
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 4,780,178 – 4,780,286 |
| Length | 108 |
| Max. P | 0.919304 |

| Location | 4,780,178 – 4,780,286 |
|---|---|
| Length | 108 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 78.43 |
| Mean single sequence MFE | -26.13 |
| Consensus MFE | -19.24 |
| Energy contribution | -20.13 |
| Covariance contribution | 0.89 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.84 |
| Structure conservation index | 0.74 |
| SVM decision value | 1.13 |
| SVM RNA-class probability | 0.919304 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 4780178 108 + 22224390 GAGGUGGUGGAGA---------AACAUCGAUGCCAAACAUCGAUAUUUUCGAAAAAUCACAUAAAUUGGUG---UUUUUUGAUGAAUAUUCAUUUUUGAUGUAUAUUCAAAUUUCUGAUG (((((..((((..---------.(((((((((.....))))(((((((((((((((.(((........)))---)))))))).))))))).......)))))...)))).)))))..... ( -25.70) >DroSec_CAF1 19184 117 + 1 GAGGUGGAGAAAAAUUGAUGCCAACAUCGAUGUCAAACAUCGAUAUUUUCGAAGAAUCACAAACAUCGGUG---UUUUUCGAGGAAUCAUCGUAUAAAUUGUAAGUACAAAUUAAUUUUG ((((((..((...(((((((....))))))).))...))))(((.(((((((((((.(((........)))---)))))))))))))).))...(((.((((....)))).)))...... ( -26.80) >DroSim_CAF1 13919 120 + 1 GAGGUGGAGAAAAAUUGAUGCCAACAUCGAUGUCAAACAUCGAUAUUUUCGAAGAAUCACACACAUCGGUUUGUUUUUUCGAGGAAUCAUCGUAUAAAUUGUAAGUACAAAUUAAUUUUG ((((((..((...(((((((....))))))).))...))))(((.(((((((((((.((.((......)).)).)))))))))))))).))...(((.((((....)))).)))...... ( -25.90) >consensus GAGGUGGAGAAAAAUUGAUGCCAACAUCGAUGUCAAACAUCGAUAUUUUCGAAGAAUCACAAACAUCGGUG___UUUUUCGAGGAAUCAUCGUAUAAAUUGUAAGUACAAAUUAAUUUUG ..(((((..................(((((((.....))))))).(((((((((((.(((........)))...))))))))))).)))))............................. (-19.24 = -20.13 + 0.89)


| Location | 4,780,178 – 4,780,286 |
|---|---|
| Length | 108 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 78.43 |
| Mean single sequence MFE | -22.73 |
| Consensus MFE | -12.53 |
| Energy contribution | -13.20 |
| Covariance contribution | 0.67 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.40 |
| Structure conservation index | 0.55 |
| SVM decision value | 0.58 |
| SVM RNA-class probability | 0.790103 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 4780178 108 - 22224390 CAUCAGAAAUUUGAAUAUACAUCAAAAAUGAAUAUUCAUCAAAAAA---CACCAAUUUAUGUGAUUUUUCGAAAAUAUCGAUGUUUGGCAUCGAUGUU---------UCUCCACCACCUC .....((((..(((((((.(((.....))).)))))))........---(((........)))...))))(((((((((((((.....))))))))))---------).))......... ( -21.70) >DroSec_CAF1 19184 117 - 1 CAAAAUUAAUUUGUACUUACAAUUUAUACGAUGAUUCCUCGAAAAA---CACCGAUGUUUGUGAUUCUUCGAAAAUAUCGAUGUUUGACAUCGAUGUUGGCAUCAAUUUUUCUCCACCUC ......(((.((((....)))).))).....((((.(((((((.((---(((........))).)).))))).((((((((((.....)))))))))))).))))............... ( -23.60) >DroSim_CAF1 13919 120 - 1 CAAAAUUAAUUUGUACUUACAAUUUAUACGAUGAUUCCUCGAAAAAACAAACCGAUGUGUGUGAUUCUUCGAAAAUAUCGAUGUUUGACAUCGAUGUUGGCAUCAAUUUUUCUCCACCUC ......(((.((((....)))).))).....((((.(((((((...(((.((....)).))).....))))).((((((((((.....)))))))))))).))))............... ( -22.90) >consensus CAAAAUUAAUUUGUACUUACAAUUUAUACGAUGAUUCCUCGAAAAA___CACCGAUGUAUGUGAUUCUUCGAAAAUAUCGAUGUUUGACAUCGAUGUUGGCAUCAAUUUUUCUCCACCUC ......................................(((((.((...(((........))).)).))))).((((((((((.....))))))))))...................... (-12.53 = -13.20 + 0.67)

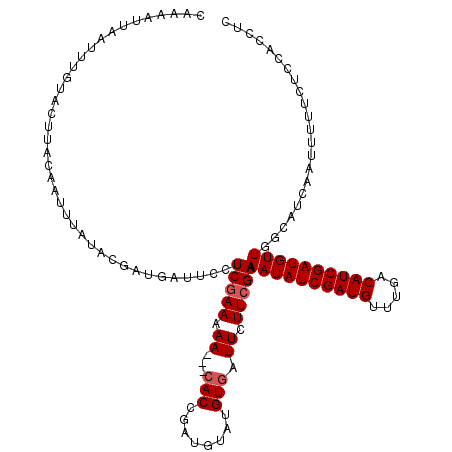

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:17:19 2006