
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 4,408,260 – 4,408,408 |
| Length | 148 |
| Max. P | 0.903465 |

| Location | 4,408,260 – 4,408,376 |
|---|---|
| Length | 116 |
| Sequences | 5 |
| Columns | 116 |
| Reading direction | reverse |
| Mean pairwise identity | 98.26 |
| Mean single sequence MFE | -29.02 |
| Consensus MFE | -25.78 |
| Energy contribution | -25.66 |
| Covariance contribution | -0.12 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.39 |
| Structure conservation index | 0.89 |
| SVM decision value | 0.15 |
| SVM RNA-class probability | 0.605460 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 4408260 116 - 22224390 CAUCCCUGCCACACAUGAUCGCAACUCGUAAAUAAACGAAAACCGAAAGGGGCACGUGCCGCUCAAUGUGCAAGGAUCAAGGAUCUCCCCCUUUGGCAAUUCUAUCUGCAACUGCA ......(((((....(((((.....((((......))))...((....)).((((((........))))))...)))))(((.......))).))))).........((....)). ( -29.60) >DroSec_CAF1 10762 114 - 1 CAUCCCUGCCACACAUGAUCGCAACUCGUAAAUAAACGAAAACCGA--GGGGCACGUGCCGCUCAAUGUGCAAGGAUCAAGGAUCUCCCCCUUUGGCAAUUCUAUCUGCAACUGCA ......(((((....((((((((..((((......)))).....((--(.(((....))).)))....)))...)))))(((.......))).))))).........((....)). ( -29.50) >DroSim_CAF1 10860 114 - 1 CAUCCCUGCCACACAUGAUCGCAACUCGUAAAUAAACGAAAACCGA--GGGGCACGUGCCGCUCAAUGUGCAAGGAUCAAGGAUCUCCCCCUUUGGCAAUUCUAUCUGCAACUGCA ......(((((....((((((((..((((......)))).....((--(.(((....))).)))....)))...)))))(((.......))).))))).........((....)). ( -29.50) >DroEre_CAF1 11610 114 - 1 CAUCCCUGCCACACAUGAUCGCAACUCGUAAAUAAACGAAAACCGA--GGGGCACGUGCCGCUCAAUGUGCAAGGAUCAAGGAUCUCCCCCUUUGGCAAUUCUAUCUGCAACUGCA ......(((((....((((((((..((((......)))).....((--(.(((....))).)))....)))...)))))(((.......))).))))).........((....)). ( -29.50) >DroYak_CAF1 10943 114 - 1 CAUCCCUGGCACACAUGAUCGCAACUCGUAAAUAAACGAAAACCGA--GGGGCACGUGCCGCUCAAUGUGCAAGAAUCAAGGAUCUCCCCCUUUGUCAAUUCUAUCUGCAACUGCA ........(((((............((((......)))).....((--(.(((....))).)))..))))).((((((((((........)))))...)))))....((....)). ( -27.00) >consensus CAUCCCUGCCACACAUGAUCGCAACUCGUAAAUAAACGAAAACCGA__GGGGCACGUGCCGCUCAAUGUGCAAGGAUCAAGGAUCUCCCCCUUUGGCAAUUCUAUCUGCAACUGCA ......(((((.......(((....((((......))))....)))..(((((((((........))))))..(((((...)))))..)))..))))).........((....)). (-25.78 = -25.66 + -0.12)


