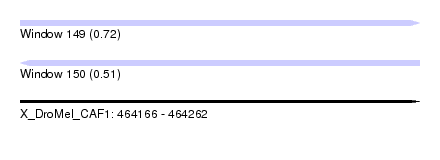

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 464,166 – 464,262 |
| Length | 96 |
| Max. P | 0.720882 |

| Location | 464,166 – 464,262 |
|---|---|
| Length | 96 |
| Sequences | 5 |
| Columns | 99 |
| Reading direction | forward |
| Mean pairwise identity | 83.25 |
| Mean single sequence MFE | -28.78 |
| Consensus MFE | -20.86 |
| Energy contribution | -20.90 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.80 |
| Structure conservation index | 0.72 |
| SVM decision value | 0.40 |
| SVM RNA-class probability | 0.720882 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 464166 96 + 22224390 ACUCAAUGGCUCUUUGUGGAUCCAUUGUGACUGGCAUUGCAGCCUAGCUGUCGGAAUAGCCGAA---UAUCCCAAUAGCCGAAAAGUCACAUGCAGAGC .((((((((.(((....))).)))))(((((((((((((((((...))))((((.....)))).---.....)))).)))....)))))).....))). ( -30.00) >DroSec_CAF1 66317 99 + 1 ACUCAAUGGCUCUUUGUGGAUCCAUUGUGACUGGCAUUGCAGCCGAGCUGUCGGAAAAGCCGAAUAAUAGCCCAAUAGCCGAAAAGUCACAUGCAGAGC .((((((((.(((....))).)))))(((((((((((((((((...))))((((.....)))).........)))).)))....)))))).....))). ( -30.00) >DroSim_CAF1 45359 96 + 1 ACUCAAUGGCUCUUUGUGGAUCCAUUGUGACUGGCAUUGCAGCCGAGCUGUCGGAAAAGCCGAA---UAGCCCAAUAGCCAAAAAGUCACAUGCAGAGC .((((((((.(((....))).)))))(((((((((......)))(.((((((((.....))).)---)))).)...........)))))).....))). ( -31.90) >DroEre_CAF1 78932 79 + 1 ACUCG-UGGGUCUUUGUGGAUCCAUUGUGACUGUCAUUGCAGCCGAGCUGUCGGAACA-------------------GCCGAAAAGUCACAUGCAGAGC .((((-(((((((....))))))))(((((((.......((((...))))((((....-------------------.))))..)))))))....))). ( -28.30) >DroYak_CAF1 65005 79 + 1 ACUCG-UGACUCUUUGUGGAUUCAUUGUGACUGUCAUUGCAGCCGAGCUGUCGGAAUA-------------------GCCGAGAAGUCACAUGCAGAGC .((((-(((.(((....))).))))(((((((.((....((((...))))((((....-------------------.)))))))))))))....))). ( -23.70) >consensus ACUCAAUGGCUCUUUGUGGAUCCAUUGUGACUGGCAUUGCAGCCGAGCUGUCGGAAAAGCCGAA___UA_CCCAAUAGCCGAAAAGUCACAUGCAGAGC .((((((((.(((....))).)))))((((((.......((((...))))((((........................))))..)))))).....))). (-20.86 = -20.90 + 0.04)

