
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 4,178,664 – 4,178,824 |
| Length | 160 |
| Max. P | 0.500000 |

| Location | 4,178,664 – 4,178,784 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 89.25 |
| Mean single sequence MFE | -23.79 |
| Consensus MFE | -16.69 |
| Energy contribution | -17.25 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.56 |
| Structure conservation index | 0.70 |
| SVM decision value | -0.07 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
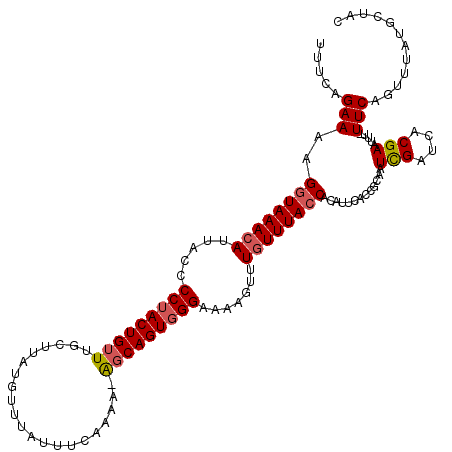
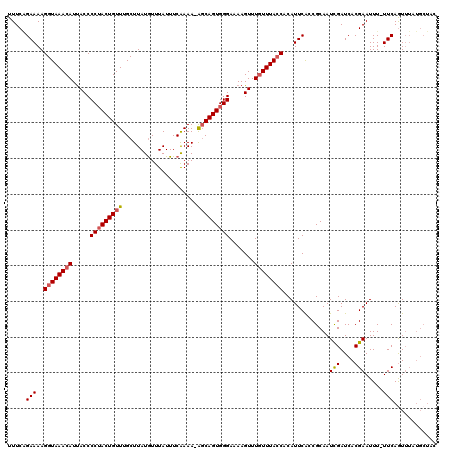
>X_DroMel_CAF1 4178664 120 - 22224390 UUUCAGAAAAGGUAAACAUUACCCCUACUGUUUGCUUAUGUUUAUUUCAAAAUAGCAGUGGGAAAAGUUUGUUUACCACAUUCACCGCAAUCGAUCACGAAUUUUUUCAGUUUAUGCUAC .....(((((((((((((.....(((((((((.....................))))))))).......)))))))).............(((....)))...)))))(((....))).. ( -24.60) >DroSec_CAF1 11296 119 - 1 UUUCAGAAAAGGUAAACAUUACCCCUACUGUUUGCUUAUGUUUAUUUCAAAAAGCCAGUGGGAAAAGUUUGUUUAACACAUUCACCGCAAUCGAUCACGAAUUU-UUCAAUUUAUGCUAC .....((((((((..........(((((((...((((.((.......))..)))))))))))....(..(((.....)))..))))....(((....)))..))-)))............ ( -19.00) >DroEre_CAF1 11476 118 - 1 UUUCGGAAAAGGUAAACAUCACCCCAACUGUUUGCUUAUGUUUAUUUCAAAA-GGCAGUGGGAAAAGUUUAUUUACCACAUUCGCCGCAAUUGAUCACGAAUUU-UUCAGUUUAUGCUAC .((((((..(((((((((..........))))))))).........((((..-(((((((((.(((.....))).)).)))).)))....)))))).))))...-...(((....))).. ( -22.00) >DroYak_CAF1 12460 108 - 1 UUUCUGAAAAGGUAAACAUUACCCCUACUGUUUGCUUAUAUUUAUUUCGAAA-AGCAGUGGGA-AAGUUUGUUUACCACAUUCACCGCAAUCGAUCACGAAUUU-UUCAGU--------- ...(((((((((((((((.....((((((((((..................)-))))))))).-.....)))))))).............(((....)))..))-))))).--------- ( -29.57) >consensus UUUCAGAAAAGGUAAACAUUACCCCUACUGUUUGCUUAUGUUUAUUUCAAAA_AGCAGUGGGAAAAGUUUGUUUACCACAUUCACCGCAAUCGAUCACGAAUUU_UUCAGUUUAUGCUAC .....(((..((((((((.....(((((((((.....................))))))))).......)))))))).............(((....))).....)))............ (-16.69 = -17.25 + 0.56)

| Location | 4,178,704 – 4,178,824 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 83.28 |
| Mean single sequence MFE | -23.80 |
| Consensus MFE | -17.84 |
| Energy contribution | -18.15 |
| Covariance contribution | 0.31 |
| Combinations/Pair | 1.14 |
| Mean z-score | -1.44 |
| Structure conservation index | 0.75 |
| SVM decision value | -0.06 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
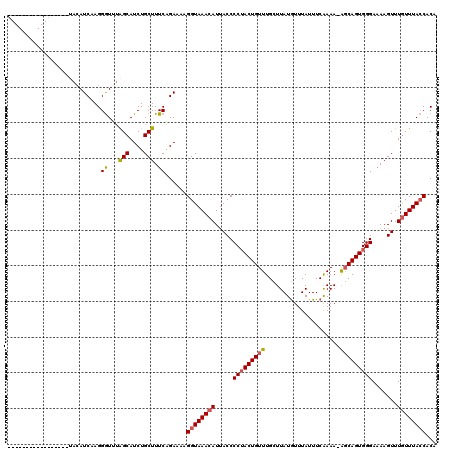
>X_DroMel_CAF1 4178704 120 - 22224390 GUACAUCCGUACAUACAUACAUCAAGGGUUUGGCAUCUGCUUUCAGAAAAGGUAAACAUUACCCCUACUGUUUGCUUAUGUUUAUUUCAAAAUAGCAGUGGGAAAAGUUUGUUUACCACA ((((....))))..............((...(((....))).))......((((((((.....(((((((((.....................))))))))).......))))))))... ( -26.00) >DroSec_CAF1 11335 103 - 1 A-----------------AGAUCAAGGAUUUGGCAUCUGCUUUCAGAAAAGGUAAACAUUACCCCUACUGUUUGCUUAUGUUUAUUUCAAAAAGCCAGUGGGAAAAGUUUGUUUAACACA .-----------------............((((.((((....))))..(((((((((.((....)).)))))))))................))))(((..(((......)))..))). ( -19.50) >DroEre_CAF1 11515 102 - 1 -----------------UACAUCAAGGGUGUAGCAUCUGCUUUCGGAAAAGGUAAACAUCACCCCAACUGUUUGCUUAUGUUUAUUUCAAAA-GGCAGUGGGAAAAGUUUAUUUACCACA -----------------(((((.....))))).....((((((..(((((((((((((..........))))))))).......))))..))-))))((((.(((......))).)))). ( -22.11) >DroYak_CAF1 12490 101 - 1 -----------------UACAUCAAGGGUAAAGCAUCUGCUUUCUGAAAAGGUAAACAUUACCCCUACUGUUUGCUUAUAUUUAUUUCGAAA-AGCAGUGGGA-AAGUUUGUUUACCACA -----------------.........((.(((((....))))))).....((((((((.....((((((((((..................)-))))))))).-.....))))))))... ( -27.57) >consensus _________________UACAUCAAGGGUUUAGCAUCUGCUUUCAGAAAAGGUAAACAUUACCCCUACUGUUUGCUUAUGUUUAUUUCAAAA_AGCAGUGGGAAAAGUUUGUUUACCACA ..........................((...(((....))).))......((((((((.....(((((((((.....................))))))))).......))))))))... (-17.84 = -18.15 + 0.31)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:08:10 2006