
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 4,586,473 – 4,586,620 |
| Length | 147 |
| Max. P | 0.999809 |

| Location | 4,586,473 – 4,586,580 |
|---|---|
| Length | 107 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 83.11 |
| Mean single sequence MFE | -27.32 |
| Consensus MFE | -15.60 |
| Energy contribution | -15.60 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.10 |
| Structure conservation index | 0.57 |
| SVM decision value | 0.14 |
| SVM RNA-class probability | 0.603785 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 4586473 107 + 22224390 CUUUAUCCCGACGCUGG-GGUCAUGCAAAUUUUAUGAAUAUCUGCCAAAAUGGGAAAUAAGCAAAUAAGAACAUUUCCCGCUUGGCUUUACGUCACA------------GCACGCAGCUU .....(((((....)))-))(((((.......)))))....((((......(((((((...(......)...)))))))(((((((.....)))).)------------))..))))... ( -25.40) >DroSec_CAF1 10602 106 + 1 CUUUACCCCGUCGCUGU-GGACAUGCAAAUUUUAUG-AUAUCUGCCAAAAUGGGAAAUAAGCAAAUAAGAACAUUUCCCGCUUGGCUUUACAUCACA------------GCACGCAGCUU ........(((.(((((-((.((((.......))))-......(((((..((((((((...(......)...)))))))).)))))......)))))------------)))))...... ( -28.00) >DroSim_CAF1 12049 107 + 1 CUUUACCCCGUCGCUGU-GGCCAUGCAAAUUUUAUGAAUAUCUGCCAAAAUGGGAAAUAAGCAAAUAAGAACAUUUCCCGCUUGGCUUUACAUCACA------------GCACGCAGCUU ........(((.(((((-((.((((.......)))).......(((((..((((((((...(......)...)))))))).)))))......)))))------------)))))...... ( -27.50) >DroEre_CAF1 13582 98 + 1 CUUC---------------GCCAUGCAAAUUUUAUGAAUGUCUGCCAAAAUGGGAAAUAAGCAAAUAAGAACAUUUCCCGCUUGGCUUUACAUCGCAGCAC-------GGCACGCAACUU ....---------------(((.(((........((.((((..(((((..((((((((...(......)...)))))))).)))))...))))..))))).-------)))......... ( -24.40) >DroYak_CAF1 10945 120 + 1 CUUUUCGCAGUUGCGCCGAGCCAUGCAAAUUUUAUGAAUAUCUGCCAAAAUGGGAAAUAAGCAAAUAAGAACAUUUCCCGCUUGGCCUUACAGCACAGCACAGCGCACAGCACGCAACUU ........(((((((((((((...(((.((.........)).)))......(((((((...(......)...))))))))))))))......((...((.....))...))..)))))). ( -31.30) >consensus CUUUACCCCGUCGCUGU_GGCCAUGCAAAUUUUAUGAAUAUCUGCCAAAAUGGGAAAUAAGCAAAUAAGAACAUUUCCCGCUUGGCUUUACAUCACA____________GCACGCAGCUU .......................(((.........(.....).(((((..((((((((...(......)...)))))))).)))))...........................))).... (-15.60 = -15.60 + -0.00)


