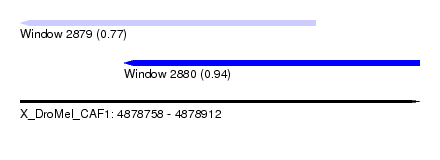
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 4,878,758 – 4,878,912 |
| Length | 154 |
| Max. P | 0.938752 |

| Location | 4,878,758 – 4,878,872 |
|---|---|
| Length | 114 |
| Sequences | 4 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 84.01 |
| Mean single sequence MFE | -18.55 |
| Consensus MFE | -12.88 |
| Energy contribution | -13.88 |
| Covariance contribution | 1.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.88 |
| Structure conservation index | 0.69 |
| SVM decision value | 0.53 |
| SVM RNA-class probability | 0.770918 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 4878758 114 - 22224390 AUAAAAA----AAAACAAACAUCUCUGCAAAAUGCGUAAAGAGUUUUUUAAUUGCCCCCCGCACUGAAAAGCUCAGAUCUCGAAAUUAAACAAAGCCCAAAACUUCAGGCUGGCAAAA .......----.....................(((.....((((((((((..(((.....))).))))))))))...................((((..........)))).)))... ( -18.60) >DroSec_CAF1 30050 106 - 1 AAAAAAAGAAAAAAAAACACAUCUCUGCAAAUUGCGUAAAGAGUUUUUUAAUUGCCCCACGCACUGAAAAGCUCAGAUCUCGAAAUUAAACAAAGCCCAAAACG------------AA ....................((((..((.....)).....((((((((((..(((.....))).))))))))))))))..........................------------.. ( -13.70) >DroEre_CAF1 32975 106 - 1 A------------AAAACACAUCUCUGCAAAUUGCGUAAAGAGUUUUUUAAUUGCCCCCCGCACUGAAAAGCUCUAAUCUCGGAAUUAAACAAAGCCCAAAACUCCAGGCUGGCAAAA .------------............(((.((((.((...(((((((((((..(((.....))).))))))))))).....)).))))......((((..........)))).)))... ( -22.10) >DroYak_CAF1 30798 106 - 1 A------------AAAACACAUCUCUGCAAAUUGCGUAAAGAGUUUUUUAAUUGCCCCCCGCACUGAAAAGCUCAGAUCUCGAAAUUAAACGAAGCCCAAAACUUCAGGCUGGCAAAA .------------..................((((.....((((((((((..(((.....))).)))))))))).....(((........)))((((..........)))).)))).. ( -19.80) >consensus A____________AAAACACAUCUCUGCAAAUUGCGUAAAGAGUUUUUUAAUUGCCCCCCGCACUGAAAAGCUCAGAUCUCGAAAUUAAACAAAGCCCAAAACUUCAGGCUGGCAAAA ..........................((.....)).....((((((((((..(((.....))).))))))))))...................((((..........))))....... (-12.88 = -13.88 + 1.00)



| Location | 4,878,798 – 4,878,912 |
|---|---|
| Length | 114 |
| Sequences | 4 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 85.69 |
| Mean single sequence MFE | -13.65 |
| Consensus MFE | -12.77 |
| Energy contribution | -12.78 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.85 |
| Structure conservation index | 0.94 |
| SVM decision value | 1.27 |
| SVM RNA-class probability | 0.938752 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
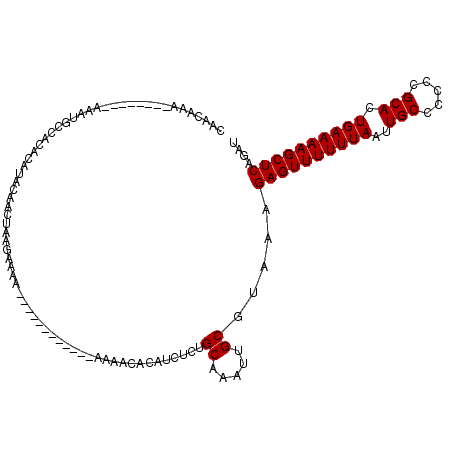
>X_DroMel_CAF1 4878798 114 - 22224390 CAACAAAAAAAAAAAAAAUGCCUCACAUACAACUAAGAAAAUAAAAA----AAAACAAACAUCUCUGCAAAAUGCGUAAAGAGUUUUUUAAUUGCCCCCCGCACUGAAAAGCUCAGAU ...............................................----.........((((..((.....)).....((((((((((..(((.....))).)))))))))))))) ( -13.60) >DroSec_CAF1 30078 110 - 1 CAACAAA--------AAAUGCCUCACAUACAACUAAGAAAAAAAAAAGAAAAAAAAACACAUCUCUGCAAAUUGCGUAAAGAGUUUUUUAAUUGCCCCACGCACUGAAAAGCUCAGAU .......--------.............................................((((..((.....)).....((((((((((..(((.....))).)))))))))))))) ( -13.60) >DroEre_CAF1 33015 98 - 1 CAACAAA--------AAAUGCCACACAUACAACUAGGAAAA------------AAAACACAUCUCUGCAAAUUGCGUAAAGAGUUUUUUAAUUGCCCCCCGCACUGAAAAGCUCUAAU .......--------............(((.....(((...------------..........)))((.....))))).(((((((((((..(((.....))).)))))))))))... ( -13.82) >DroYak_CAF1 30838 98 - 1 CAACAAA--------AAAUGCCACACAUACAACUAGGAAAA------------AAAACACAUCUCUGCAAAUUGCGUAAAGAGUUUUUUAAUUGCCCCCCGCACUGAAAAGCUCAGAU .......--------..........................------------.......((((..((.....)).....((((((((((..(((.....))).)))))))))))))) ( -13.60) >consensus CAACAAA________AAAUGCCACACAUACAACUAAGAAAA____________AAAACACAUCUCUGCAAAUUGCGUAAAGAGUUUUUUAAUUGCCCCCCGCACUGAAAAGCUCAGAU ..................................................................((.....)).....((((((((((..(((.....))).)))))))))).... (-12.77 = -12.78 + 0.00)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:18:17 2006