
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 5,019,246 – 5,019,394 |
| Length | 148 |
| Max. P | 0.991345 |

| Location | 5,019,246 – 5,019,356 |
|---|---|
| Length | 110 |
| Sequences | 4 |
| Columns | 114 |
| Reading direction | forward |
| Mean pairwise identity | 92.86 |
| Mean single sequence MFE | -18.99 |
| Consensus MFE | -15.68 |
| Energy contribution | -15.93 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.51 |
| Structure conservation index | 0.83 |
| SVM decision value | 1.25 |
| SVM RNA-class probability | 0.935682 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 5019246 110 + 22224390 UGCAACAUGUUCAAUGCAAAAACAAGCAUAUUUCUUCUUACUCCAA----ACUUAAAUUUCACCAUGAAUAAGACAUGAGUUUGAGUAUAAAAUAACUCACAUUUAUGCACAUU ((((..((((...((((........)))).........(((((.((----(((((...(((.....))).......))))))))))))...........))))...)))).... ( -16.90) >DroSec_CAF1 85163 110 + 1 UGCAACAUGUUCAAUGCAUAAACAAGCAUAUUUCUUCUUACUCCAA----ACUCAAAUUUCACCAUGAAUAAGACAUGAGUUUGAGUAUGAAAUAACUCACAUUUAUGCACAUU ..............((((((((......((((((....(((((.((----(((((...(((.....))).......)))))))))))).)))))).......)))))))).... ( -22.62) >DroEre_CAF1 89506 110 + 1 UGCAACAUGUUCAAUGCAUAAACAAGCAUAUUUUAUCUUACUCCGA----ACUCAAAUUUCACCAUGAAUAAGACAUGAGUUUGAGUAUGAAAUAACUCACACUUAUGCACAUU .......((((((((((........))))...............((----(.......)))....)))))).(.(((((((.(((((........))))).))))))))..... ( -20.30) >DroYak_CAF1 86582 114 + 1 UGCAACAUGUUCAACGCAUAAACAAGCAUAUUUCAUCUUACUUCGUAUACACUCAAAUUUCACCAUGAAUAAGACAUGAGUUCGAGUAUGAAAUAACUCACACUUAUGCACAUU ...............((((((.......(((((((...((((.((.....(((((...(((.....))).......))))).)))))))))))))........))))))..... ( -16.16) >consensus UGCAACAUGUUCAAUGCAUAAACAAGCAUAUUUCAUCUUACUCCAA____ACUCAAAUUUCACCAUGAAUAAGACAUGAGUUUGAGUAUGAAAUAACUCACACUUAUGCACAUU ((((.((((((..((((........)))).............................(((.....)))...))))))(((.(((((........))))).)))..)))).... (-15.68 = -15.93 + 0.25)


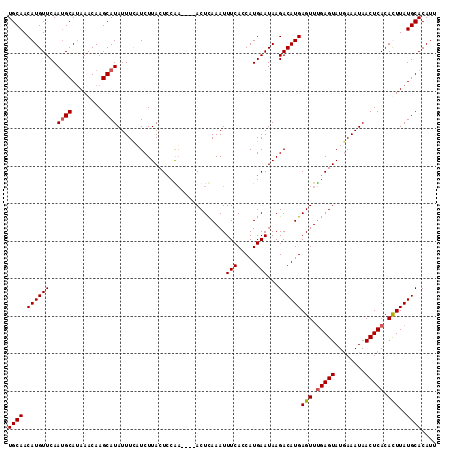
| Location | 5,019,246 – 5,019,356 |
|---|---|
| Length | 110 |
| Sequences | 4 |
| Columns | 114 |
| Reading direction | reverse |
| Mean pairwise identity | 92.86 |
| Mean single sequence MFE | -25.85 |
| Consensus MFE | -19.90 |
| Energy contribution | -19.65 |
| Covariance contribution | -0.25 |
| Combinations/Pair | 1.15 |
| Mean z-score | -1.84 |
| Structure conservation index | 0.77 |
| SVM decision value | 0.09 |
| SVM RNA-class probability | 0.580167 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 5019246 110 - 22224390 AAUGUGCAUAAAUGUGAGUUAUUUUAUACUCAAACUCAUGUCUUAUUCAUGGUGAAAUUUAAGU----UUGGAGUAAGAAGAAAUAUGCUUGUUUUUGCAUUGAACAUGUUGCA ....(((((((((.(((((........))))).((.((((.......))))))...))))).((----(..(.(((((((.((......)).))))))).)..)))....)))) ( -21.70) >DroSec_CAF1 85163 110 - 1 AAUGUGCAUAAAUGUGAGUUAUUUCAUACUCAAACUCAUGUCUUAUUCAUGGUGAAAUUUGAGU----UUGGAGUAAGAAGAAAUAUGCUUGUUUAUGCAUUGAACAUGUUGCA ...((((((((((..((((((((((.((((((((((((.((.((((.....)))).)).)))))----)).)))))....)))))).))))))))))))))............. ( -29.50) >DroEre_CAF1 89506 110 - 1 AAUGUGCAUAAGUGUGAGUUAUUUCAUACUCAAACUCAUGUCUUAUUCAUGGUGAAAUUUGAGU----UCGGAGUAAGAUAAAAUAUGCUUGUUUAUGCAUUGAACAUGUUGCA ....((((.(((((((..((((((..(((((.((((((.((.((((.....)))).)).)))))----)..)))))))))))..)))))))(((((.....)))))....)))) ( -25.50) >DroYak_CAF1 86582 114 - 1 AAUGUGCAUAAGUGUGAGUUAUUUCAUACUCGAACUCAUGUCUUAUUCAUGGUGAAAUUUGAGUGUAUACGAAGUAAGAUGAAAUAUGCUUGUUUAUGCGUUGAACAUGUUGCA ..(((((...((((((((....))))))))...(((((.((.((((.....)))).)).))))))))))....((((.(((....((((........))))....))).)))). ( -26.70) >consensus AAUGUGCAUAAAUGUGAGUUAUUUCAUACUCAAACUCAUGUCUUAUUCAUGGUGAAAUUUGAGU____UCGGAGUAAGAAGAAAUAUGCUUGUUUAUGCAUUGAACAUGUUGCA ....((((...((((...((((((((.(((((((((((((.......))))).)...))))))).....))))))))........((((........))))...))))..)))) (-19.90 = -19.65 + -0.25)