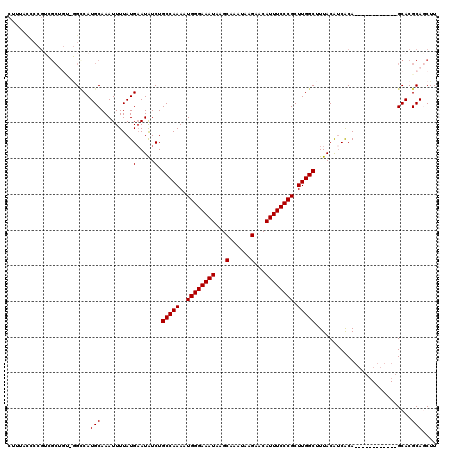
| Location | 4,586,473 – 4,586,580 |
|---|---|
| Length | 107 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 83.11 |
| Mean single sequence MFE | -36.30 |
| Consensus MFE | -22.39 |
| Energy contribution | -22.83 |
| Covariance contribution | 0.44 |
| Combinations/Pair | 1.04 |
| Mean z-score | -3.06 |
| Structure conservation index | 0.62 |
| SVM decision value | 1.77 |
| SVM RNA-class probability | 0.976311 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 4586473 107 - 22224390 AAGCUGCGUGC------------UGUGACGUAAAGCCAAGCGGGAAAUGUUCUUAUUUGCUUAUUUCCCAUUUUGGCAGAUAUUCAUAAAAUUUGCAUGACC-CCAGCGUCGGGAUAAAG ..(((((((((------------(((((......((((((.((((((((..(......)..))))))))..))))))......)))))......)))))...-.))))............ ( -28.80) >DroSec_CAF1 10602 106 - 1 AAGCUGCGUGC------------UGUGAUGUAAAGCCAAGCGGGAAAUGUUCUUAUUUGCUUAUUUCCCAUUUUGGCAGAUAU-CAUAAAAUUUGCAUGUCC-ACAGCGACGGGGUAAAG ..(((.(((((------------((((.......((((((.((((((((..(......)..))))))))..)))))).(((((-((.......)).))))))-))))).))).))).... ( -38.40) >DroSim_CAF1 12049 107 - 1 AAGCUGCGUGC------------UGUGAUGUAAAGCCAAGCGGGAAAUGUUCUUAUUUGCUUAUUUCCCAUUUUGGCAGAUAUUCAUAAAAUUUGCAUGGCC-ACAGCGACGGGGUAAAG ..(((.(((((------------(((((((((((((((((.((((((((..(......)..))))))))..)))))).(.....)......)))))))...)-))))).))).))).... ( -38.00) >DroEre_CAF1 13582 98 - 1 AAGUUGCGUGCC-------GUGCUGCGAUGUAAAGCCAAGCGGGAAAUGUUCUUAUUUGCUUAUUUCCCAUUUUGGCAGACAUUCAUAAAAUUUGCAUGGC---------------GAAG ........((((-------((((((.(((((...((((((.((((((((..(......)..))))))))..))))))..)))))))........)))))))---------------)... ( -32.30) >DroYak_CAF1 10945 120 - 1 AAGUUGCGUGCUGUGCGCUGUGCUGUGCUGUAAGGCCAAGCGGGAAAUGUUCUUAUUUGCUUAUUUCCCAUUUUGGCAGAUAUUCAUAAAAUUUGCAUGGCUCGGCGCAACUGCGAAAAG ........(((..(((((((.(((((((......((((((.((((((((..(......)..))))))))..)))))).(.....).........))))))).)))))))...)))..... ( -44.00) >consensus AAGCUGCGUGC____________UGUGAUGUAAAGCCAAGCGGGAAAUGUUCUUAUUUGCUUAUUUCCCAUUUUGGCAGAUAUUCAUAAAAUUUGCAUGGCC_ACAGCGACGGGGUAAAG ......(((((............((((((((...((((((.((((((((..(......)..))))))))..))))))..)))).))))......)))))..................... (-22.39 = -22.83 + 0.44)



| Location | 4,586,512 – 4,586,620 |
|---|---|
| Length | 108 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 91.69 |
| Mean single sequence MFE | -29.24 |
| Consensus MFE | -28.12 |
| Energy contribution | -28.12 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.27 |
| Structure conservation index | 0.96 |
| SVM decision value | 4.13 |
| SVM RNA-class probability | 0.999809 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 4586512 108 + 22224390 UCUGCCAAAAUGGGAAAUAAGCAAAUAAGAACAUUUCCCGCUUGGCUUUACGUCACA------------GCACGCAGCUUGUGCAGCUCGCAAAUUCAUAGUGGGCAACUUUAAAUUUAA ...(((((..((((((((...(......)...)))))))).)))))...........------------(((((.....))))).((((((.........)))))).............. ( -28.40) >DroSec_CAF1 10640 108 + 1 UCUGCCAAAAUGGGAAAUAAGCAAAUAAGAACAUUUCCCGCUUGGCUUUACAUCACA------------GCACGCAGCUUGUGCAGCUCGCAAAUUCAUGGUGGGCAACUUUAAAUUUAA ...(((((..((((((((...(......)...)))))))).)))))...........------------(((((.....))))).((((((.........)))))).............. ( -28.30) >DroSim_CAF1 12088 108 + 1 UCUGCCAAAAUGGGAAAUAAGCAAAUAAGAACAUUUCCCGCUUGGCUUUACAUCACA------------GCACGCAGCUUGUGCAGCUCGCAAAUUCAUGGUGGGCAACUUUAAAUUUAA ...(((((..((((((((...(......)...)))))))).)))))...........------------(((((.....))))).((((((.........)))))).............. ( -28.30) >DroEre_CAF1 13607 113 + 1 UCUGCCAAAAUGGGAAAUAAGCAAAUAAGAACAUUUCCCGCUUGGCUUUACAUCGCAGCAC-------GGCACGCAACUUGUGCAGCUCGCAAAUUCAUAGUGGGCAACUUUAAAUUUCA ...(((((..((((((((...(......)...)))))))).)))))...........((((-------((........)))))).((((((.........)))))).............. ( -28.90) >DroYak_CAF1 10985 120 + 1 UCUGCCAAAAUGGGAAAUAAGCAAAUAAGAACAUUUCCCGCUUGGCCUUACAGCACAGCACAGCGCACAGCACGCAACUUGUGCAGCUCGCAAAUUCAUAGUGGGCAACUUUAAAUUCAA ...(((((..((((((((...(......)...)))))))).)))))......((((((....(((.......)))...)))))).((((((.........)))))).............. ( -32.30) >consensus UCUGCCAAAAUGGGAAAUAAGCAAAUAAGAACAUUUCCCGCUUGGCUUUACAUCACA____________GCACGCAGCUUGUGCAGCUCGCAAAUUCAUAGUGGGCAACUUUAAAUUUAA ...(((((..((((((((...(......)...)))))))).))))).......................(((((.....))))).((((((.........)))))).............. (-28.12 = -28.12 + -0.00)



