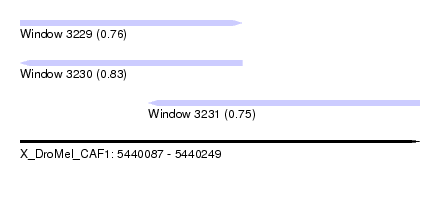

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 5,440,087 – 5,440,249 |
| Length | 162 |
| Max. P | 0.833471 |

| Location | 5,440,087 – 5,440,177 |
|---|---|
| Length | 90 |
| Sequences | 4 |
| Columns | 99 |
| Reading direction | forward |
| Mean pairwise identity | 85.89 |
| Mean single sequence MFE | -39.20 |
| Consensus MFE | -27.14 |
| Energy contribution | -28.82 |
| Covariance contribution | 1.69 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.93 |
| Structure conservation index | 0.69 |
| SVM decision value | 0.51 |
| SVM RNA-class probability | 0.763164 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 5440087 90 + 22224390 CAGCGGCGGCGG-CAGCGGCUGCCG----CCCACUGCCAAU--UGCCCGUCUUCAAUCCGGC-CAACUUCUAUGCCGCCGCCGG-CUAUGCCGCUGCAG ((((((((((((-((((((...)))----).....(((.((--((........))))..)))-.........))))))))))((-(...)))))))... ( -41.80) >DroSec_CAF1 46912 97 + 1 CAG--GCGGCGGAGAGCGAGUGCUGUCCGCACAAAGACAAAUAUGCCCGUCUUCAAUCCGGCCCAAUUUCUAUGCCGCCACCGGAGUAUGCCGCUGCAG ..(--((((((((.(((....))).)))((...(((((..........)))))...(((((...................)))))))..)))))).... ( -30.01) >DroEre_CAF1 57819 90 + 1 CAGCGGCGGCGG-CGGCGGCUGCCG----CCCACUGCCAAC--UGCCCGUCUUCAAUCCGGC-CAACUUCUAUGCCGCCGCCGG-CUAUGCCGCUGCAG ..((((((((((-(((((((.....----......)))...--.(((.(........).)))-..........)))))))))((-(...)))))))).. ( -43.70) >DroYak_CAF1 48650 90 + 1 CAGCAGCGGCAG-CGGCGGCUGCCG----CCCACUGCCAAC--UGCCCGUCUUCAAUCCGGC-CAACUUCUAUGCCGCCGCCGG-CUAUGCCGCUGCAG ..(((((((((.-(((((((.((.(----(.....))....--.(((.(........).)))-..........)).))))))).-...))))))))).. ( -41.30) >consensus CAGCGGCGGCGG_CAGCGGCUGCCG____CCCACUGCCAAC__UGCCCGUCUUCAAUCCGGC_CAACUUCUAUGCCGCCGCCGG_CUAUGCCGCUGCAG ..(((((((((..(((((((.((.(......)............(((.(........).)))...........)).))))))).....))))))))).. (-27.14 = -28.82 + 1.69)


| Location | 5,440,087 – 5,440,177 |
|---|---|
| Length | 90 |
| Sequences | 4 |
| Columns | 99 |
| Reading direction | reverse |
| Mean pairwise identity | 85.89 |
| Mean single sequence MFE | -45.42 |
| Consensus MFE | -31.56 |
| Energy contribution | -32.56 |
| Covariance contribution | 1.00 |
| Combinations/Pair | 1.15 |
| Mean z-score | -2.11 |
| Structure conservation index | 0.69 |
| SVM decision value | 0.72 |
| SVM RNA-class probability | 0.833471 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 5440087 90 - 22224390 CUGCAGCGGCAUAG-CCGGCGGCGGCAUAGAAGUUG-GCCGGAUUGAAGACGGGCA--AUUGGCAGUGGG----CGGCAGCCGCUG-CCGCCGCCGCUG ......((((...)-)))((((((((.....(((((-.(((.........))).))--)))((((((((.----(....)))))))-)))))))))).. ( -50.40) >DroSec_CAF1 46912 97 - 1 CUGCAGCGGCAUACUCCGGUGGCGGCAUAGAAAUUGGGCCGGAUUGAAGACGGGCAUAUUUGUCUUUGUGCGGACAGCACUCGCUCUCCGCCGC--CUG .(((....)))......(((((((((..(....)...)))(((..(((((((((....)))))))))((((.....))))......))))))))--).. ( -34.30) >DroEre_CAF1 57819 90 - 1 CUGCAGCGGCAUAG-CCGGCGGCGGCAUAGAAGUUG-GCCGGAUUGAAGACGGGCA--GUUGGCAGUGGG----CGGCAGCCGCCG-CCGCCGCCGCUG ...(((((((...(-(.(((((((((........((-.(((.........))).))--((((.(....).----)))).)))))))-)))).))))))) ( -49.00) >DroYak_CAF1 48650 90 - 1 CUGCAGCGGCAUAG-CCGGCGGCGGCAUAGAAGUUG-GCCGGAUUGAAGACGGGCA--GUUGGCAGUGGG----CGGCAGCCGCCG-CUGCCGCUGCUG ..(((((((((...-...((((((((........((-.(((.........))).))--((((.(....).----)))).)))))))-)))))))))).. ( -48.00) >consensus CUGCAGCGGCAUAG_CCGGCGGCGGCAUAGAAGUUG_GCCGGAUUGAAGACGGGCA__GUUGGCAGUGGG____CGGCAGCCGCCG_CCGCCGCCGCUG ...(((((((.......(((((((((...........(((.(........).)))......(((.((.(.....).)).))))))).)))))))))))) (-31.56 = -32.56 + 1.00)