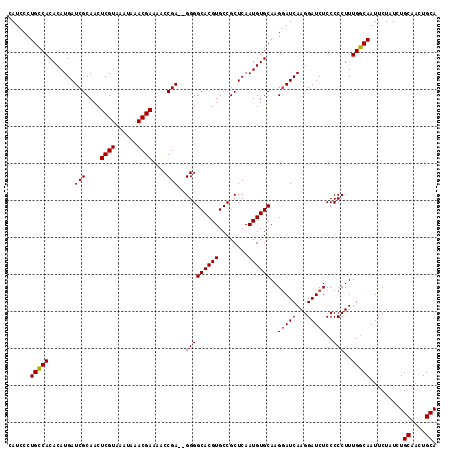
| Location | 4,408,296 – 4,408,408 |
|---|---|
| Length | 112 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 93.07 |
| Mean single sequence MFE | -39.00 |
| Consensus MFE | -35.18 |
| Energy contribution | -35.42 |
| Covariance contribution | 0.24 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.42 |
| Structure conservation index | 0.90 |
| SVM decision value | 1.03 |
| SVM RNA-class probability | 0.903465 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 4408296 112 + 22224390 UUGAUCCUUGCACAUUGAGCGGCACGUGCCCCUUUCGGUUUUCGUUUAUUUACGAGUUGCGAUCAUGUGUGGCAGGGAUGCAGCGCGUGCGAGGAUCCUCCGCUUCUCA--------AUU ..(((((((((((...(((.(((....))).)))(((...(((((......)))))...)))....((((.(((....))).)))))))))))))))............--------... ( -39.20) >DroSec_CAF1 10798 110 + 1 UUGAUCCUUGCACAUUGAGCGGCACGUGCCCC--UCGGUUUUCGUUUAUUUACGAGUUGCGAUCAUGUGUGGCAGGGAUGCAGCGCGUGCGAGGAUCCUCCGCUCCUCG--------AUU ..(((((((((((((((((.(((....))).)--)))))..((((......))))(((((.(((.((.....))..))))))))..)))))))))))............--------... ( -42.30) >DroSim_CAF1 10896 118 + 1 UUGAUCCUUGCACAUUGAGCGGCACGUGCCCC--UCGGUUUUCGUUUAUUUACGAGUUGCGAUCAUGUGUGGCAGGGAUGCAGCGCGUGCGAGUAUCCUCCGCUCCUCGAUUCCUCGAUU ................((((((((((((((((--(......((((......))))((..((......))..)))))).....))))))))(((....)))))))).((((....)))).. ( -37.80) >DroEre_CAF1 11646 110 + 1 UUGAUCCUUGCACAUUGAGCGGCACGUGCCCC--UCGGUUUUCGUUUAUUUACGAGUUGCGAUCAUGUGUGGCAGGGAUGCAGCGCGUGCAACGAUCCUCUGCUCCUCA--------AUU ..((((.((((((((((((.(((....))).)--)))))..((((......))))(((((.(((.((.....))..))))))))..)))))).))))............--------... ( -36.10) >DroYak_CAF1 10979 110 + 1 UUGAUUCUUGCACAUUGAGCGGCACGUGCCCC--UCGGUUUUCGUUUAUUUACGAGUUGCGAUCAUGUGUGCCAGGGAUGCAGCGCGUGCAAGGAUCCUCGGCUCCUCG--------AUU ..(((((((((((((((((.(((....))).)--)))))..((((......))))(((((.(((.((.....))..))))))))..))))))))))).((((....)))--------).. ( -39.60) >consensus UUGAUCCUUGCACAUUGAGCGGCACGUGCCCC__UCGGUUUUCGUUUAUUUACGAGUUGCGAUCAUGUGUGGCAGGGAUGCAGCGCGUGCGAGGAUCCUCCGCUCCUCG________AUU ..(((((((((((.....(((((....(((......)))..((((......)))))))))......((((.(((....))).)))))))))))))))....................... (-35.18 = -35.42 + 0.24)

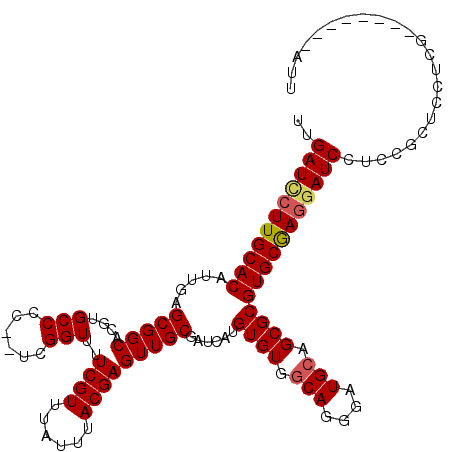
