
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 5,198,749 – 5,198,849 |
| Length | 100 |
| Max. P | 0.999470 |

| Location | 5,198,749 – 5,198,849 |
|---|---|
| Length | 100 |
| Sequences | 5 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 77.98 |
| Mean single sequence MFE | -23.32 |
| Consensus MFE | -11.53 |
| Energy contribution | -11.85 |
| Covariance contribution | 0.32 |
| Combinations/Pair | 1.35 |
| Mean z-score | -2.17 |
| Structure conservation index | 0.49 |
| SVM decision value | 0.94 |
| SVM RNA-class probability | 0.886427 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
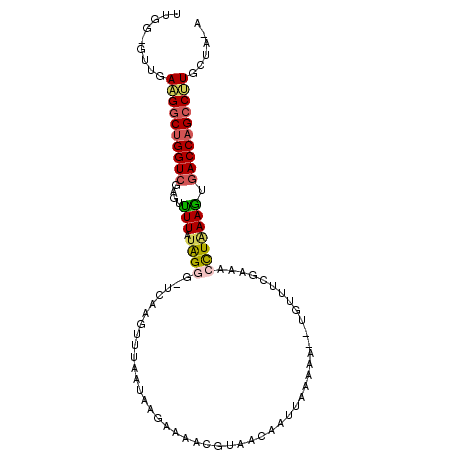
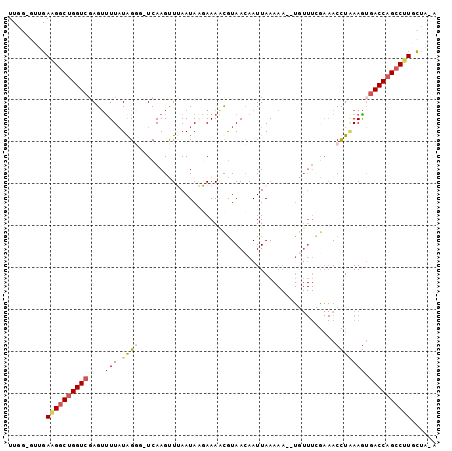
>X_DroMel_CAF1 5198749 100 + 22224390 UUGG-GUUGAAGGCUGGUAGAGUUUUAUGGGUUUCAAGUUUAAUAAGAAAACGUAACAAUUAAAAAUAUGUUUCGAAACCUUAAGUGACCAGCCUUUCGA-A ....-.((((((((((((.....((((.(((((((..((((.......))))(.((((..........)))).))))))))))))..))))))).)))))-. ( -27.10) >DroSec_CAF1 19472 98 + 1 UUGGGGUUGAAGGCUGGUCGAGUUUUAUAGGG-UCAAGCUUAAUAAGAAAACGUAACAAUUAAAAA--UGUUUCGAAACCUAAAGUGACCAGCCUUGCUA-A ....(((..(((((((((((..((((((..((-.....))..))))))...((.((((........--)))).))..........)))))))))))))).-. ( -26.70) >DroSim_CAF1 13251 97 + 1 UUGG-GUUGAAGGCUGGUCGAGUUUUAUAGGG-UCAAGCUUAAUAAGAAAACGUAACAAUUAAAAA--UGUUUCGAAACCUAAAGUGACCAGCCUUGCUA-A ...(-((..(((((((((((..((((((..((-.....))..))))))...((.((((........--)))).))..........)))))))))))))).-. ( -26.70) >DroEre_CAF1 22369 86 + 1 UUGG-GUUGAUGGCUGGUCCAGAUUUAUGGCA-UCAAGUUAAAUAUUAAAACAUAACAAUUAA--------------ACCUGAAAUGACCAGACAUUUUAAA .((.-.(((((((((((((((......))).)-)).)))....))))))..))..........--------------...(((((((......))))))).. ( -13.10) >DroYak_CAF1 19859 97 + 1 UUGG-CUUGAAGGCCGGUCGAGUCUUCUAGCC-UCAAGUUAAAUAUUAAAGUGUAAGAAUUAAAAA--UAUUUUAAAACUUAAAGGGACCAGCCCUGCUG-A ..((-((.(((((((....).)))))).))))-..(((((.....(((((((((...........)--)))))))))))))..((((.....))))....-. ( -23.00) >consensus UUGG_GUUGAAGGCUGGUCGAGUUUUAUAGGG_UCAAGUUUAAUAAGAAAACGUAACAAUUAAAAA__UGUUUCGAAACCUAAAGUGACCAGCCUUGCUA_A .........((((((((((....(((.((((...............................................))))))).))))))))))...... (-11.53 = -11.85 + 0.32)