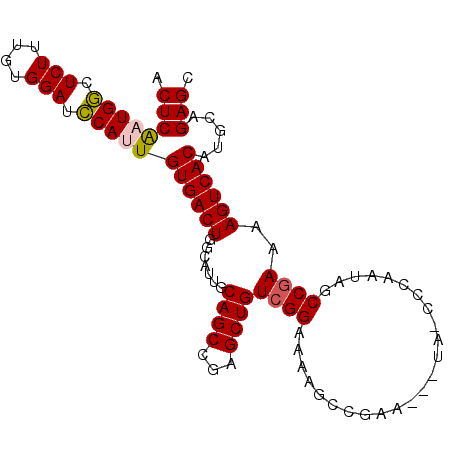
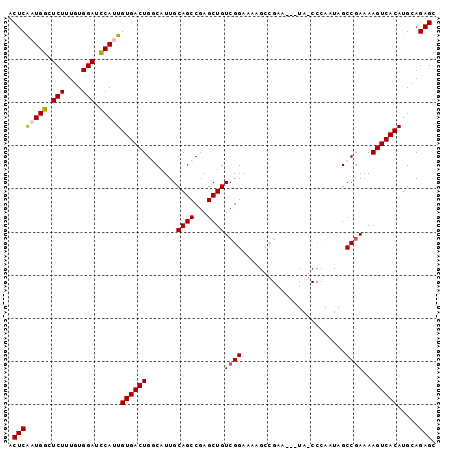
| Location | 464,166 – 464,262 |
|---|---|
| Length | 96 |
| Sequences | 5 |
| Columns | 99 |
| Reading direction | reverse |
| Mean pairwise identity | 83.25 |
| Mean single sequence MFE | -27.74 |
| Consensus MFE | -18.58 |
| Energy contribution | -18.10 |
| Covariance contribution | -0.48 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.63 |
| Structure conservation index | 0.67 |
| SVM decision value | -0.05 |
| SVM RNA-class probability | 0.508646 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 464166 96 - 22224390 GCUCUGCAUGUGACUUUUCGGCUAUUGGGAUA---UUCGGCUAUUCCGACAGCUAGGCUGCAAUGCCAGUCACAAUGGAUCCACAAAGAGCCAUUGAGU ((((.....((((((....((((.((((((((---......)))))))).)))).(((......)))))))))(((((.((......)).))))))))) ( -33.60) >DroSec_CAF1 66317 99 - 1 GCUCUGCAUGUGACUUUUCGGCUAUUGGGCUAUUAUUCGGCUUUUCCGACAGCUCGGCUGCAAUGCCAGUCACAAUGGAUCCACAAAGAGCCAUUGAGU ((((.....((((((...........(((((.....((((.....)))).)))))(((......)))))))))(((((.((......)).))))))))) ( -29.60) >DroSim_CAF1 45359 96 - 1 GCUCUGCAUGUGACUUUUUGGCUAUUGGGCUA---UUCGGCUUUUCCGACAGCUCGGCUGCAAUGCCAGUCACAAUGGAUCCACAAAGAGCCAUUGAGU ((((.....((((((...........(((((.---.((((.....)))).)))))(((......)))))))))(((((.((......)).))))))))) ( -31.30) >DroEre_CAF1 78932 79 - 1 GCUCUGCAUGUGACUUUUCGGC-------------------UGUUCCGACAGCUCGGCUGCAAUGACAGUCACAAUGGAUCCACAAAGACCCA-CGAGU ((((....(((((((..(((..-------------------(((.((((....))))..))).))).))))))).(((.((......)).)))-.)))) ( -22.90) >DroYak_CAF1 65005 79 - 1 GCUCUGCAUGUGACUUCUCGGC-------------------UAUUCCGACAGCUCGGCUGCAAUGACAGUCACAAUGAAUCCACAAAGAGUCA-CGAGU ((((....(((((((((...((-------------------....((((....))))..))...)).))))))).(((.((......)).)))-.)))) ( -21.30) >consensus GCUCUGCAUGUGACUUUUCGGCUAUUGGG_UA___UUCGGCUAUUCCGACAGCUCGGCUGCAAUGCCAGUCACAAUGGAUCCACAAAGAGCCAUUGAGU ((((....(((((((..((((........................))))((((...)))).......))))))).(((.((......)).)))..)))) (-18.58 = -18.10 + -0.48)

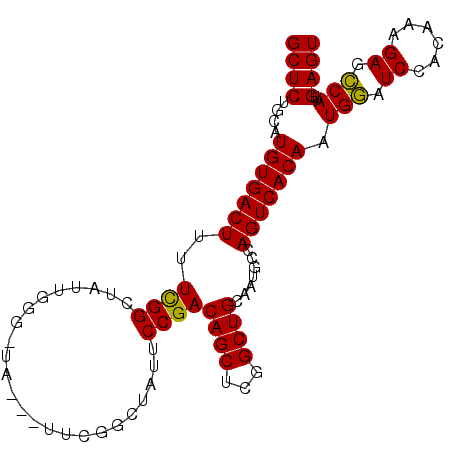
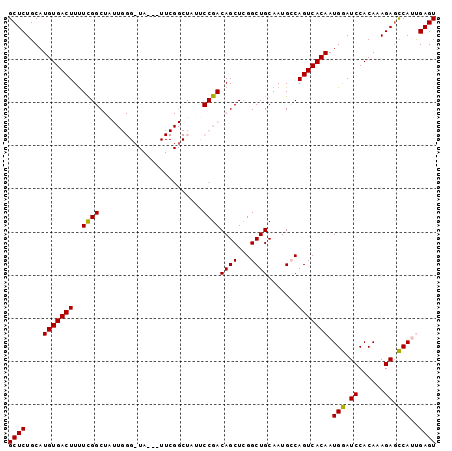
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:34:38 2006