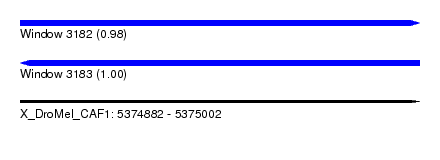

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 5,374,882 – 5,375,002 |
| Length | 120 |
| Max. P | 0.997089 |

| Location | 5,374,882 – 5,375,002 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 95.25 |
| Mean single sequence MFE | -59.03 |
| Consensus MFE | -51.62 |
| Energy contribution | -52.10 |
| Covariance contribution | 0.48 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.84 |
| Structure conservation index | 0.87 |
| SVM decision value | 1.84 |
| SVM RNA-class probability | 0.979467 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
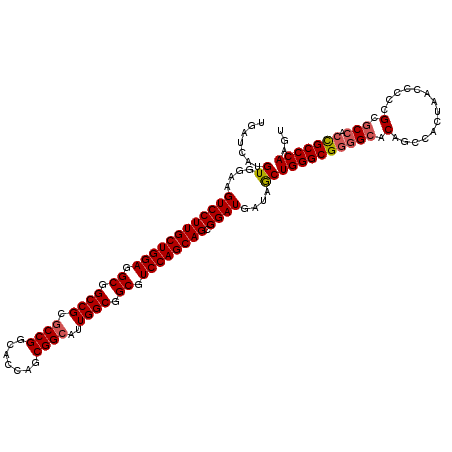
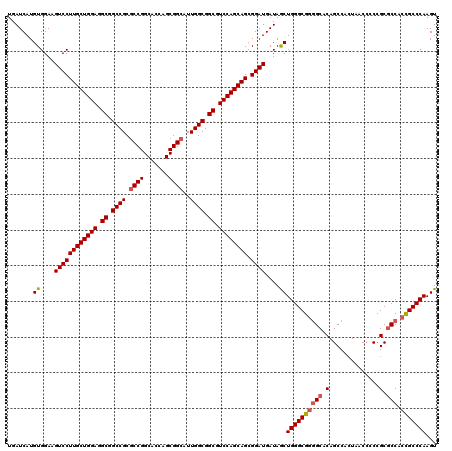
>X_DroMel_CAF1 5374882 120 + 22224390 UGAUCAUGUGGAAGUCCUUGCUGGAGGCGGCCGCGCCGGCACCAGCGGCAUUGGCGGCGUCCAGCAGCGGAUGAUAGCUGGGCGGGGCACAGCCAGUAACCACCGCGCCAGCGCCCAAGU ........(((..((((((((((((.((.((((.((((.......))))..)))).)).)))))))).))))....(((((((((((.((.....))..)).)))).)))))..)))... ( -59.30) >DroSec_CAF1 56342 120 + 1 UGAUCAUGUGGAAGUCCUUGCUGGAGGCAGCCGCGCCGGCACCAGCGGCAUUGGCGGCGUCCAGCAGCGGAUGAUAACUGGGCGGGGCACAGCCACUAACCGCCGCGCCACCGCCCAAGU .......((....((((((((((((.((.((((.((((.......))))..)))).)).)))))))).))))....))((((((((((.(.((........)).).))).)))))))... ( -60.10) >DroSim_CAF1 43296 120 + 1 UGAUCAUGUGGAAGUCCUUGCUGGAAGCGGCCGCGCCGGCACCAGCGGCAUUGGCGGCGUCCAGCAGCGGAUGAUAGCUGGGCGGGGCACAGCCACUAACCGCCGCGCCACCGCCCAAGU .......((....((((((((((((.((.((((.((((.......))))..)))).)).)))))))).))))....))((((((((((.(.((........)).).))).)))))))... ( -60.10) >DroEre_CAF1 49605 120 + 1 UGAUCAUGUGGAAGUCCUUGCUGGAGGCGGCCGCGCCGGCACCAGCGGCAUUGGCGGCGUCCAGCAGCGGAUGAUAGCUGGGCGGGGCACAGCCACUAACGCCCGCCCCGCUGCCCAAGU ........(((..((((((((((((.((.((((.((((.......))))..)))).)).)))))))).))))..((((.(((((((((...))).(....).)))))).)))).)))... ( -60.60) >DroYak_CAF1 48594 120 + 1 UGAUCAUGUGGAAGUCCUUGCUGGAGGCCGCCGCUCCGGCACCAGCGGCAUUGGCGGCGUCCAGCAGCGGAUGAUAGCUGGGCGGAGCACAGCCACUAACCCCCGCGCCGCCGCCCAAGU .......((....((((((((((((.(((((((..(((.......)))...))))))).)))))))).))))....))(((((((.((.(..............).))..)))))))... ( -55.04) >consensus UGAUCAUGUGGAAGUCCUUGCUGGAGGCGGCCGCGCCGGCACCAGCGGCAUUGGCGGCGUCCAGCAGCGGAUGAUAGCUGGGCGGGGCACAGCCACUAACCCCCGCGCCACCGCCCAAGU .......((....((((((((((((.((.((((.((((.......))))..)))).)).)))))))).))))....))((((((((((.(..............).))).)))))))... (-51.62 = -52.10 + 0.48)

