
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 3,625,178 – 3,625,273 |
| Length | 95 |
| Max. P | 0.999953 |

| Location | 3,625,178 – 3,625,273 |
|---|---|
| Length | 95 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 74.42 |
| Mean single sequence MFE | -36.79 |
| Consensus MFE | -25.36 |
| Energy contribution | -26.54 |
| Covariance contribution | 1.19 |
| Combinations/Pair | 1.12 |
| Mean z-score | -3.19 |
| Structure conservation index | 0.69 |
| SVM decision value | 4.82 |
| SVM RNA-class probability | 0.999953 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 3625178 95 + 22224390 ---GCAGCUGCUUUCCGUUUUGGAUUGGAAGUUCG----------UUGGAAUGGAAGGCAGCAGC----AAAAGAA----GA-AGAAGAAGCUGCAACGCC---GCGAAGCUGGCAGCGC ---((.((((((((((((((..(...((....)).----------)..)))))))))))))).))----.......----..-.......(((((...((.---.....))..))))).. ( -36.80) >DroSec_CAF1 4528 95 + 1 ---GCAGCUGCUUUCCGUUUUGGAUUGGAAGUUCG----------UUGGAAUGGAAGGCAGCAGC----GAAAGAA----GA-AGAAGAAGCUGCAACGCC---GCGAAGCUGGCAGCGC ---((.((((((((((((((..(...((....)).----------)..)))))))))))))).))----.......----..-.......(((((...((.---.....))..))))).. ( -36.80) >DroEre_CAF1 4573 95 + 1 GCAGCAGCUGCUUUCCGUUUUGGAUUGGAAGUUCG----------GUGGAAUGGAAGGCAGCAGC----AAAAGAA----G----AAGAAGCUGCAACGCC---GCGAAGCUGGCAGCGC ((.((.(((((((((((((((((((.....)))))----------...)))))))))))))).))----.......----.----.....(((((...((.---.....))..))))))) ( -37.40) >DroAna_CAF1 9693 110 + 1 ---GCAGCUGCUUUCCGU-------UGGAAGUUGCUGGCUAUGUUGUGGAAUGGUAAGAAAUAGAGGGGAAGGGAAGCAGGAAGGAAGAGGCUGCAGCGCUAAAGCGACGCAGUCAGCGU ---((..(((((((((..-------.........((.(((((........))))).)).............))))))))).........((((((..(((....)))..)))))).)).. ( -36.15) >consensus ___GCAGCUGCUUUCCGUUUUGGAUUGGAAGUUCG__________GUGGAAUGGAAGGCAGCAGC____AAAAGAA____GA_AGAAGAAGCUGCAACGCC___GCGAAGCUGGCAGCGC ...((.(((((((((((((((..........................)))))))))))))))............................(((((..(((....)))......))))))) (-25.36 = -26.54 + 1.19)


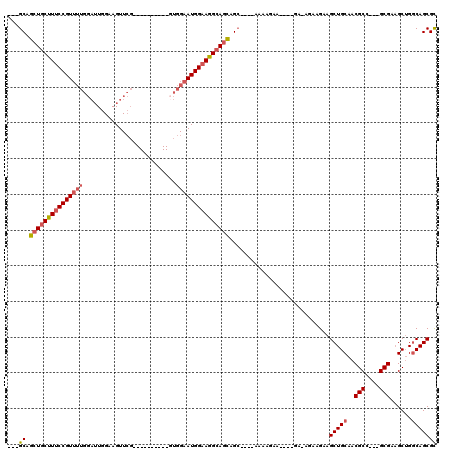
| Location | 3,625,178 – 3,625,273 |
|---|---|
| Length | 95 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 74.42 |
| Mean single sequence MFE | -23.61 |
| Consensus MFE | -16.53 |
| Energy contribution | -17.59 |
| Covariance contribution | 1.06 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.38 |
| Structure conservation index | 0.70 |
| SVM decision value | 0.04 |
| SVM RNA-class probability | 0.552400 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 3625178 95 - 22224390 GCGCUGCCAGCUUCGC---GGCGUUGCAGCUUCUUCU-UC----UUCUUUU----GCUGCUGCCUUCCAUUCCAA----------CGAACUUCCAAUCCAAAACGGAAAGCAGCUGC--- ..(((((..(((....---)))...))))).......-..----.......----((.(((((.((((.......----------...................)))).))))).))--- ( -24.77) >DroSec_CAF1 4528 95 - 1 GCGCUGCCAGCUUCGC---GGCGUUGCAGCUUCUUCU-UC----UUCUUUC----GCUGCUGCCUUCCAUUCCAA----------CGAACUUCCAAUCCAAAACGGAAAGCAGCUGC--- ..(((((..(((....---)))...))))).......-..----.......----((.(((((.((((.......----------...................)))).))))).))--- ( -25.07) >DroEre_CAF1 4573 95 - 1 GCGCUGCCAGCUUCGC---GGCGUUGCAGCUUCUU----C----UUCUUUU----GCUGCUGCCUUCCAUUCCAC----------CGAACUUCCAAUCCAAAACGGAAAGCAGCUGCUGC (((((((.......))---))))).(((((.....----.----.......----...(((((.((((.......----------...................)))).))))).))))) ( -26.60) >DroAna_CAF1 9693 110 - 1 ACGCUGACUGCGUCGCUUUAGCGCUGCAGCCUCUUCCUUCCUGCUUCCCUUCCCCUCUAUUUCUUACCAUUCCACAACAUAGCCAGCAACUUCCA-------ACGGAAAGCAGCUGC--- ..((((.(((((.(((....))).)))))........((((((((...((..............................))..)))).......-------..))))..))))...--- ( -18.01) >consensus GCGCUGCCAGCUUCGC___GGCGUUGCAGCUUCUUCU_UC____UUCUUUU____GCUGCUGCCUUCCAUUCCAA__________CGAACUUCCAAUCCAAAACGGAAAGCAGCUGC___ ..(((((..((.........))...))))).........................((.(((((.((((....................................)))).))))).))... (-16.53 = -17.59 + 1.06)


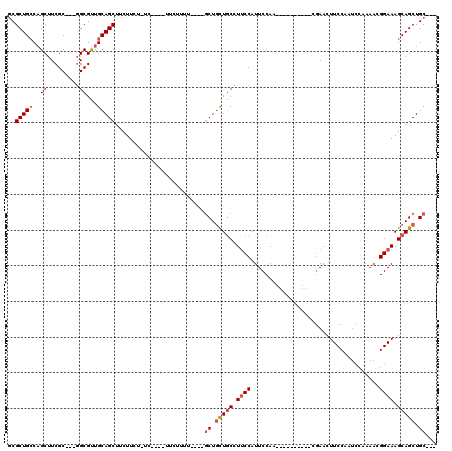
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:04:10 2006