
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 4,529,559 – 4,529,660 |
| Length | 101 |
| Max. P | 0.989137 |

| Location | 4,529,559 – 4,529,660 |
|---|---|
| Length | 101 |
| Sequences | 5 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 94.48 |
| Mean single sequence MFE | -22.34 |
| Consensus MFE | -17.50 |
| Energy contribution | -17.70 |
| Covariance contribution | 0.20 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.01 |
| Structure conservation index | 0.78 |
| SVM decision value | 0.73 |
| SVM RNA-class probability | 0.834963 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 4529559 101 + 22224390 UAUAUCGAA-ACGACGGAUAUUGAUAUUUCCCACACAGAUACACACCUACAAACGUGUAUCUGCGUUUCUGGCAUCCCUUGAAUUUCUGUGUACAUUGUUUU (((((.(((-((((.((((...((....))..((.(((((((((..........))))))))).)).......)))).)))..)))).)))))......... ( -20.90) >DroSec_CAF1 2825 101 + 1 AAUAUCGAA-ACGACGGGUAUUGAUAUUUCCCACACAGAUACACACCUACAAACGUGUAUCUGCGUUUCUGGCAUCCCUUGAAUUUCUGUGUACUUUGUUUU ......(((-(((..(((...........)))...(((((((((..........)))))))))))))))................................. ( -21.90) >DroSim_CAF1 2450 99 + 1 AAUAUCGAA-ACGA--GGUAUUGAUAUUUCCCACACAGAUACACACCUACAAACGUGUAUCUGCGUUUCUGGCAUCCCUUGAAUUUCUGUGUACUUUGUUUU ......(((-((((--(((((.((...(((..((.(((((((((..........))))))))).))....((....))..)))..))...)))))))))))) ( -25.60) >DroEre_CAF1 3762 102 + 1 AAUAUCGAAAACGACGAAUAUUGAUAUUUCCCACACAGAUACACACCAACAAACGUGUAUCUGCGUUUCUGGCAUCCCUUGAAUUUCUGUGUACGUUGUGUU ..........((((((.((((.((...(((..((.(((((((((..........))))))))).))....((....))..)))..)).)))).))))))... ( -23.80) >DroYak_CAF1 3714 101 + 1 AAUAUCGAA-AUGACGAAUAUUGAUAUUUUCCACACAGAUACACACAUACAAACGUGUAUCUGCGUUUCUGGCAUCCCUUGAAUUUCUGUGUACAUUGUUUU ((((((((.-((.....)).))))))))...(((((((((((((..........))))))))).((((..((....))..))))...))))........... ( -19.50) >consensus AAUAUCGAA_ACGACGGAUAUUGAUAUUUCCCACACAGAUACACACCUACAAACGUGUAUCUGCGUUUCUGGCAUCCCUUGAAUUUCUGUGUACAUUGUUUU ((((((((............))))))))...(((((((((((((..........))))))))).((((..((....))..))))...))))........... (-17.50 = -17.70 + 0.20)



| Location | 4,529,559 – 4,529,660 |
|---|---|
| Length | 101 |
| Sequences | 5 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 94.48 |
| Mean single sequence MFE | -25.32 |
| Consensus MFE | -21.94 |
| Energy contribution | -22.34 |
| Covariance contribution | 0.40 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.99 |
| Structure conservation index | 0.87 |
| SVM decision value | 2.15 |
| SVM RNA-class probability | 0.989137 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
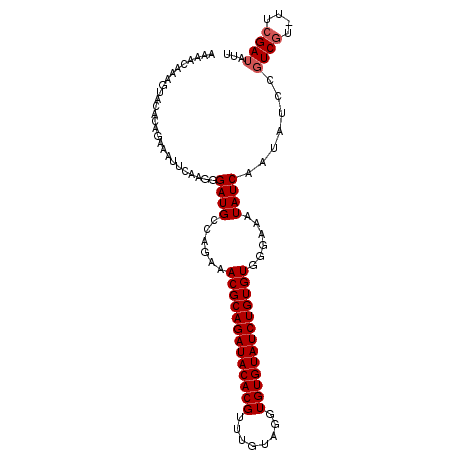
>X_DroMel_CAF1 4529559 101 - 22224390 AAAACAAUGUACACAGAAAUUCAAGGGAUGCCAGAAACGCAGAUACACGUUUGUAGGUGUGUAUCUGUGUGGGAAAUAUCAAUAUCCGUCGU-UUCGAUAUA ...............(((((...(.(((((((....(((((((((((((........))))))))))))).)).........))))).).))-)))...... ( -26.20) >DroSec_CAF1 2825 101 - 1 AAAACAAAGUACACAGAAAUUCAAGGGAUGCCAGAAACGCAGAUACACGUUUGUAGGUGUGUAUCUGUGUGGGAAAUAUCAAUACCCGUCGU-UUCGAUAUU ...............(((((....(((((.((....(((((((((((((........))))))))))))).))........)).)))...))-)))...... ( -26.90) >DroSim_CAF1 2450 99 - 1 AAAACAAAGUACACAGAAAUUCAAGGGAUGCCAGAAACGCAGAUACACGUUUGUAGGUGUGUAUCUGUGUGGGAAAUAUCAAUACC--UCGU-UUCGAUAUU ...............(((((...(((((((((....(((((((((((((........))))))))))))).))...))))....))--).))-)))...... ( -25.30) >DroEre_CAF1 3762 102 - 1 AACACAACGUACACAGAAAUUCAAGGGAUGCCAGAAACGCAGAUACACGUUUGUUGGUGUGUAUCUGUGUGGGAAAUAUCAAUAUUCGUCGUUUUCGAUAUU ..........................((((((....(((((((((((((........))))))))))))).))...)))).......((((....))))... ( -24.40) >DroYak_CAF1 3714 101 - 1 AAAACAAUGUACACAGAAAUUCAAGGGAUGCCAGAAACGCAGAUACACGUUUGUAUGUGUGUAUCUGUGUGGAAAAUAUCAAUAUUCGUCAU-UUCGAUAUU ........................(..(((...((((((((((((((((........))))))))))))).((.....))....)))..)))-..)...... ( -23.80) >consensus AAAACAAAGUACACAGAAAUUCAAGGGAUGCCAGAAACGCAGAUACACGUUUGUAGGUGUGUAUCUGUGUGGGAAAUAUCAAUAUCCGUCGU_UUCGAUAUU ..........................((((......(((((((((((((........)))))))))))))......)))).......((((....))))... (-21.94 = -22.34 + 0.40)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:13:56 2006