| Location | 5,374,882 – 5,375,002 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 95.25 |
| Mean single sequence MFE | -59.66 |
| Consensus MFE | -55.14 |
| Energy contribution | -56.10 |
| Covariance contribution | 0.96 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.93 |
| Structure conservation index | 0.92 |
| SVM decision value | 2.80 |
| SVM RNA-class probability | 0.997089 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
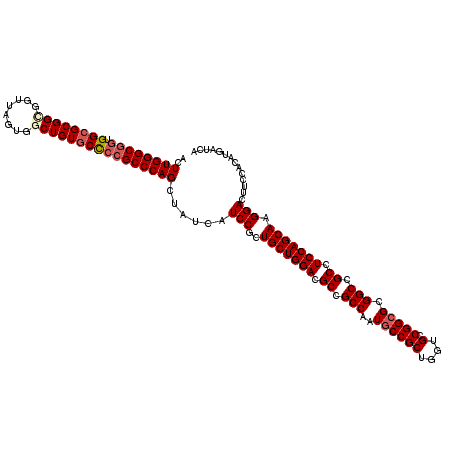
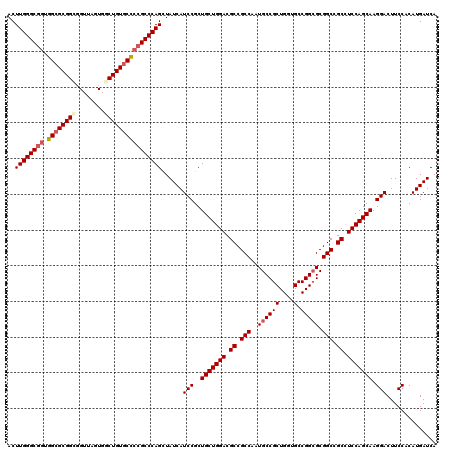
>X_DroMel_CAF1 5374882 120 - 22224390 ACUUGGGCGCUGGCGCGGUGGUUACUGGCUGUGCCCCGCCCAGCUAUCAUCCGCUGCUGGACGCCGCCAAUGCCGCUGGUGCCGGCGCGGCCGCCUCCAGCAAGGACUUCCACAUGAUCA ...((((.(((((.((((.(((.((.....)))))))))))))).....(((..(((((((.((.(((..((((((....).))))).))).)).))))))).)))..))))........ ( -56.80) >DroSec_CAF1 56342 120 - 1 ACUUGGGCGGUGGCGCGGCGGUUAGUGGCUGUGCCCCGCCCAGUUAUCAUCCGCUGCUGGACGCCGCCAAUGCCGCUGGUGCCGGCGCGGCUGCCUCCAGCAAGGACUUCCACAUGAUCA ..((((((((.((((((((........))))))))))))))))......(((..(((((((.((.(((..((((((....).))))).))).)).))))))).))).............. ( -63.60) >DroSim_CAF1 43296 120 - 1 ACUUGGGCGGUGGCGCGGCGGUUAGUGGCUGUGCCCCGCCCAGCUAUCAUCCGCUGCUGGACGCCGCCAAUGCCGCUGGUGCCGGCGCGGCCGCUUCCAGCAAGGACUUCCACAUGAUCA ..((((((((.((((((((........))))))))))))))))......(((..(((((((.((.(((..((((((....).))))).))).)).))))))).))).............. ( -63.50) >DroEre_CAF1 49605 120 - 1 ACUUGGGCAGCGGGGCGGGCGUUAGUGGCUGUGCCCCGCCCAGCUAUCAUCCGCUGCUGGACGCCGCCAAUGCCGCUGGUGCCGGCGCGGCCGCCUCCAGCAAGGACUUCCACAUGAUCA ...((((.(((.(((((((.((..........))))))))).)))....(((..(((((((.((.(((..((((((....).))))).))).)).))))))).)))..))))........ ( -55.60) >DroYak_CAF1 48594 120 - 1 ACUUGGGCGGCGGCGCGGGGGUUAGUGGCUGUGCUCCGCCCAGCUAUCAUCCGCUGCUGGACGCCGCCAAUGCCGCUGGUGCCGGAGCGGCGGCCUCCAGCAAGGACUUCCACAUGAUCA ..((((((((.(((((((..........)))))))))))))))......(((..(((((((.((((((....((((....).)))...)))))).))))))).))).............. ( -58.80) >consensus ACUUGGGCGGUGGCGCGGCGGUUAGUGGCUGUGCCCCGCCCAGCUAUCAUCCGCUGCUGGACGCCGCCAAUGCCGCUGGUGCCGGCGCGGCCGCCUCCAGCAAGGACUUCCACAUGAUCA ..((((((((.((((((((........))))))))))))))))......(((..(((((((.((.(((..((((((....).))))).))).)).))))))).))).............. (-55.14 = -56.10 + 0.96)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:22:48 2006