
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 547,518 – 547,631 |
| Length | 113 |
| Max. P | 0.755127 |

| Location | 547,518 – 547,631 |
|---|---|
| Length | 113 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 77.16 |
| Mean single sequence MFE | -25.60 |
| Consensus MFE | -18.57 |
| Energy contribution | -19.64 |
| Covariance contribution | 1.06 |
| Combinations/Pair | 1.19 |
| Mean z-score | -1.63 |
| Structure conservation index | 0.73 |
| SVM decision value | 0.49 |
| SVM RNA-class probability | 0.755127 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 547518 113 + 22224390 UUUGUACAUAUAUUACAAGAAUUUUUUCUCAGGUGUAUGAAUGUUCUGUCCUUAAUAAUCGUGUGGGCUAUGGGCCUAAGGAAAUGAGGC-------ACAUCCACCUAUGGAAAUUCGAU .(((((.......)))))((((((((....(((((.(((..(((((..((((...........(((((.....)))))))))...)).))-------)))).)))))..))))))))... ( -26.40) >DroSec_CAF1 10257 94 + 1 U----------------ACAAUUUUUUCUUAGGUGUAUGAAU---CUGUCCUUAAUAAUCCAGUGGGCUAUGGGCCCGAGGAAAUGAGGC-------ACAUCCACCUAUGGAAAUUCGAU .----------------......(((((.((((((.(((...---.((.(((((....(((..(((((.....))))).)))..))))))-------)))).)))))).)))))...... ( -28.70) >DroSim_CAF1 10715 94 + 1 U----------------ACAAUUUUUUCUUAGGUGUAUGAAU---CUGUCCAUAAUAAUCCAGUGGGCUAUGGGCCCGAGGAAAUGAGGC-------ACAUCCACCUAUGGAAAUUCGAU .----------------......(((((.((((((.(((...---.(((((((.....(((..(((((.....))))).))).))).)))-------)))).)))))).)))))...... ( -27.10) >DroEre_CAF1 12018 104 + 1 U----------------ACUAUUUCAUUUUAGUUGUAUGAACGAUCCCUCUUUAAUUAUCCAUUGGGCUAUGGGCCCGAGGAAAAACGGCUCAUCCACCAUCCACCUAUGGAAAUUCGAU .----------------...(((((....(((.((.(((...................(((.((((((.....))))))))).....((.....))..))).)).)))..)))))..... ( -20.20) >consensus U________________ACAAUUUUUUCUUAGGUGUAUGAAU___CUGUCCUUAAUAAUCCAGUGGGCUAUGGGCCCGAGGAAAUGAGGC_______ACAUCCACCUAUGGAAAUUCGAU .......................(((((.((((((.(((..........(((((....(((..(((((.....))))).)))..))))).........))).)))))).)))))...... (-18.57 = -19.64 + 1.06)

