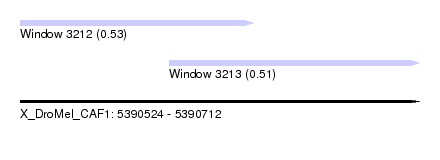
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 5,390,524 – 5,390,712 |
| Length | 188 |
| Max. P | 0.525454 |

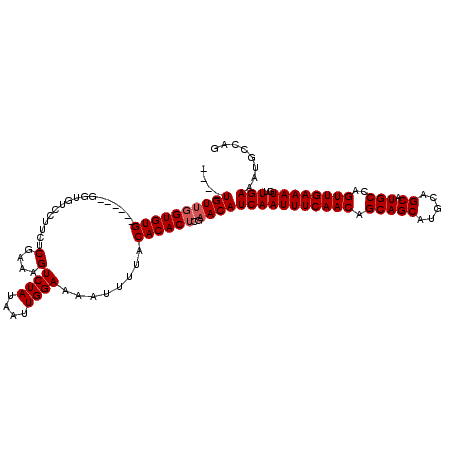
| Location | 5,390,524 – 5,390,634 |
|---|---|
| Length | 110 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 89.11 |
| Mean single sequence MFE | -30.54 |
| Consensus MFE | -20.79 |
| Energy contribution | -21.35 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.74 |
| Structure conservation index | 0.68 |
| SVM decision value | -0.02 |
| SVM RNA-class probability | 0.525454 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 5390524 110 + 22224390 ----UGUUGGUGUG------GGUGUCCUUCUCGAAAGUCUAUAAUUGGAAAAUUUUACACACUCGAACAUCAAUUUCAACAGCAGCAUGCAGCAUGCCAGUUGAAAUGUUGAAAUGCCAA ----..(((((...------(((((.....((((..((.((.((((....)))).)).))..))))))))).(((((((((.((((..((.....))..))))...)))))))))))))) ( -28.80) >DroSec_CAF1 71653 103 + 1 ----UGUGGGUGUG------GGUGUCCUUCUCGAAAGUCUAUAAUUGGACAAUUUUACACACUCGAACAUCAAUUUCAACAGCAG-------CAUGCCAGUUGAAAUGUUGAAAUGCCAG ----..((((((((------..(((((....(....).........)))))......)))))))).......(((((((((.(((-------(......))))...)))))))))..... ( -30.22) >DroSim_CAF1 59384 114 + 1 ----UGUUGGUGUGG--GUGGGUGUCCUUCUCGAAAGUCUAUAAUUGGAAAAUUUUACACACUCGAACAUCAAUUUCAACAGCAGCAUGCAGCAUGCCAGUUGAAAUGUUGAAAUGCCAG ----...(((((((.--.(((((((......(....)((((....)))).........)))))))..)))..(((((((((.((((..((.....))..))))...))))))))))))). ( -32.00) >DroYak_CAF1 65190 120 + 1 UGUGUGUUUGUGUGGGAGUGAGAGUCCUUCUCGAAAGUCUAUAAUUGGAAAAUUUUACACACUCGAACAUCAAUUUCAACAGCAGCAUGCAGCAUGCCAGUUGAAAUGCUGAAAUGCCAG ((.(((((((.(((...((((((.(((....(....).........)))...)))))).))).)))))))))((((((((.(((((.....)).)))..))))))))((......))... ( -31.12) >consensus ____UGUUGGUGUG______GGUGUCCUUCUCGAAAGUCUAUAAUUGGAAAAUUUUACACACUCGAACAUCAAUUUCAACAGCAGCAUGCAGCAUGCCAGUUGAAAUGUUGAAAUGCCAG ....((((((((((.................(....)((((....))))........))))))..))))(((((((((((.(((((.....)).)))..))))))))..)))........ (-20.79 = -21.35 + 0.56)

