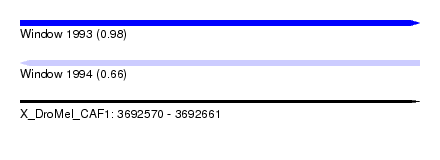

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 3,692,570 – 3,692,661 |
| Length | 91 |
| Max. P | 0.983669 |

| Location | 3,692,570 – 3,692,661 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 98 |
| Reading direction | forward |
| Mean pairwise identity | 86.98 |
| Mean single sequence MFE | -38.00 |
| Consensus MFE | -28.06 |
| Energy contribution | -27.90 |
| Covariance contribution | -0.16 |
| Combinations/Pair | 1.15 |
| Mean z-score | -3.00 |
| Structure conservation index | 0.74 |
| SVM decision value | 1.95 |
| SVM RNA-class probability | 0.983669 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
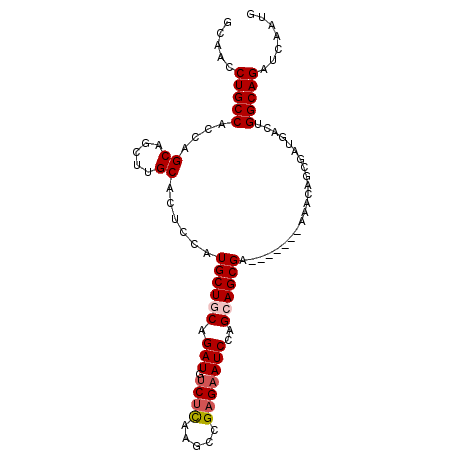
>X_DroMel_CAF1 3692570 91 + 22224390 CAUUGAUCUGCCAGUCAUCGCUGUUU-------UCGCUGCUGGAUUCUCGGCUUAAGACAUCUGCAGCAUGGAGUGCAAGCUGCUGGUGGCAGGUUGC ....((((((((.(((...((.....-------..)).(((((....)))))....)))(((.(((((.((.....)).))))).))))))))))).. ( -32.10) >DroSec_CAF1 26149 90 + 1 CAUAGAUCUGCCAGUCAUCGCU-UUU-------UCGCUGCUGGAUUCUCGGUCUGAGACAUCUGUAGCAUGGAGUGCAAGCUGCUGGCGGCAGGUUGC ....((((((((.((((..(((-((.-------..(((((.((((((((.....)))).)))))))))..)))))((.....)))))))))))))).. ( -33.00) >DroSim_CAF1 29205 90 + 1 CAUAGAUCUGCCAGUCAUCGCU-UUU-------UCGCUGCUGGAUUCUCGGCCUGAGACAUCUGUAGCAUGAAGUGCAAGCUGCUGGCGGCAGGUUGC ....((((((((.((((..(((-((.-------..(((((.((((((((.....)))).)))))))))..)))))((.....)))))))))))))).. ( -33.40) >DroEre_CAF1 31099 91 + 1 CAUUGAUCUGCCUGCCACCGCUGAUU-------UCGCUGCUGGAUUCUCGGCUUGCGGCAUCUGCAGCAUGGGUUGCAUGCUGCUGGUGGCAGGUUGC .........((((((((((...(((.-------((((.(((((....)))))..)))).))).((((((((.....)))))))).))))))))))... ( -46.90) >DroYak_CAF1 26663 98 + 1 CAUUGAUCUGCCAGCCAUCGCUGUUUUAGAUUCUCGCUGCUGGAUUCUCGGCUUGAGACAUCUGCAGCAUGGGUUGCAUGCUGCUGCUGGCAGGUUUC ....(((((((((((............(((((((((..(((((....))))).))))).))))((((((((.....)))))))).))))))))))).. ( -44.60) >consensus CAUUGAUCUGCCAGUCAUCGCUGUUU_______UCGCUGCUGGAUUCUCGGCUUGAGACAUCUGCAGCAUGGAGUGCAAGCUGCUGGUGGCAGGUUGC ....((((((((.((((..................(((((.((((((((.....)))).))))))))).......((.....)))))))))))))).. (-28.06 = -27.90 + -0.16)



| Location | 3,692,570 – 3,692,661 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 98 |
| Reading direction | reverse |
| Mean pairwise identity | 86.98 |
| Mean single sequence MFE | -30.02 |
| Consensus MFE | -20.38 |
| Energy contribution | -21.02 |
| Covariance contribution | 0.64 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.08 |
| Structure conservation index | 0.68 |
| SVM decision value | 0.27 |
| SVM RNA-class probability | 0.663974 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 3692570 91 - 22224390 GCAACCUGCCACCAGCAGCUUGCACUCCAUGCUGCAGAUGUCUUAAGCCGAGAAUCCAGCAGCGA-------AAACAGCGAUGACUGGCAGAUCAAUG ((((.((((.....)))).)))).(((((((((((.(((.((((.....)))))))..)))))).-------...((....))..))).))....... ( -24.50) >DroSec_CAF1 26149 90 - 1 GCAACCUGCCGCCAGCAGCUUGCACUCCAUGCUACAGAUGUCUCAGACCGAGAAUCCAGCAGCGA-------AAA-AGCGAUGACUGGCAGAUCUAUG ((((.((((.....)))).)))).(((((((((...(((.((((.....))))))).))))((..-------...-.))......))).))....... ( -23.60) >DroSim_CAF1 29205 90 - 1 GCAACCUGCCGCCAGCAGCUUGCACUUCAUGCUACAGAUGUCUCAGGCCGAGAAUCCAGCAGCGA-------AAA-AGCGAUGACUGGCAGAUCUAUG .....((((((.((...((((....(((.((((...(((.((((.....))))))).))))..))-------).)-)))..))..))))))....... ( -24.30) >DroEre_CAF1 31099 91 - 1 GCAACCUGCCACCAGCAGCAUGCAACCCAUGCUGCAGAUGCCGCAAGCCGAGAAUCCAGCAGCGA-------AAUCAGCGGUGGCAGGCAGAUCAAUG ....(((((((((.((((((((.....)))))))).(((..(((..((.(......).)).))).-------.)))...))))))))).......... ( -40.50) >DroYak_CAF1 26663 98 - 1 GAAACCUGCCAGCAGCAGCAUGCAACCCAUGCUGCAGAUGUCUCAAGCCGAGAAUCCAGCAGCGAGAAUCUAAAACAGCGAUGGCUGGCAGAUCAAUG .....((((((((.((((((((.....))))))))((((.((((..((.(......).))...))))))))............))))))))....... ( -37.20) >consensus GCAACCUGCCACCAGCAGCUUGCACUCCAUGCUGCAGAUGUCUCAAGCCGAGAAUCCAGCAGCGA_______AAACAGCGAUGACUGGCAGAUCAAUG .....(((((....((.....))......((((((.(((.((((.....)))))))..))))))......................)))))....... (-20.38 = -21.02 + 0.64)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:04:44 2006