| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 4,863,597 – 4,863,692 |
| Length | 95 |
| Max. P | 0.999665 |

| Location | 4,863,597 – 4,863,692 |
|---|---|
| Length | 95 |
| Sequences | 5 |
| Columns | 95 |
| Reading direction | forward |
| Mean pairwise identity | 87.37 |
| Mean single sequence MFE | -21.28 |
| Consensus MFE | -18.16 |
| Energy contribution | -18.00 |
| Covariance contribution | -0.16 |
| Combinations/Pair | 1.19 |
| Mean z-score | -2.66 |
| Structure conservation index | 0.85 |
| SVM decision value | 2.87 |
| SVM RNA-class probability | 0.997483 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 4863597 95 + 22224390 AACUAAAAACAAAAAAAAAAAAAACCACUUGCAAUCCAAUUCGGACGGGAAGGAACCAUUUCAAUCGGGCCGCAAGUGGAUUUGAAUUGAGAUCU ........................((((((((..(((.(((..((.((.......)).))..))).)))..))))))))................ ( -20.30) >DroSec_CAF1 15352 85 + 1 AACCAAA----------AAAGAAACCACUUGCAAUCCAAUCCGGACGGAAAGGAACCAUUUCAAUCGGGCCGCAAGUGGAUUUGAAUUGAGAUCU ...(((.----------...(((.((((((((..(((..(((....)))..))).((.........))...)))))))).)))...)))...... ( -22.60) >DroSim_CAF1 15339 85 + 1 AACCAAA----------AAAGAAACCACUUGCAAUCCAAUCGGGACGGAAAGGAACCAUUUCAAUCGGGCCGCAAGUGGAUUUGAAUUGAGAUCU ...(((.----------...(((.((((((((..(((.((.(..(.((.......)).)..).)).)))..)))))))).)))...)))...... ( -21.10) >DroEre_CAF1 18309 80 + 1 AACUAAA---------------AACCACUUGCAAUCCAACCCAGACGGGAAGGAACCAUUUCAAUCAGGUCGCAAGUGGAUUUGAAUUGAGAUCU .......---------------..((((((((..(((..(((....)))..)))(((..........))).))))))))................ ( -23.80) >DroYak_CAF1 15540 82 + 1 AACAAAA----------A---AAAACACUUGCAAUCCAAUCCGGACGGGAAGGAACCAUUUCAAUCAGGCCGCAAGUGGAUUUGAAUUGAGAUCU ..(((..----------(---((..(((((((..(((..(((....)))..))).((..........))..)))))))..)))...)))...... ( -18.60) >consensus AACCAAA__________AAA_AAACCACUUGCAAUCCAAUCCGGACGGGAAGGAACCAUUUCAAUCGGGCCGCAAGUGGAUUUGAAUUGAGAUCU ........................((((((((..(((..(((....)))..))).((..........))..))))))))................ (-18.16 = -18.00 + -0.16)



| Location | 4,863,597 – 4,863,692 |
|---|---|
| Length | 95 |
| Sequences | 5 |
| Columns | 95 |
| Reading direction | reverse |
| Mean pairwise identity | 87.37 |
| Mean single sequence MFE | -24.74 |
| Consensus MFE | -22.44 |
| Energy contribution | -21.92 |
| Covariance contribution | -0.52 |
| Combinations/Pair | 1.20 |
| Mean z-score | -2.90 |
| Structure conservation index | 0.91 |
| SVM decision value | 3.86 |
| SVM RNA-class probability | 0.999665 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 4863597 95 - 22224390 AGAUCUCAAUUCAAAUCCACUUGCGGCCCGAUUGAAAUGGUUCCUUCCCGUCCGAAUUGGAUUGCAAGUGGUUUUUUUUUUUUUUGUUUUUAGUU ....((.((..((((.(((((((((..((((((...((((.......))))...))))))..)))))))))...........))))..)).)).. ( -23.80) >DroSec_CAF1 15352 85 - 1 AGAUCUCAAUUCAAAUCCACUUGCGGCCCGAUUGAAAUGGUUCCUUUCCGUCCGGAUUGGAUUGCAAGUGGUUUCUUU----------UUUGGUU .....((((.......(((((((((..((((((...((((.......))))...))))))..))))))))).......----------.)))).. ( -25.76) >DroSim_CAF1 15339 85 - 1 AGAUCUCAAUUCAAAUCCACUUGCGGCCCGAUUGAAAUGGUUCCUUUCCGUCCCGAUUGGAUUGCAAGUGGUUUCUUU----------UUUGGUU .....((((.......(((((((((..(((((((..((((.......))))..)))))))..))))))))).......----------.)))).. ( -29.16) >DroEre_CAF1 18309 80 - 1 AGAUCUCAAUUCAAAUCCACUUGCGACCUGAUUGAAAUGGUUCCUUCCCGUCUGGGUUGGAUUGCAAGUGGUU---------------UUUAGUU .......((((.(((.((((((((((.(..(((.(.((((.......)))).).)))..).)))))))))).)---------------)).)))) ( -24.80) >DroYak_CAF1 15540 82 - 1 AGAUCUCAAUUCAAAUCCACUUGCGGCCUGAUUGAAAUGGUUCCUUCCCGUCCGGAUUGGAUUGCAAGUGUUUU---U----------UUUUGUU ......(((...(((..((((((((..(..(((...((((.......))))...)))..)..))))))))..))---)----------..))).. ( -20.20) >consensus AGAUCUCAAUUCAAAUCCACUUGCGGCCCGAUUGAAAUGGUUCCUUCCCGUCCGGAUUGGAUUGCAAGUGGUUU_UUU__________UUUAGUU ................((((((((((.((((((...((((.......))))...)))))).))))))))))........................ (-22.44 = -21.92 + -0.52)


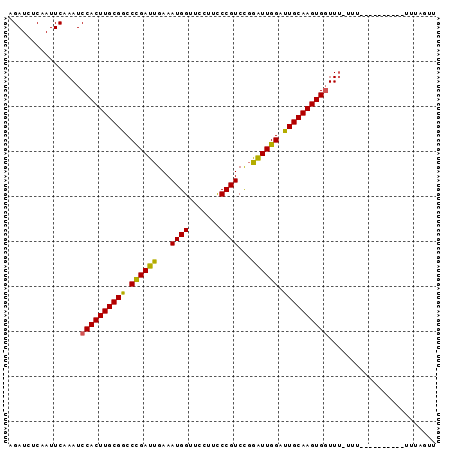
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:18:05 2006