| Location | 5,019,280 – 5,019,394 |
|---|---|
| Length | 114 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 91.49 |
| Mean single sequence MFE | -28.27 |
| Consensus MFE | -25.25 |
| Energy contribution | -25.62 |
| Covariance contribution | 0.38 |
| Combinations/Pair | 1.11 |
| Mean z-score | -2.74 |
| Structure conservation index | 0.89 |
| SVM decision value | 2.26 |
| SVM RNA-class probability | 0.991345 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 5019280 114 + 22224390 UUCUUACUCCAA----ACUUAAAUUUCACCAUGAAUAAGACAUGAGUUUGAGUAUAAAAUAACUCACAUUUAUGCACAUUCAGUUGCCAGACAGUUUGAAUUG--CUCUUCGGGGUAACU .........(((----(((............(((((..(.(((((((.(((((........))))).))))))))..)))))(((....)))))))))..(((--(((....)))))).. ( -27.20) >DroSec_CAF1 85197 116 + 1 UUCUUACUCCAA----ACUCAAAUUUCACCAUGAAUAAGACAUGAGUUUGAGUAUGAAAUAACUCACAUUUAUGCACAUUCAGUUGCCAGACAGUUUGAAUUGGCUUUUUUGGGGUAACU ...(((((((((----(..............(((((..(.(((((((.(((((........))))).))))))))..)))))...(((((..........)))))...)))))))))).. ( -29.00) >DroEre_CAF1 89540 115 + 1 AUCUUACUCCGA----ACUCAAAUUUCACCAUGAAUAAGACAUGAGUUUGAGUAUGAAAUAACUCACACUUAUGCACAUUCAGUUGCCAGACAGCUUGAAUUGG-CUCUUCGGGGUAACU ...(((((((((----(.((((..((((...(((((..(.(((((((.(((((........))))).))))))))..)))))((((.....)))).))))))))-...)))))))))).. ( -33.30) >DroYak_CAF1 86616 119 + 1 AUCUUACUUCGUAUACACUCAAAUUUCACCAUGAAUAAGACAUGAGUUCGAGUAUGAAAUAACUCACACUUAUGCACAUUCAGUUGCCAGACAGUUUGAAUUGG-CUCUUCGGUGUAACU ..........((.((((((............(((((..(.(((((((..((((........))))..))))))))..)))))...(((((..........))))-).....)))))))). ( -23.60) >consensus AUCUUACUCCAA____ACUCAAAUUUCACCAUGAAUAAGACAUGAGUUUGAGUAUGAAAUAACUCACACUUAUGCACAUUCAGUUGCCAGACAGUUUGAAUUGG_CUCUUCGGGGUAACU ...(((((((((......((((..((((...(((((..(.(((((((.(((((........))))).))))))))..)))))(((....)))....)))))))).....))))))))).. (-25.25 = -25.62 + 0.38)

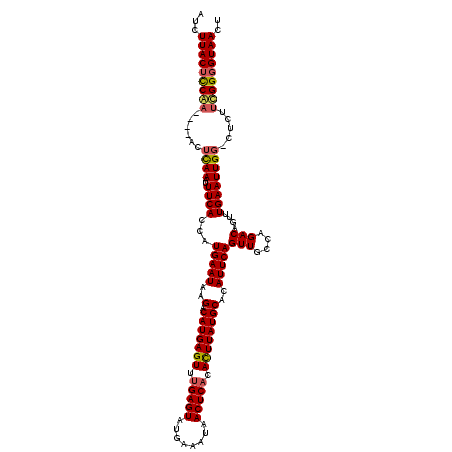
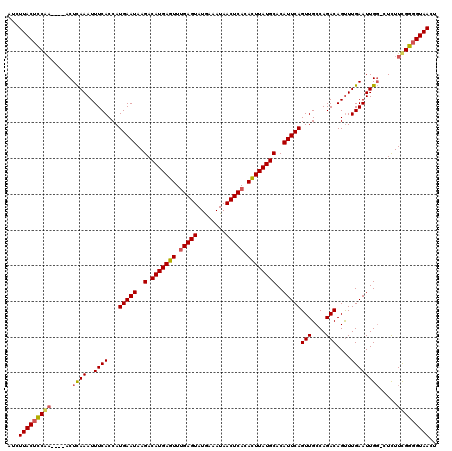
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:19:24 2006