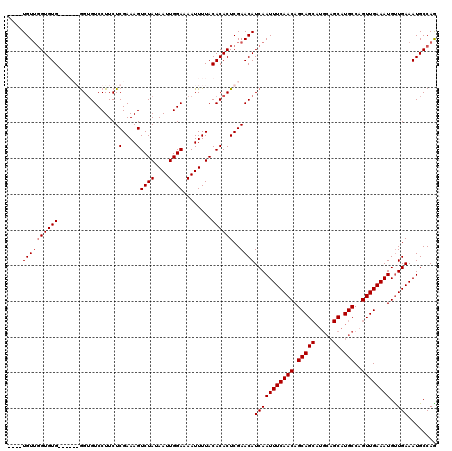
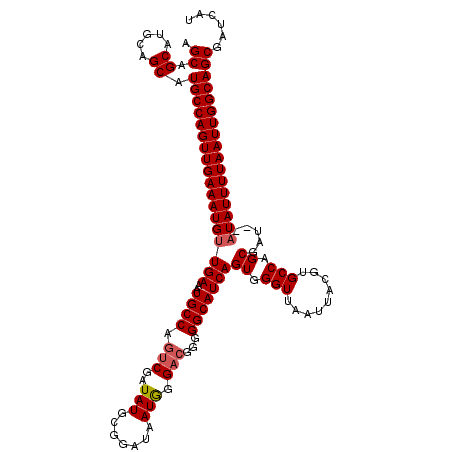
| Location | 5,390,594 – 5,390,712 |
|---|---|
| Length | 118 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 92.02 |
| Mean single sequence MFE | -34.05 |
| Consensus MFE | -29.90 |
| Energy contribution | -30.53 |
| Covariance contribution | 0.63 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.34 |
| Structure conservation index | 0.88 |
| SVM decision value | -0.04 |
| SVM RNA-class probability | 0.513401 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 5390594 118 + 22224390 AGCAGCAUGCAGCAUGCCAGUUGAAAUGUUGAAAUGCCAAUCGAUAUGCGGAUAAUGGGACGGGGGCAUCAGUGGGUUAAUUACGUGCCAGCUAU--AUAUUUUAAUUGGCAGCGAUCAU .((.....)).((.(((((((((((((((....(((((..(((.(((.......)))...))).))))).(((.(((.........))).)))..--)))))))))))))))))...... ( -32.40) >DroSec_CAF1 71723 110 + 1 AGCAG-------CAUGCCAGUUGAAAUGUUGAAAUGCCAGUCGAUAUGCGGAUAAUGGGACAGGGGCAUCAGUGGGUUAAU-GUGUGCCAGCGAU--AUAUUUUAAUUGGCAGCGAUCAU ....(-------(.((((((((((((((((((..((((.(((..(((.......))).)))...)))))))((.(((....-....))).))...--)))))))))))))))))...... ( -33.50) >DroSim_CAF1 59458 118 + 1 AGCAGCAUGCAGCAUGCCAGUUGAAAUGUUGAAAUGCCAGUCGAUAUGCGGAUAAUGGGACGGGGGCAUCAGUGGGUUAAUUACGUGCCAGCGAU--AUAUUUUAAUUGGCAGCGAUCAU .((.....)).((.(((((((((((((((.......(((.(((.....)))....)))..((..(((((..((((.....)))))))))..))..--)))))))))))))))))...... ( -35.60) >DroYak_CAF1 65270 120 + 1 AGCAGCAUGCAGCAUGCCAGUUGAAAUGCUGAAAUGCCAGUCGAUAUGCGGAUAAUAGGCCGGGGGCAUCAGUGGGUUAAUUACGUGCCAGCGAUAUAUAUUUUAAUUGGCAGCGAUCUU .((.....)).((.(((((((((((((((......))..((((...(.(((........))).)(((((..((((.....)))))))))..))))....)))))))))))))))...... ( -34.70) >consensus AGCAGCAUGCAGCAUGCCAGUUGAAAUGUUGAAAUGCCAGUCGAUAUGCGGAUAAUGGGACGGGGGCAUCAGUGGGUUAAUUACGUGCCAGCGAU__AUAUUUUAAUUGGCAGCGAUCAU .((.((.....)).((((((((((((((((((..((((.(((..(((.......))).)))...)))))))((.(((.........))).)).....)))))))))))))))))...... (-29.90 = -30.53 + 0.63)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:23:14 2006