
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 5,586,962 – 5,587,060 |
| Length | 98 |
| Max. P | 0.965750 |

| Location | 5,586,962 – 5,587,060 |
|---|---|
| Length | 98 |
| Sequences | 5 |
| Columns | 98 |
| Reading direction | forward |
| Mean pairwise identity | 81.28 |
| Mean single sequence MFE | -47.36 |
| Consensus MFE | -35.76 |
| Energy contribution | -35.78 |
| Covariance contribution | 0.02 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.68 |
| Structure conservation index | 0.76 |
| SVM decision value | 0.42 |
| SVM RNA-class probability | 0.729101 |
| Prediction | RNA |
| WARNING | Sequence 3: Base composition out of range. |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 5586962 98 + 22224390 CCGGCAUUCCGGGCAUCCCGGGACCCGAACCACCGCCGGGCGAAGAGCCCGGGGAUGAGGGACCCGGUCCAGUGGGCGAGCCCGAUCCGGAAGGUGAG .((.(.((((((((...(((((.(((....((((.((((((.....)))))))).)).))).)))))......(((....)))).)))))))).)).. ( -49.50) >DroSec_CAF1 15108 98 + 1 CGGCCAUUCCGGGCAUCCCGGGACCCGAAACACCGCCGGGCGAAGAGCCCGGGGAUGAGGGGCCCGGUCCAGUGGGCGGGCCCGAUCCGGAAGGUGAG ..(((.(((((((((((((((...........)))((((((.....)))))))))))..(((((((.(((...))))))))))..))))))))))... ( -55.10) >DroSim_CAF1 15081 98 + 1 CGGCCAUUCCGGGCAUCCCGGGACCCGAAACACCGCCGGGCGAAGAGCCCGGGGAUGAGGGGCCCGGUCCAGUGGGCGGGCCNNNNNNNNNNNNNNNN .((((.(((((((...)))))))(((....((((.((((((.....)))))))).)).)))(((((......))))).))))................ ( -46.50) >DroEre_CAF1 15577 90 + 1 --------CCAGGCAUCCCGGGACCCGAGCCACCGCCUGGCGACGAGCCAGGGGAUGAGGGCCCCGGUCCAGUGGGCGAACCCGAUCCGGAUGGUGAG --------.....(((((((((.(((....((((.((((((.....)))))))).)).))).)))))(((..((((....))))....))).)))).. ( -40.70) >DroYak_CAF1 15638 90 + 1 --------CCAGGCAUCCCGGGACCGGAGCCACCGCCUGGCGACGAGCCAGGGGACGAGGGGCCCGGUCCAGUGGGCGAACCCGAUCCGGAAGGUGAG --------.....((((((((((((((.(((.((.((((((.....))))))......))))))))))))..((((....))))...)))..)))).. ( -45.00) >consensus C_G_CAUUCCGGGCAUCCCGGGACCCGAACCACCGCCGGGCGAAGAGCCCGGGGAUGAGGGGCCCGGUCCAGUGGGCGAGCCCGAUCCGGAAGGUGAG ...........((....(((((.(((......((.((((((.....))))))))....))).))))).))...(((....)))...((....)).... (-35.76 = -35.78 + 0.02)



| Location | 5,586,962 – 5,587,060 |
|---|---|
| Length | 98 |
| Sequences | 5 |
| Columns | 98 |
| Reading direction | reverse |
| Mean pairwise identity | 81.28 |
| Mean single sequence MFE | -48.76 |
| Consensus MFE | -39.04 |
| Energy contribution | -38.60 |
| Covariance contribution | -0.44 |
| Combinations/Pair | 1.15 |
| Mean z-score | -2.23 |
| Structure conservation index | 0.80 |
| SVM decision value | 1.59 |
| SVM RNA-class probability | 0.965750 |
| Prediction | RNA |
| WARNING | Sequence 3: Base composition out of range. |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 5586962 98 - 22224390 CUCACCUUCCGGAUCGGGCUCGCCCACUGGACCGGGUCCCUCAUCCCCGGGCUCUUCGCCCGGCGGUGGUUCGGGUCCCGGGAUGCCCGGAAUGCCGG ........((((.((((((((.((....)).(((((.((((((((.((((((.....)))))).)))))...))).))))))).))))))....)))) ( -52.90) >DroSec_CAF1 15108 98 - 1 CUCACCUUCCGGAUCGGGCCCGCCCACUGGACCGGGCCCCUCAUCCCCGGGCUCUUCGCCCGGCGGUGUUUCGGGUCCCGGGAUGCCCGGAAUGGCCG ....((((((((((((((....)))......((((((((..((((.((((((.....)))))).))))....))).))))))))..)))))).))... ( -51.50) >DroSim_CAF1 15081 98 - 1 NNNNNNNNNNNNNNNNGGCCCGCCCACUGGACCGGGCCCCUCAUCCCCGGGCUCUUCGCCCGGCGGUGUUUCGGGUCCCGGGAUGCCCGGAAUGGCCG ................((((..((....((.((((((((..((((.((((((.....)))))).))))....))).)))))....)).))...)))). ( -44.90) >DroEre_CAF1 15577 90 - 1 CUCACCAUCCGGAUCGGGUUCGCCCACUGGACCGGGGCCCUCAUCCCCUGGCUCGUCGCCAGGCGGUGGCUCGGGUCCCGGGAUGCCUGG-------- ....((((((((...(((....))).)))))((((((((((((((.((((((.....)))))).)))))...)))))))))......)))-------- ( -48.50) >DroYak_CAF1 15638 90 - 1 CUCACCUUCCGGAUCGGGUUCGCCCACUGGACCGGGCCCCUCGUCCCCUGGCUCGUCGCCAGGCGGUGGCUCCGGUCCCGGGAUGCCUGG-------- ....((....))..(((((...(((...((((((((((......((((((((.....)))))).)).))).))))))).)))..))))).-------- ( -46.00) >consensus CUCACCUUCCGGAUCGGGCUCGCCCACUGGACCGGGCCCCUCAUCCCCGGGCUCUUCGCCCGGCGGUGGUUCGGGUCCCGGGAUGCCCGGAAUG_C_G .............((((((...(((...(((((.((.....((((.((((((.....)))))).))))..)).))))).)))..))))))........ (-39.04 = -38.60 + -0.44)


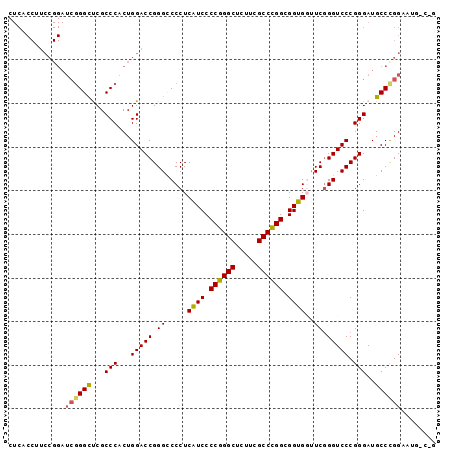
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:24:37 2006