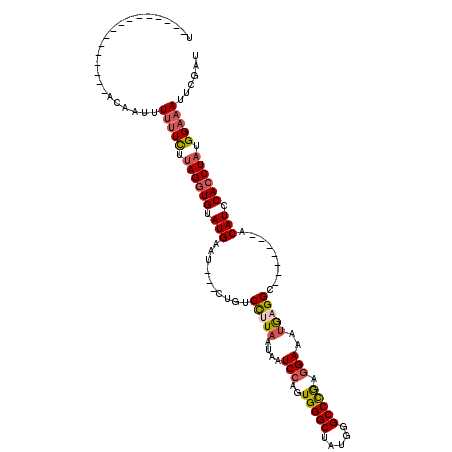
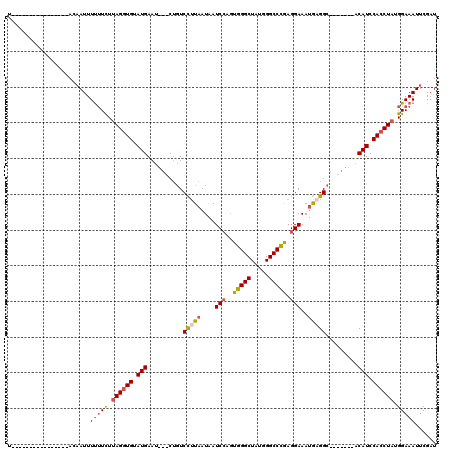
| Location | 547,518 – 547,631 |
|---|---|
| Length | 113 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 77.16 |
| Mean single sequence MFE | -23.67 |
| Consensus MFE | -15.87 |
| Energy contribution | -16.68 |
| Covariance contribution | 0.81 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.53 |
| Structure conservation index | 0.67 |
| SVM decision value | -0.03 |
| SVM RNA-class probability | 0.519443 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 547518 113 - 22224390 AUCGAAUUUCCAUAGGUGGAUGU-------GCCUCAUUUCCUUAGGCCCAUAGCCCACACGAUUAUUAAGGACAGAACAUUCAUACACCUGAGAAAAAAUUCUUGUAAUAUAUGUACAAA ...((((((.(.((((((.(((.-------........(((((((((.....)))...........)))))).........))).)))))).)...))))))(((((.......))))). ( -20.77) >DroSec_CAF1 10257 94 - 1 AUCGAAUUUCCAUAGGUGGAUGU-------GCCUCAUUUCCUCGGGCCCAUAGCCCACUGGAUUAUUAAGGACAG---AUUCAUACACCUAAGAAAAAAUUGU----------------A ......((((..(((((((((.(-------((((....(((..((((.....))))...)))......))).)).---)))....)))))).)))).......----------------. ( -24.00) >DroSim_CAF1 10715 94 - 1 AUCGAAUUUCCAUAGGUGGAUGU-------GCCUCAUUUCCUCGGGCCCAUAGCCCACUGGAUUAUUAUGGACAG---AUUCAUACACCUAAGAAAAAAUUGU----------------A ......((((..(((((((((.(-------(((.....(((..((((.....))))...))).......)).)).---)))....)))))).)))).......----------------. ( -23.10) >DroEre_CAF1 12018 104 - 1 AUCGAAUUUCCAUAGGUGGAUGGUGGAUGAGCCGUUUUUCCUCGGGCCCAUAGCCCAAUGGAUAAUUAAAGAGGGAUCGUUCAUACAACUAAAAUGAAAUAGU----------------A .(((....((((....))))((((..((((((.(...((((((((((.....))))(((.....)))...)))))).)))))))...))))...)))......----------------. ( -26.80) >consensus AUCGAAUUUCCAUAGGUGGAUGU_______GCCUCAUUUCCUCGGGCCCAUAGCCCACUGGAUUAUUAAGGACAG___AUUCAUACACCUAAGAAAAAAUUGU________________A ............((((((.(((.........((.....(((..((((.....))))...))).......))..........))).))))))............................. (-15.87 = -16.68 + 0.81)

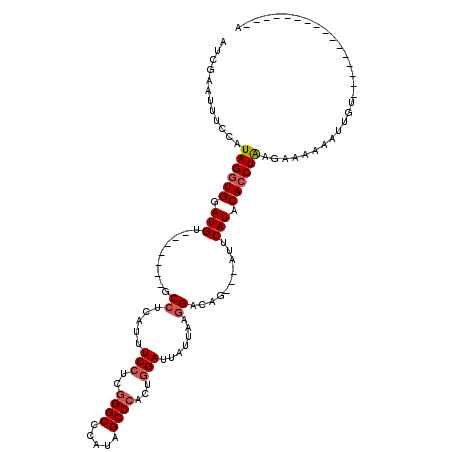
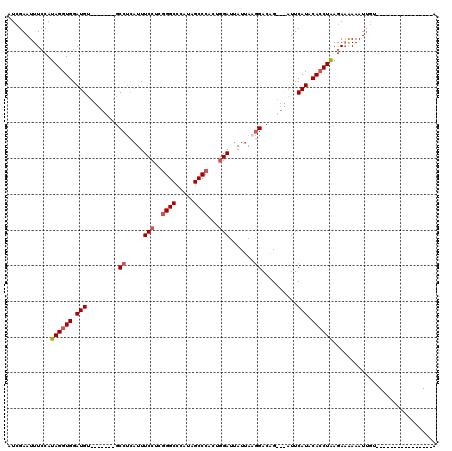
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:37:24 2006