| Location | 5,440,139 – 5,440,249 |
|---|---|
| Length | 110 |
| Sequences | 6 |
| Columns | 116 |
| Reading direction | reverse |
| Mean pairwise identity | 83.45 |
| Mean single sequence MFE | -56.15 |
| Consensus MFE | -39.38 |
| Energy contribution | -39.03 |
| Covariance contribution | -0.36 |
| Combinations/Pair | 1.21 |
| Mean z-score | -2.51 |
| Structure conservation index | 0.70 |
| SVM decision value | 0.47 |
| SVM RNA-class probability | 0.747655 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 5440139 110 - 22224390 CCCGGAGAUAGGCGGCCGCCGCCAAGGUGGUCAUGUGGUGGUGAUGGGCUGCAGCCGCG------UUGCCCGGAUCCGCUGCAGCGGCAUAGCCGGCGGCGGCAUAGAAGUUGGCC .((((...((....(((((((((..(((.(((.((..((((...(((((.((......)------).)))))...))))..))..)))...)))))))))))).))....)))).. ( -54.50) >DroVir_CAF1 119248 110 - 1 CGCGCAAAUAGGCAGCCGCCGCCAGCGUCGUCAUAUGAUGAUGAUGGGCGGCCGCUGCA------UUGGCCGGAUCGGCGGCAGCGGCAUAGCCGGCGGCAGCAUAGAAAUUCGCC .((((......)).((.((((((..(((((((((...)))))))))(((.((((((((.------...((((...)))).))))))))...))))))))).))..........)). ( -59.50) >DroGri_CAF1 99544 110 - 1 CCCGCAGAUAGGCGGCCGCUGCCAGCGUCGUCAUGUGAUGAUGAUGGGCGGCGGCAGCG------UUCGCUGGAUCGGCGGCAGCCGCAUAGCCGGCGGCGGCAUAGAAAUUUGCC ...((((((...(.(((((((((..(((((((((...)))))))))(((.(((((.((.------..(((((...))))))).)))))...))))))))))))...)..)))))). ( -59.30) >DroWil_CAF1 111938 116 - 1 CUCUUAGAUAGGCGGCUGCUGCCAACGUUGUCAUGUGAUGAUGAUGGGCGGCUGCUGCGGCUGAAUUAUUUGGAUCGGCGGCUGCUGCAUAGCCGGCGGCUGCAUAAAAAUUGGCC ...........(((((.((((((..(((..((....))..)))...)))))).)))))((((.((((.((((.....((((((((((......)))))))))).)))))))))))) ( -50.50) >DroMoj_CAF1 125075 110 - 1 CGCGCAGAUAGGCGGCCGCUGCCAACGUCGUCAUGUGAUGGUGAUGGGCGGCAGCUGCA------UUAGCCGGAUCGGCGGCAGCGGCAUAGCCGGCGGCUGCAUAGAAAUUCGCC .(((....((.(((((((((((((.(..((((....))))..).)))((.((.(((((.------...((((...)))).))))).))...)))))))))))).))......))). ( -52.60) >DroAna_CAF1 65130 110 - 1 CCCGGAGAUAGGCGGCCGCCGCCAGCGUCGUCAUGUGGUGAUGGUGGGCAGCUGCCGCA------UUGCCCGGAUCGGCGGCGGCCGCAUAGCCGGCGGCAGCGUAGAAGUUGGCC .((((...((.((((((((((((..((((..(....)..)))).((((((((....)).------.))))))....)))))))))))).)).)))).(.((((......)))).). ( -60.50) >consensus CCCGCAGAUAGGCGGCCGCCGCCAACGUCGUCAUGUGAUGAUGAUGGGCGGCAGCUGCA______UUGGCCGGAUCGGCGGCAGCGGCAUAGCCGGCGGCAGCAUAGAAAUUGGCC ...........((.((((((((((.(((((((....))))))).)))((.((.(((((...................))))).)).)).....))))))).))............. (-39.38 = -39.03 + -0.36)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:23:30 2006