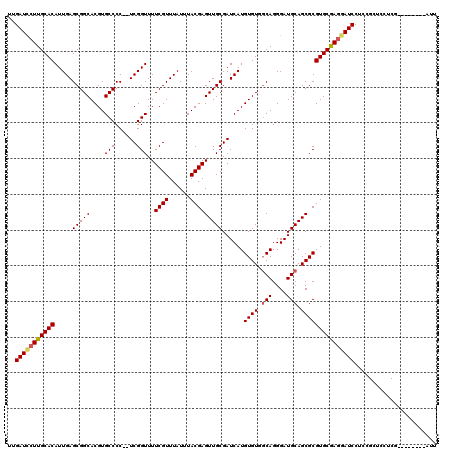
| Location | 4,408,296 – 4,408,408 |
|---|---|
| Length | 112 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 93.07 |
| Mean single sequence MFE | -34.74 |
| Consensus MFE | -26.96 |
| Energy contribution | -27.76 |
| Covariance contribution | 0.80 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.56 |
| Structure conservation index | 0.78 |
| SVM decision value | 0.03 |
| SVM RNA-class probability | 0.549754 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 4408296 112 - 22224390 AAU--------UGAGAAGCGGAGGAUCCUCGCACGCGCUGCAUCCCUGCCACACAUGAUCGCAACUCGUAAAUAAACGAAAACCGAAAGGGGCACGUGCCGCUCAAUGUGCAAGGAUCAA ...--------............((((((.(((((.((.((((...((((...............((((......))))...((....)))))).)))).))....))))).)))))).. ( -35.90) >DroSec_CAF1 10798 110 - 1 AAU--------CGAGGAGCGGAGGAUCCUCGCACGCGCUGCAUCCCUGCCACACAUGAUCGCAACUCGUAAAUAAACGAAAACCGA--GGGGCACGUGCCGCUCAAUGUGCAAGGAUCAA ..(--------((.....)))..((((((.(((((((...(((...........)))..)))...((((......)))).....((--(.(((....))).)))...)))).)))))).. ( -33.40) >DroSim_CAF1 10896 118 - 1 AAUCGAGGAAUCGAGGAGCGGAGGAUACUCGCACGCGCUGCAUCCCUGCCACACAUGAUCGCAACUCGUAAAUAAACGAAAACCGA--GGGGCACGUGCCGCUCAAUGUGCAAGGAUCAA ..((((....))))(..((((.((((.(.((....))..).))))))))..)...((((((((..((((......)))).....((--(.(((....))).)))....)))...))))). ( -36.10) >DroEre_CAF1 11646 110 - 1 AAU--------UGAGGAGCAGAGGAUCGUUGCACGCGCUGCAUCCCUGCCACACAUGAUCGCAACUCGUAAAUAAACGAAAACCGA--GGGGCACGUGCCGCUCAAUGUGCAAGGAUCAA ...--------((.(..((((.((((.((.((....)).))))))))))..).))((((((((..((((......)))).....((--(.(((....))).)))....)))...))))). ( -36.00) >DroYak_CAF1 10979 110 - 1 AAU--------CGAGGAGCCGAGGAUCCUUGCACGCGCUGCAUCCCUGGCACACAUGAUCGCAACUCGUAAAUAAACGAAAACCGA--GGGGCACGUGCCGCUCAAUGUGCAAGAAUCAA ..(--------((......))).(((.((((((((...((((((..((.....)).))).)))..((((......)))).....((--(.(((....))).)))..)))))))).))).. ( -32.30) >consensus AAU________CGAGGAGCGGAGGAUCCUCGCACGCGCUGCAUCCCUGCCACACAUGAUCGCAACUCGUAAAUAAACGAAAACCGA__GGGGCACGUGCCGCUCAAUGUGCAAGGAUCAA .......................((((((.(((((.((.((((...((((...............((((......))))...((....)))))).)))).))....))))).)))))).. (-26.96 = -27.76 + 0.80)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:10:17 2006