| Location | 5,198,749 – 5,198,849 |
|---|---|
| Length | 100 |
| Sequences | 5 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 77.98 |
| Mean single sequence MFE | -22.78 |
| Consensus MFE | -14.29 |
| Energy contribution | -13.97 |
| Covariance contribution | -0.32 |
| Combinations/Pair | 1.35 |
| Mean z-score | -3.37 |
| Structure conservation index | 0.63 |
| SVM decision value | 3.63 |
| SVM RNA-class probability | 0.999470 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
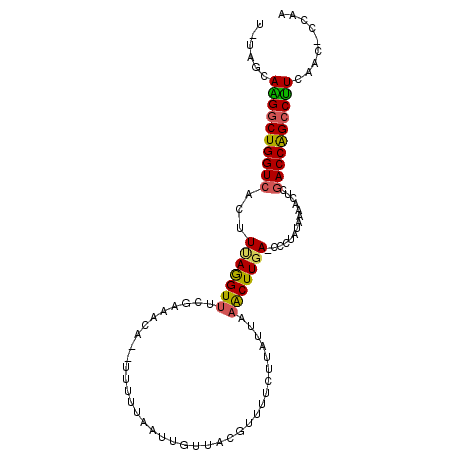
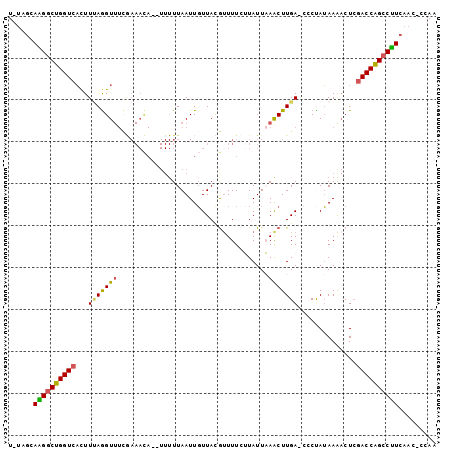
>X_DroMel_CAF1 5198749 100 - 22224390 U-UCGAAAGGCUGGUCACUUAAGGUUUCGAAACAUAUUUUUAAUUGUUACGUUUUCUUAUUAAACUUGAAACCCAUAAAACUCUACCAGCCUUCAAC-CCAA .-..(((.(((((((.......((((((((((((..........))))..((((.......))))))))))))...........))))))))))...-.... ( -23.07) >DroSec_CAF1 19472 98 - 1 U-UAGCAAGGCUGGUCACUUUAGGUUUCGAAACA--UUUUUAAUUGUUACGUUUUCUUAUUAAGCUUGA-CCCUAUAAAACUCGACCAGCCUUCAACCCCAA .-..(.((((((((((....((((..((((((((--........))))..((((.......))))))))-.))))........)))))))))))........ ( -23.80) >DroSim_CAF1 13251 97 - 1 U-UAGCAAGGCUGGUCACUUUAGGUUUCGAAACA--UUUUUAAUUGUUACGUUUUCUUAUUAAGCUUGA-CCCUAUAAAACUCGACCAGCCUUCAAC-CCAA .-..(.((((((((((....((((..((((((((--........))))..((((.......))))))))-.))))........)))))))))))...-.... ( -23.80) >DroEre_CAF1 22369 86 - 1 UUUAAAAUGUCUGGUCAUUUCAGGU--------------UUAAUUGUUAUGUUUUAAUAUUUAACUUGA-UGCCAUAAAUCUGGACCAGCCAUCAAC-CCAA ......(((.((((((...((((((--------------(....(((((.....)))))...)))))))-...((......)))))))).)))....-.... ( -18.50) >DroYak_CAF1 19859 97 - 1 U-CAGCAGGGCUGGUCCCUUUAAGUUUUAAAAUA--UUUUUAAUUCUUACACUUUAAUAUUUAACUUGA-GGCUAGAAGACUCGACCGGCCUUCAAG-CCAA .-..((((((((((((.(((((((((...(((((--((.................))))))))))))))-))...((....))))))))))))...)-)... ( -24.73) >consensus U_UAGCAAGGCUGGUCACUUUAGGUUUCGAAACA__UUUUUAAUUGUUACGUUUUCUUAUUAAACUUGA_CCCUAUAAAACUCGACCAGCCUUCAAC_CCAA ......((((((((((...(((((((....................................)))))))..............))))))))))......... (-14.29 = -13.97 + -0.32)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:20:48 2006