
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 3,588,185 – 3,588,292 |
| Length | 107 |
| Max. P | 0.794487 |

| Location | 3,588,185 – 3,588,292 |
|---|---|
| Length | 107 |
| Sequences | 4 |
| Columns | 115 |
| Reading direction | forward |
| Mean pairwise identity | 85.56 |
| Mean single sequence MFE | -17.70 |
| Consensus MFE | -14.68 |
| Energy contribution | -14.68 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.75 |
| Structure conservation index | 0.83 |
| SVM decision value | 0.60 |
| SVM RNA-class probability | 0.794487 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
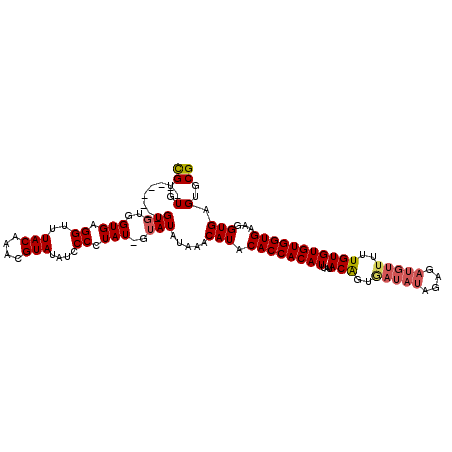
>X_DroMel_CAF1 3588185 107 + 22224390 CGCACUCACCUUCACCACACACAAAAACAUCUCUAUAUCCCCGUAUAAUGUGGUGUAUGUUUAUAUACAAUAGGGGAUAUACGUUUGUAAACCUCACCACAC--------ACACG ..........................(((....(((((((((.(((..((((.((((....))))))))))))))))))))....)))..............--------..... ( -17.90) >DroSec_CAF1 47158 107 + 1 CGCACUCACCUUCACCACACACAAAAACAUCUCUAUAUCAACGUAUAAUGUGGUGUAUGUUUAUAUACAAUAGGGGAUAUACGUUUGUAAACCUCACCACAC--------ACACG .........................................(((....(((((((...(((((((.((.(((.....)))..)).)))))))..))))))).--------..))) ( -16.70) >DroEre_CAF1 42038 112 + 1 CGCACUCACCUUCACCACACACAAAAACAUCUCUAUAUUGCUGUAGAAUGUGGUGUAUGUUUAUAU---AUAGGGGAUAUACGAUUGUAAACCCCACCACACACACACACACACA ...............................((((((....)))))).((((((((((((....))---)))(((..((((....))))..)))))))))).............. ( -20.20) >DroYak_CAF1 42926 97 + 1 CGCACUCACCUUCACCACACACAAAAACAU---------AAUGUAUAAUGUGGUGUAUGUUUAUAUAU-AUAUGGUAUAUACGUUUGUAAACCUCACCACAC--------ACACG ..............................---------.........(((((((...(((((((.(.-.((((...))))..).)))))))..))))))).--------..... ( -16.00) >consensus CGCACUCACCUUCACCACACACAAAAACAUCUCUAUAUCAACGUAUAAUGUGGUGUAUGUUUAUAUAC_AUAGGGGAUAUACGUUUGUAAACCUCACCACAC________ACACG ................................................(((((((...(((((((....(((.....))).....)))))))..))))))).............. (-14.68 = -14.68 + -0.00)
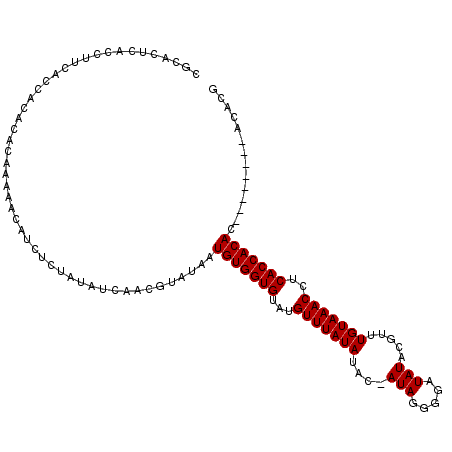



| Location | 3,588,185 – 3,588,292 |
|---|---|
| Length | 107 |
| Sequences | 4 |
| Columns | 115 |
| Reading direction | reverse |
| Mean pairwise identity | 85.56 |
| Mean single sequence MFE | -30.87 |
| Consensus MFE | -17.96 |
| Energy contribution | -18.90 |
| Covariance contribution | 0.94 |
| Combinations/Pair | 1.09 |
| Mean z-score | -2.49 |
| Structure conservation index | 0.58 |
| SVM decision value | 0.20 |
| SVM RNA-class probability | 0.630768 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 3588185 107 - 22224390 CGUGU--------GUGUGGUGAGGUUUACAAACGUAUAUCCCCUAUUGUAUAUAAACAUACACCACAUUAUACGGGGAUAUAGAGAUGUUUUUGUGUGUGGUGAAGGUGAGUGCG ..(((--------(((..(((.((..(((....)))....)).)))..))))))..(((.((((((((...((((((((((....))))))))))))))))))...)))...... ( -28.70) >DroSec_CAF1 47158 107 - 1 CGUGU--------GUGUGGUGAGGUUUACAAACGUAUAUCCCCUAUUGUAUAUAAACAUACACCACAUUAUACGUUGAUAUAGAGAUGUUUUUGUGUGUGGUGAAGGUGAGUGCG ..(((--------(((..(((.((..(((....)))....)).)))..))))))..(((.((((((((...(((..(((((....)))))..)))))))))))...)))...... ( -26.40) >DroEre_CAF1 42038 112 - 1 UGUGUGUGUGUGUGUGUGGUGGGGUUUACAAUCGUAUAUCCCCUAU---AUAUAAACAUACACCACAUUCUACAGCAAUAUAGAGAUGUUUUUGUGUGUGGUGAAGGUGAGUGCG .((((((((.(((((((((.(((((.(((....))).)))))))))---))))).))))))))(((.(((.((.(((.(((((((....)))))))))).))))).)))...... ( -39.60) >DroYak_CAF1 42926 97 - 1 CGUGU--------GUGUGGUGAGGUUUACAAACGUAUAUACCAUAU-AUAUAUAAACAUACACCACAUUAUACAUU---------AUGUUUUUGUGUGUGGUGAAGGUGAGUGCG .((((--------((((((((.....(((....)))..))))))))-)))).....(((.((((.((((((((((.---------........))))))))))..)))).))).. ( -28.80) >consensus CGUGU________GUGUGGUGAGGUUUACAAACGUAUAUCCCCUAU_GUAUAUAAACAUACACCACAUUAUACAGUGAUAUAGAGAUGUUUUUGUGUGUGGUGAAGGUGAGUGCG ((..(........(((..(((.((..(((....)))....)).)))..))).....(((.((((((((...(((..(((((....)))))..)))))))))))...))).)..)) (-17.96 = -18.90 + 0.94)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:03:57 2006