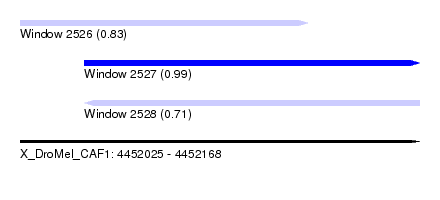

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 4,452,025 – 4,452,168 |
| Length | 143 |
| Max. P | 0.990510 |

| Location | 4,452,025 – 4,452,128 |
|---|---|
| Length | 103 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 86.29 |
| Mean single sequence MFE | -27.74 |
| Consensus MFE | -17.72 |
| Energy contribution | -17.76 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.74 |
| Structure conservation index | 0.64 |
| SVM decision value | 0.70 |
| SVM RNA-class probability | 0.826810 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 4452025 103 + 22224390 -------UCACCAUU------AUAGAG----AGAGUAUCUCAGCUCCCCUUCCCCUAAAUGGAAACCACCAUGUUUUUGGGGCCAAAUCCCCAUCACGGAUAGUUUUUCCGGAUACACAA -------...(((((------.(((.(----(((((......))))....))..)))))))).........(((....((((......))))(((.((((.......)))))))..))). ( -21.90) >DroSec_CAF1 53494 107 + 1 -------CCACCAUU------AUAGAGAGAGAGAGUAUCUCAGCUCCCCUUCCCCUAAAUGGAAGCCCCUCUGUUUUUGGGGCCAAAUCCCCAUCACGGAUACUUUUUCCGGAUACACAA -------...(((((------.(((.(..((.((((......))))..))..).))))))))..(((((.........))))).........(((.((((.......)))))))...... ( -29.40) >DroSim_CAF1 54831 105 + 1 -------CCACCAUU------AUAGAGA--GAGAGUAUCUCAGCUCCCCUUCCCCUAAAUGGAAGCCCCUCUGUUUUUGGGGCCAAAUCCCCAUCACGGAUACUUUUUCCGGAUACACAA -------........------...(.((--(((((((((((((.....(((((.......))))).....)))....(((((......)))))....))))))))))))).......... ( -27.50) >DroEre_CAF1 55475 107 + 1 CUGUCAUCCACCAUU------AUAGAG----AGAGUAUCUCGACUCCUCUUCCCCUAAAUGGAAACCCC--UGUU-UUGGGGCCAAAUUCCCAUCACGGAUACUUUUUCCGGAUACACAA .(((.((((.(((((------..((((----.((((......)))))))).......)))))......(--(((.-.(((((......)))))..))))...........)))))))... ( -27.90) >DroYak_CAF1 54201 114 + 1 CUGUCAUCCACCAUUAUGAUUAUGGAG----AGAGUAUCUCAGCUCCUCUUCCCCUAAAUGGAAACCCC--UGUUUUUGGGGCCGAAUCGCCAUCACGGAUACUUUUUCCGGAUACACAA .(((.((((..(.....).....((((----(((((((((..((...((..((((.((((((......)--)))))..))))..))...))......))))))))))))))))))))... ( -32.00) >consensus _______CCACCAUU______AUAGAG____AGAGUAUCUCAGCUCCCCUUCCCCUAAAUGGAAACCCC__UGUUUUUGGGGCCAAAUCCCCAUCACGGAUACUUUUUCCGGAUACACAA ..................................(((((..........((((.......)))).............(((((......)))))...((((.......))))))))).... (-17.72 = -17.76 + 0.04)

