU______UGCUUGAGUGCUGAUGUUGCCAAUAUUUGUAGUAAAG_UGG_UAGAAAAUAUA_U_G .....((((....((((((((((......(((((....))))).(((((............))))))).))))))))....))))........................... (-23.07 = -23.73 + 0.67)


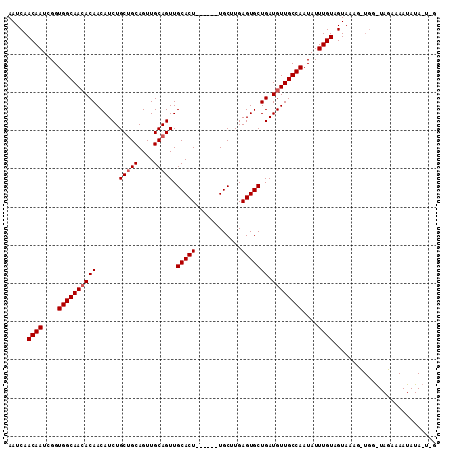
| Location | 5,676,636 – 5,676,748 |
|---|---|
| Length | 112 |
| Sequences | 3 |
| Columns | 112 |
| Reading direction | reverse |
| Mean pairwise identity | 80.61 |
| Mean single sequence MFE | -28.73 |
| Consensus MFE | -24.47 |
| Energy contribution | -24.80 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -3.53 |
| Structure conservation index | 0.85 |
| SVM decision value | 3.09 |
| SVM RNA-class probability | 0.998414 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 5676636 112 - 22224390 CCAAUAUAUUUUCUACCCAGCUUUACAACAAAUAUUGGCAACAUCAGCACUCAAGCAAGUGCAAGUGCAACUGCAACUGCAGCAGAUGUUGUCUUGCCACCGAUUGUUGAUU .........................((((((....((((((((((.(((((...((....)).)))))..((((....))))..))))).....)))))....))))))... ( -29.70) >DroSec_CAF1 15589 86 - 1 --------------------CUUUACUACAAAUAUUGGCAACAUCAGCACUCAUGCA------AGUGCAACUGCAACUGCUGCAGAUGUUGUGUUGCCACCGAUUGUUGAUU --------------------.....(.((((....((((((((.(((((((((.(((------..(((....)))..))))).)).)))))))))))))....)))).)... ( -29.10) >DroSim_CAF1 16166 106 - 1 CUAUUAUAUUUUCUAUCCAACUUUACUACAAAUAUUGGCAACAUCAGCACUCAAGCA------AGUGCAACUGCAACUGCAGCAGAUGUUGUGUUGCCACCGAUUGUUGAUU .........................(.((((....((((((((.(((((((...(((------..(((....)))..)))...)).)))))))))))))....)))).)... ( -27.40) >consensus C_A_UAUAUUUUCUA_CCA_CUUUACUACAAAUAUUGGCAACAUCAGCACUCAAGCA______AGUGCAACUGCAACUGCAGCAGAUGUUGUGUUGCCACCGAUUGUUGAUU .........................(.((((....((((((((.(((((.....(((........)))..((((.......)))).)))))))))))))....)))).)... (-24.47 = -24.80 + 0.33)

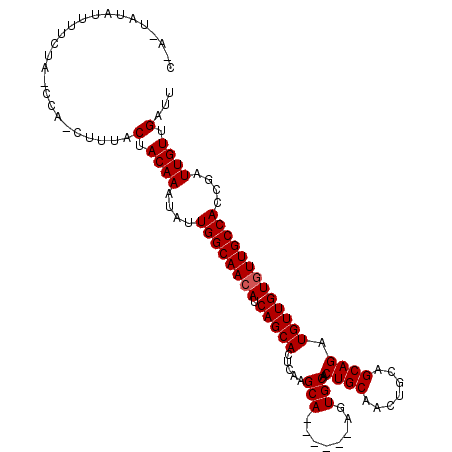

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:25:48 2006