| Location | 4,586,512 – 4,586,620 |
|---|---|
| Length | 108 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 91.69 |
| Mean single sequence MFE | -31.14 |
| Consensus MFE | -27.08 |
| Energy contribution | -27.08 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.46 |
| Structure conservation index | 0.87 |
| SVM decision value | 0.79 |
| SVM RNA-class probability | 0.850353 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 4586512 108 - 22224390 UUAAAUUUAAAGUUGCCCACUAUGAAUUUGCGAGCUGCACAAGCUGCGUGC------------UGUGACGUAAAGCCAAGCGGGAAAUGUUCUUAUUUGCUUAUUUCCCAUUUUGGCAGA .....((((.(((.....))).))))((((((.((.((((.......))))------------.))..))))))((((((.((((((((..(......)..))))))))..))))))... ( -30.50) >DroSec_CAF1 10640 108 - 1 UUAAAUUUAAAGUUGCCCACCAUGAAUUUGCGAGCUGCACAAGCUGCGUGC------------UGUGAUGUAAAGCCAAGCGGGAAAUGUUCUUAUUUGCUUAUUUCCCAUUUUGGCAGA ............(((((.....((..((((((.((.((((.......))))------------.))..))))))..))...((((((((..(......)..)))))))).....))))). ( -28.50) >DroSim_CAF1 12088 108 - 1 UUAAAUUUAAAGUUGCCCACCAUGAAUUUGCGAGCUGCACAAGCUGCGUGC------------UGUGAUGUAAAGCCAAGCGGGAAAUGUUCUUAUUUGCUUAUUUCCCAUUUUGGCAGA ............(((((.....((..((((((.((.((((.......))))------------.))..))))))..))...((((((((..(......)..)))))))).....))))). ( -28.50) >DroEre_CAF1 13607 113 - 1 UGAAAUUUAAAGUUGCCCACUAUGAAUUUGCGAGCUGCACAAGUUGCGUGCC-------GUGCUGCGAUGUAAAGCCAAGCGGGAAAUGUUCUUAUUUGCUUAUUUCCCAUUUUGGCAGA ..(((((((.(((.....))).)))))))((.(((.((((.......)))).-------..)))))........((((((.((((((((..(......)..))))))))..))))))... ( -31.90) >DroYak_CAF1 10985 120 - 1 UUGAAUUUAAAGUUGCCCACUAUGAAUUUGCGAGCUGCACAAGUUGCGUGCUGUGCGCUGUGCUGUGCUGUAAGGCCAAGCGGGAAAUGUUCUUAUUUGCUUAUUUCCCAUUUUGGCAGA ..(((((((.(((.....))).)))))))....((.(((((.((.((((.....))))...))))))).))...((((((.((((((((..(......)..))))))))..))))))... ( -36.30) >consensus UUAAAUUUAAAGUUGCCCACUAUGAAUUUGCGAGCUGCACAAGCUGCGUGC____________UGUGAUGUAAAGCCAAGCGGGAAAUGUUCUUAUUUGCUUAUUUCCCAUUUUGGCAGA ..(((((((.(((.....))).)))))))(((((((.....)))).))).........................((((((.((((((((..(......)..))))))))..))))))... (-27.08 = -27.08 + 0.00)

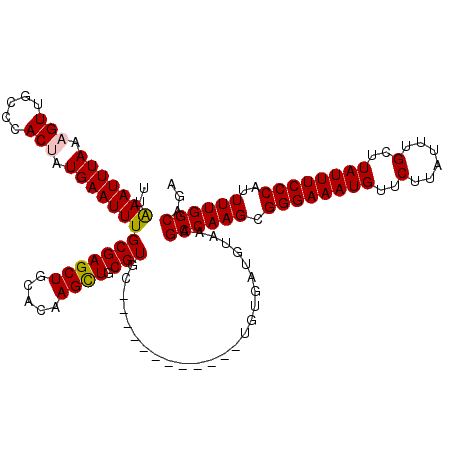
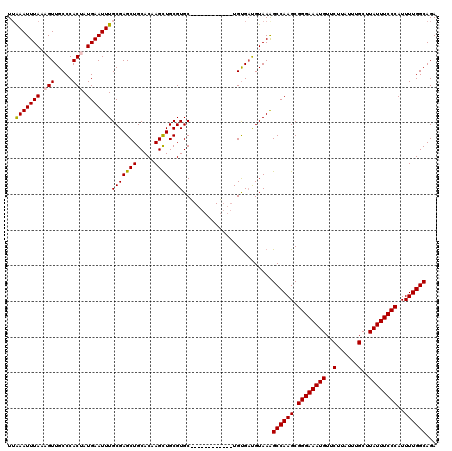
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:14:17 2006