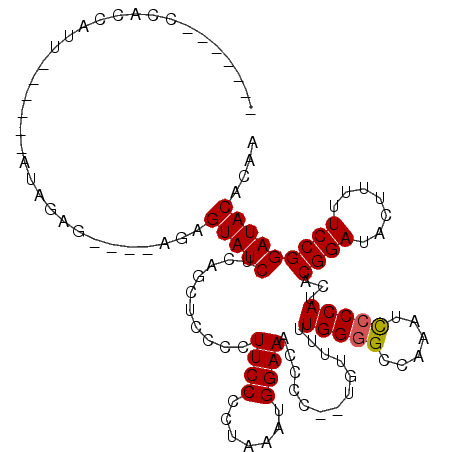
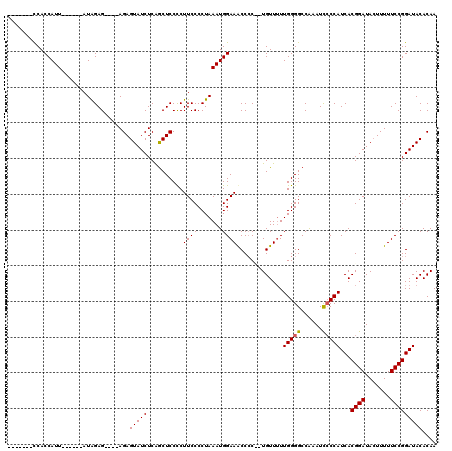
| Location | 4,452,048 – 4,452,168 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 93.82 |
| Mean single sequence MFE | -25.15 |
| Consensus MFE | -20.98 |
| Energy contribution | -21.18 |
| Covariance contribution | 0.20 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.27 |
| Structure conservation index | 0.83 |
| SVM decision value | 2.22 |
| SVM RNA-class probability | 0.990510 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 4452048 120 + 22224390 CAGCUCCCCUUCCCCUAAAUGGAAACCACCAUGUUUUUGGGGCCAAAUCCCCAUCACGGAUAGUUUUUCCGGAUACACAACUUGCAUUCCACUUUAGCUGUUUACUUAGCACUCCACAGA ..(((.........((((((((((..............((((......))))(((.((((.......)))))))............))))).)))))..........))).......... ( -23.71) >DroSec_CAF1 53521 120 + 1 CAGCUCCCCUUCCCCUAAAUGGAAGCCCCUCUGUUUUUGGGGCCAAAUCCCCAUCACGGAUACUUUUUCCGGAUACACAACUUGCAUUCCACUUUAGCUGUUUACUCAGCACUCCACAGA ...............((((((((((((((.........))))).........(((.((((.......)))))))............))))).))))((((......)))).......... ( -27.80) >DroSim_CAF1 54856 120 + 1 CAGCUCCCCUUCCCCUAAAUGGAAGCCCCUCUGUUUUUGGGGCCAAAUCCCCAUCACGGAUACUUUUUCCGGAUACACAACUUGCAUUCCACUUUAGCUGUUUACUCAGCACUCCACAGA ...............((((((((((((((.........))))).........(((.((((.......)))))))............))))).))))((((......)))).......... ( -27.80) >DroEre_CAF1 55505 117 + 1 CGACUCCUCUUCCCCUAAAUGGAAACCCC--UGUU-UUGGGGCCAAAUUCCCAUCACGGAUACUUUUUCCGGAUACACAACUUGCAUUCCACUUUAGCUGUUUACUUUGCACUCCACAGA ..............((((((((((.((((--....-..))))..........(((.((((.......)))))))............))))).)))))((((..............)))). ( -23.04) >DroYak_CAF1 54237 118 + 1 CAGCUCCUCUUCCCCUAAAUGGAAACCCC--UGUUUUUGGGGCCGAAUCGCCAUCACGGAUACUUUUUCCGGAUACACAACUUGCACUCCACUUUAGCUGUUUACUUAGCACUCCACAGA ..((...((..((((.((((((......)--)))))..))))..))......(((.((((.......))))))).........))...........((((......)))).......... ( -23.40) >consensus CAGCUCCCCUUCCCCUAAAUGGAAACCCC__UGUUUUUGGGGCCAAAUCCCCAUCACGGAUACUUUUUCCGGAUACACAACUUGCAUUCCACUUUAGCUGUUUACUUAGCACUCCACAGA ...............(((((((((.......(((....((((......))))(((.((((.......)))))))..))).......))))).))))((((......)))).......... (-20.98 = -21.18 + 0.20)


