
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 4,699,075 – 4,699,208 |
| Length | 133 |
| Max. P | 0.925769 |

| Location | 4,699,075 – 4,699,178 |
|---|---|
| Length | 103 |
| Sequences | 4 |
| Columns | 103 |
| Reading direction | forward |
| Mean pairwise identity | 94.50 |
| Mean single sequence MFE | -29.30 |
| Consensus MFE | -24.74 |
| Energy contribution | -25.55 |
| Covariance contribution | 0.81 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.54 |
| Structure conservation index | 0.84 |
| SVM decision value | 1.17 |
| SVM RNA-class probability | 0.925769 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
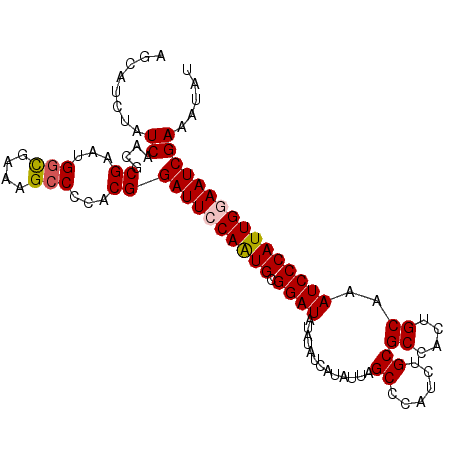
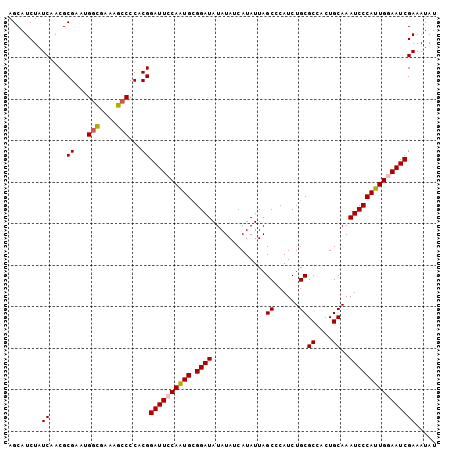
>X_DroMel_CAF1 4699075 103 + 22224390 AGCAUCUAUCAACGCGAAUGGCGAAAGCCCCACGGAUUCCAGUGCGGAUAUAUAUCAUAUUAGCCCAUCUGCGCCACUGCAAAUCCCAUUGGAAUCGAAAUAU ........((....((...(((....)))...))((((((((((.((((.............((......))((....))..))))))))))))))))..... ( -30.10) >DroSec_CAF1 52272 103 + 1 AGCAUCUAUCAACGCGAAUGGCGAAAGCCCCACGGAUUCCAGUGCGGAUAUAUAUCAUAUUAGCCCAUCUGCGCCACUGCAAAUCCCAUUGGAAUCGAAAUAU ........((....((...(((....)))...))((((((((((.((((.............((......))((....))..))))))))))))))))..... ( -30.10) >DroEre_CAF1 53593 103 + 1 AGCAUCUAUCAACGCGUGAGGCGAAAGCCCCACGGAUUGCAAUGCGGAUAUAUAUCAUAUUAGCCCAUCUGCGCCACUGCAAAUCCCAUUGGAAUCGAAAUAU ........((....((((.(((....))).))))((((.(((((.((((.............((......))((....))..))))))))).))))))..... ( -31.40) >DroYak_CAF1 50372 103 + 1 AGCAUCUAUCGACGCGGAUGUUGAAAGCCCCACGGAUUGCAAUGGGGAUAUAUAUCAUAUUAGCCCAUCUGCGCUACUGCAAAUCCCAUUGGAAUCGAAAUAU (((((((........)))))))............((((.((((((((...............((......))((....))...)))))))).))))....... ( -25.60) >consensus AGCAUCUAUCAACGCGAAUGGCGAAAGCCCCACGGAUUCCAAUGCGGAUAUAUAUCAUAUUAGCCCAUCUGCGCCACUGCAAAUCCCAUUGGAAUCGAAAUAU ........((....((...(((....)))...))((((((((((.((((.............((......))((....))..))))))))))))))))..... (-24.74 = -25.55 + 0.81)

| Location | 4,699,114 – 4,699,208 |
|---|---|
| Length | 94 |
| Sequences | 4 |
| Columns | 100 |
| Reading direction | forward |
| Mean pairwise identity | 89.21 |
| Mean single sequence MFE | -17.10 |
| Consensus MFE | -13.45 |
| Energy contribution | -13.70 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.56 |
| Structure conservation index | 0.79 |
| SVM decision value | 0.20 |
| SVM RNA-class probability | 0.631469 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
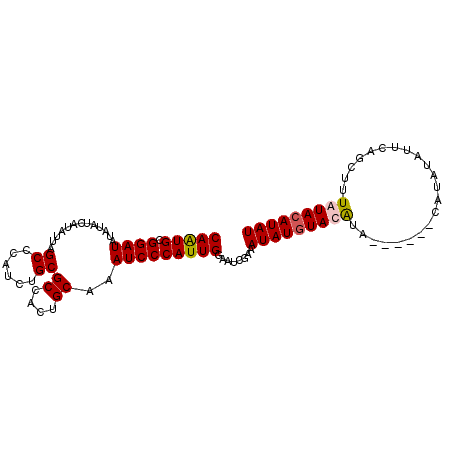
>X_DroMel_CAF1 4699114 94 + 22224390 CAGUGCGGAUAUAUAUCAUAUUAGCCCAUCUGCGCCACUGCAAAUCCCAUUGGAAUCGAAAUAUGUAUGUA------CAUAUAUUUAGCUUUAUACAUAU (((((.((((.............((......))((....))..)))))))))........(((((((((..------..((....))....))))))))) ( -16.90) >DroSec_CAF1 52311 94 + 1 CAGUGCGGAUAUAUAUCAUAUUAGCCCAUCUGCGCCACUGCAAAUCCCAUUGGAAUCGAAAUAUGUAUGCA------CAUAUAUUCAGCUUUAUACAUAU ..((((((((...((......))....))))))))....((...(((....)))...(((((((((....)------))))).))).))........... ( -17.60) >DroEre_CAF1 53632 90 + 1 CAAUGCGGAUAUAUAUCAUAUUAGCCCAUCUGCGCCACUGCAAAUCCCAUUGGAAUCGAAAUAUAUACAUAU----------AUUCAGCUUUAUACAUAU (((((.((((.............((......))((....))..)))))))))(((.(..((((((....)))----------)))..).)))........ ( -12.50) >DroYak_CAF1 50411 100 + 1 CAAUGGGGAUAUAUAUCAUAUUAGCCCAUCUGCGCUACUGCAAAUCCCAUUGGAAUCGAAAUAUGUACAUAUAUUCGUAUAUAUUCAGCUUUAUACAUAU ((((((((...............((......))((....))...))))))))(((.(..(((((((((........)))))))))..).)))........ ( -21.40) >consensus CAAUGCGGAUAUAUAUCAUAUUAGCCCAUCUGCGCCACUGCAAAUCCCAUUGGAAUCGAAAUAUGUACAUA______CAUAUAUUCAGCUUUAUACAUAU (((((.((((.............((......))((....))..)))))))))........(((((((((......................))))))))) (-13.45 = -13.70 + 0.25)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:16:27 2006