
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 5,675,663 – 5,675,756 |
| Length | 93 |
| Max. P | 0.972988 |

| Location | 5,675,663 – 5,675,756 |
|---|---|
| Length | 93 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 78.42 |
| Mean single sequence MFE | -26.60 |
| Consensus MFE | -16.60 |
| Energy contribution | -15.85 |
| Covariance contribution | -0.75 |
| Combinations/Pair | 1.16 |
| Mean z-score | -1.91 |
| Structure conservation index | 0.62 |
| SVM decision value | 0.55 |
| SVM RNA-class probability | 0.779126 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 5675663 93 + 22224390 AUAAUCGCUCGAGUACU-UGGUAAAAAGCCAAUAAAGAAGAGGAAAAUGAAAAUACUUUCCUUUGAGCGCAGGUG-----------------AGCGG-GGCACGGAGAAUGG-------- ....(((((((.(..((-((((.....)))........((((((((((....))..)))))))))))..)...))-----------------)))))-..............-------- ( -21.60) >DroSec_CAF1 14617 95 + 1 AUAAUCGCUCGAGUACU-UGGUAAAAAGCCAAUAAAGAAGAGGAAAAUGAAAAUACUUUCCUUUGAGCUCAGGUG----------------GAGCGGGGGCACGGAGAAUGG-------- ....((((((((((..(-((((.....)))))......((((((((((....))..))))))))..)))).....----------------))))))...............-------- ( -21.80) >DroAna_CAF1 18784 120 + 1 AUAAUCGCUCGAAUACAACGGCAAAAAGCCAAUAAAGGAGAGGAAAAUGAAAAUAUUUUCCUUUGAGCUCAGUCCAGGGAGGUGCUGGCUGGAGGGAGGUCCUGGUGGGCGGGAGGUCCU ......((((.........(((.....)))........(((((((((((....)))))))))))))))((((.((((.......)))))))).((((..(((((.....)))))..)))) ( -40.30) >DroSim_CAF1 15187 94 + 1 AUAAUCGCUCGAGUACU-UGGUAAAAAGCCAAUAAAGAAGAGGAAAAUGAAAAUACUUUCCUUUGAGCUCAGGUG-----------------AGCGGGGGCACGGAGAAUGG-------- ....((((..((((..(-((((.....)))))......((((((((((....))..))))))))..))))..)))-----------------)((....))...........-------- ( -22.70) >consensus AUAAUCGCUCGAGUACU_UGGUAAAAAGCCAAUAAAGAAGAGGAAAAUGAAAAUACUUUCCUUUGAGCUCAGGUG_________________AGCGGGGGCACGGAGAAUGG________ ....((((((.........(((.....)))........((((((((..........)))))))))))).........................((....)).....))............ (-16.60 = -15.85 + -0.75)

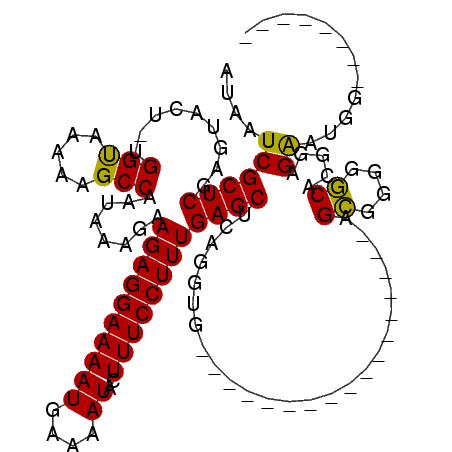
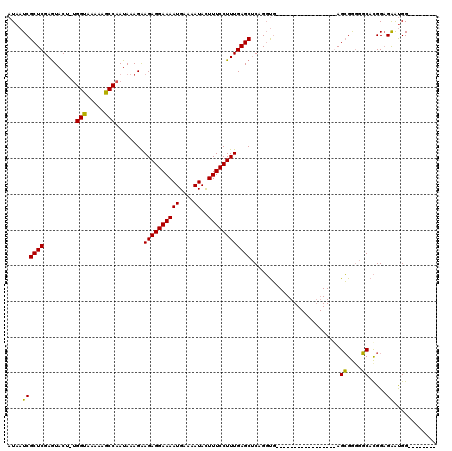
| Location | 5,675,663 – 5,675,756 |
|---|---|
| Length | 93 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 78.42 |
| Mean single sequence MFE | -21.48 |
| Consensus MFE | -14.93 |
| Energy contribution | -14.42 |
| Covariance contribution | -0.50 |
| Combinations/Pair | 1.17 |
| Mean z-score | -2.21 |
| Structure conservation index | 0.69 |
| SVM decision value | 1.70 |
| SVM RNA-class probability | 0.972988 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 5675663 93 - 22224390 --------CCAUUCUCCGUGCC-CCGCU-----------------CACCUGCGCUCAAAGGAAAGUAUUUUCAUUUUCCUCUUCUUUAUUGGCUUUUUACCA-AGUACUCGAGCGAUUAU --------..............-.((((-----------------(...(((......((((((((......))))))))........((((.......)))-))))...)))))..... ( -18.30) >DroSec_CAF1 14617 95 - 1 --------CCAUUCUCCGUGCCCCCGCUC----------------CACCUGAGCUCAAAGGAAAGUAUUUUCAUUUUCCUCUUCUUUAUUGGCUUUUUACCA-AGUACUCGAGCGAUUAU --------.....(((.((((....((((----------------.....))))....((((((((......))))))))........((((.......)))-)))))..)))....... ( -17.90) >DroAna_CAF1 18784 120 - 1 AGGACCUCCCGCCCACCAGGACCUCCCUCCAGCCAGCACCUCCCUGGACUGAGCUCAAAGGAAAAUAUUUUCAUUUUCCUCUCCUUUAUUGGCUUUUUGCCGUUGUAUUCGAGCGAUUAU .(((..(((.(.....).)))..)))...(((((((.......)))).))).((((..((((((((......))))))))..........(((.....))).........))))...... ( -30.80) >DroSim_CAF1 15187 94 - 1 --------CCAUUCUCCGUGCCCCCGCU-----------------CACCUGAGCUCAAAGGAAAGUAUUUUCAUUUUCCUCUUCUUUAUUGGCUUUUUACCA-AGUACUCGAGCGAUUAU --------.....(((.((((....(((-----------------(....))))....((((((((......))))))))........((((.......)))-)))))..)))....... ( -18.90) >consensus ________CCAUUCUCCGUGCCCCCGCU_________________CACCUGAGCUCAAAGGAAAGUAUUUUCAUUUUCCUCUUCUUUAUUGGCUUUUUACCA_AGUACUCGAGCGAUUAU .................(((....))).........................((((..((((((((......)))))))).........(((.......)))........))))...... (-14.93 = -14.42 + -0.50)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:25:44 2006