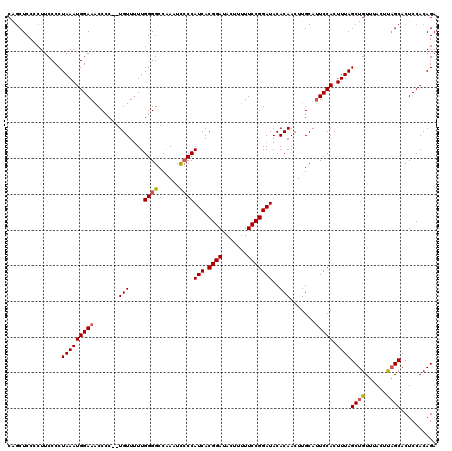
| Location | 4,452,048 – 4,452,168 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 93.82 |
| Mean single sequence MFE | -33.56 |
| Consensus MFE | -28.52 |
| Energy contribution | -28.36 |
| Covariance contribution | -0.16 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.37 |
| Structure conservation index | 0.85 |
| SVM decision value | 0.37 |
| SVM RNA-class probability | 0.707888 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 4452048 120 - 22224390 UCUGUGGAGUGCUAAGUAAACAGCUAAAGUGGAAUGCAAGUUGUGUAUCCGGAAAAACUAUCCGUGAUGGGGAUUUGGCCCCAAAAACAUGGUGGUUUCCAUUUAGGGGAAGGGGAGCUG ....................(((((.....(((.((((.....)))))))((((..((((.(((((.(((((......)))))....)))))))))))))...............))))) ( -31.80) >DroSec_CAF1 53521 120 - 1 UCUGUGGAGUGCUGAGUAAACAGCUAAAGUGGAAUGCAAGUUGUGUAUCCGGAAAAAGUAUCCGUGAUGGGGAUUUGGCCCCAAAAACAGAGGGGCUUCCAUUUAGGGGAAGGGGAGCUG ((((((((..((((......)))).....((((.((((.....)))))))).........)))....(((((......)))))...)))))..(((((((.(((....))).))))))). ( -37.00) >DroSim_CAF1 54856 120 - 1 UCUGUGGAGUGCUGAGUAAACAGCUAAAGUGGAAUGCAAGUUGUGUAUCCGGAAAAAGUAUCCGUGAUGGGGAUUUGGCCCCAAAAACAGAGGGGCUUCCAUUUAGGGGAAGGGGAGCUG ((((((((..((((......)))).....((((.((((.....)))))))).........)))....(((((......)))))...)))))..(((((((.(((....))).))))))). ( -37.00) >DroEre_CAF1 55505 117 - 1 UCUGUGGAGUGCAAAGUAAACAGCUAAAGUGGAAUGCAAGUUGUGUAUCCGGAAAAAGUAUCCGUGAUGGGAAUUUGGCCCCAA-AACA--GGGGUUUCCAUUUAGGGGAAGAGGAGUCG .((((..(.(....).)..))))((.((((((((..((((((.(.((((((((.......)))).)))).)))))))(((((..-....--))))))))))))).))............. ( -31.90) >DroYak_CAF1 54237 118 - 1 UCUGUGGAGUGCUAAGUAAACAGCUAAAGUGGAGUGCAAGUUGUGUAUCCGGAAAAAGUAUCCGUGAUGGCGAUUCGGCCCCAAAAACA--GGGGUUUCCAUUUAGGGGAAGAGGAGCUG .((((..(.(....).)..))))((.((((((((..(.((((((.((((((((.......)))).)))))))))).)(((((.......--))))))))))))).))............. ( -30.10) >consensus UCUGUGGAGUGCUAAGUAAACAGCUAAAGUGGAAUGCAAGUUGUGUAUCCGGAAAAAGUAUCCGUGAUGGGGAUUUGGCCCCAAAAACA__GGGGUUUCCAUUUAGGGGAAGGGGAGCUG .((((....(((...))).))))((.((((((((..((((((.(.((((((((.......)))).))))).))))))(((((.........))))))))))))).))............. (-28.52 = -28.36 + -0.16)


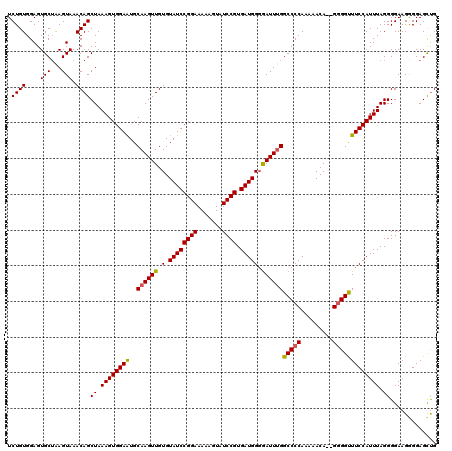
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:12:59 2006