
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 4,728,456 – 4,728,552 |
| Length | 96 |
| Max. P | 0.802859 |

| Location | 4,728,456 – 4,728,552 |
|---|---|
| Length | 96 |
| Sequences | 3 |
| Columns | 110 |
| Reading direction | forward |
| Mean pairwise identity | 79.81 |
| Mean single sequence MFE | -9.09 |
| Consensus MFE | -7.20 |
| Energy contribution | -7.20 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.61 |
| Structure conservation index | 0.79 |
| SVM decision value | 0.62 |
| SVM RNA-class probability | 0.802859 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 4728456 96 + 22224390 CUUUCUCCACAUUUUGCAGCUUUACAGCUCGCCACAAUGAAACUUUUUAGUUGAUUUUAUUAUUUUUUCCCCACUUC-----------CAUUCAAUUU---UUGUUUUUU .........((((..((((((....)))).))...))))((((.....((((((.......................-----------...)))))).---..))))... ( -8.87) >DroSec_CAF1 21484 89 + 1 CUUUCUCGACAUUUUGCAGCUUUGCAGCUCGCCACAAUGAAACUUUUUAGUUUAUUUUAUUAUUUUUUCCCCACUUC-----------CAUUCA----------UUUUUU .........((((..((((((....)))).))...))))(((((....)))))........................-----------......----------...... ( -8.40) >DroEre_CAF1 24815 105 + 1 CUUUUUCGACAUUUUGCAGCUUUACAGCUCGCCACAAUGAAACUUUUUAGUUGAUUUUAUUAUUUUUUCCCCACUUUCCCUUACAUUCGGUUUG----GUAUUUUUUUU- ((...((((((((..((((((....)))).))...)))).((((....))))..................................))))...)----)..........- ( -10.00) >consensus CUUUCUCGACAUUUUGCAGCUUUACAGCUCGCCACAAUGAAACUUUUUAGUUGAUUUUAUUAUUUUUUCCCCACUUC___________CAUUCA_______UU_UUUUUU .........((((..((((((....)))).))...)))).((((....)))).......................................................... ( -7.20 = -7.20 + 0.00)

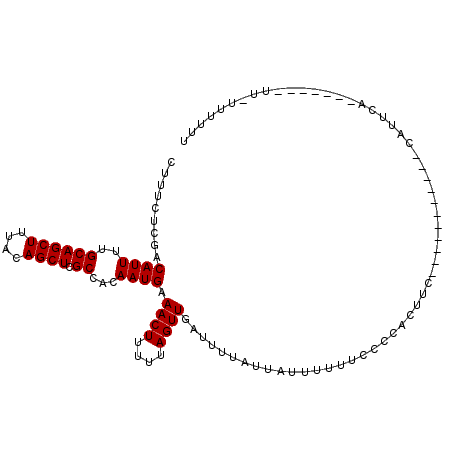
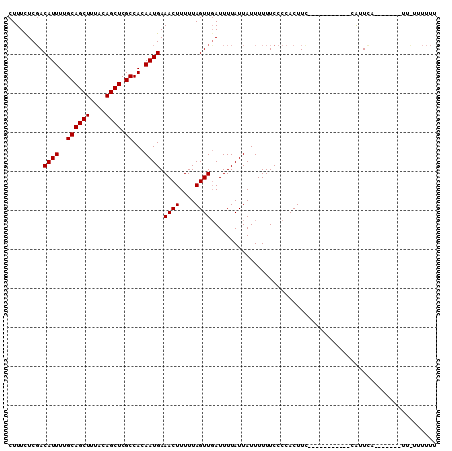
| Location | 4,728,456 – 4,728,552 |
|---|---|
| Length | 96 |
| Sequences | 3 |
| Columns | 110 |
| Reading direction | reverse |
| Mean pairwise identity | 79.81 |
| Mean single sequence MFE | -12.09 |
| Consensus MFE | -9.10 |
| Energy contribution | -9.10 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.44 |
| Structure conservation index | 0.75 |
| SVM decision value | 0.14 |
| SVM RNA-class probability | 0.601492 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 4728456 96 - 22224390 AAAAAACAA---AAAUUGAAUG-----------GAAGUGGGGAAAAAAUAAUAAAAUCAACUAAAAAGUUUCAUUGUGGCGAGCUGUAAAGCUGCAAAAUGUGGAGAAAG .....(((.---......((((-----------(((.(.((...................))....).)))))))...((.((((....))))))....)))........ ( -11.21) >DroSec_CAF1 21484 89 - 1 AAAAAA----------UGAAUG-----------GAAGUGGGGAAAAAAUAAUAAAAUAAACUAAAAAGUUUCAUUGUGGCGAGCUGCAAAGCUGCAAAAUGUCGAGAAAG ....((----------((((..-----------....(((....................)))......))))))...((.((((....))))))............... ( -10.65) >DroEre_CAF1 24815 105 - 1 -AAAAAAAAUAC----CAAACCGAAUGUAAGGGAAAGUGGGGAAAAAAUAAUAAAAUCAACUAAAAAGUUUCAUUGUGGCGAGCUGUAAAGCUGCAAAAUGUCGAAAAAG -..........(----((..((........)).....)))................((((((....)))).((((...((.((((....))))))..))))..))..... ( -14.40) >consensus AAAAAA_AA_______UGAAUG___________GAAGUGGGGAAAAAAUAAUAAAAUCAACUAAAAAGUUUCAUUGUGGCGAGCUGUAAAGCUGCAAAAUGUCGAGAAAG .......................................................................((((...((.((((....))))))..))))......... ( -9.10 = -9.10 + -0.00)


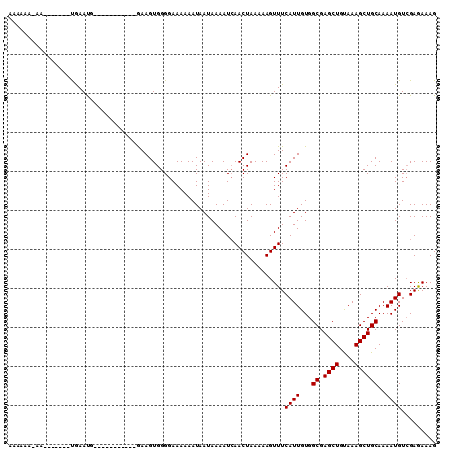
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:16:51 2006