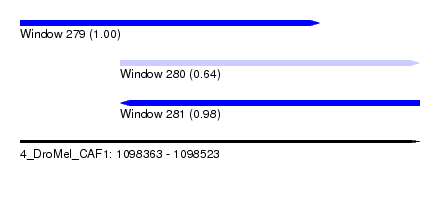
| Sequence ID | 4_DroMel_CAF1 |
|---|---|
| Location | 1,098,363 – 1,098,523 |
| Length | 160 |
| Max. P | 0.999676 |

| Location | 1,098,363 – 1,098,483 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 96.67 |
| Mean single sequence MFE | -28.24 |
| Consensus MFE | -26.76 |
| Energy contribution | -27.20 |
| Covariance contribution | 0.44 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.36 |
| Structure conservation index | 0.95 |
| SVM decision value | 3.87 |
| SVM RNA-class probability | 0.999676 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
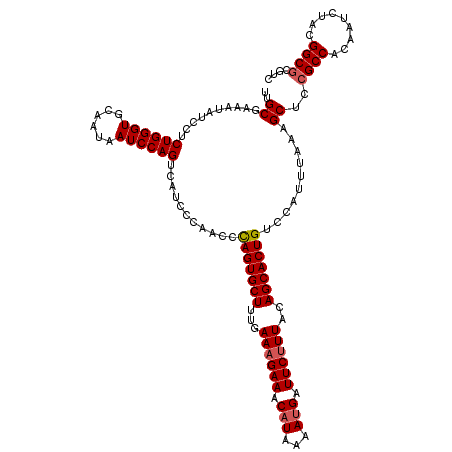
>4_DroMel_CAF1 1098363 120 + 1281640 UUGCAAAAUAUCCUCUGGGUGCAAUAAUCCAGUCAUCCCAACCUAGUGCUUUGAAAGAAACAUAAAAAGAUUCUUUACAGCACUGUCCAUUUAAAGCUCCGCCACAAUCUACGGCGCGUC ..((..........((((((......))))))...........(((((((...((((((.(.......).))))))..)))))))..........))..((((.........)))).... ( -26.10) >DroSec_CAF1 13080 120 + 1 UUGCGAAAUAUCCUCUGGGUGCAAUAAUCCAGUCAUCCCAACCCAGUGCUUUGAAAGAAACAUAAAAUGAUUCUUUACAGCACUGUCCAUUUAAAGCUCCGCCACAAUCUACGGCGCGUC ..((.((((.....((((((......))))))...........(((((((...((((((.(((...))).))))))..)))))))...))))...))..((((.........)))).... ( -29.60) >DroSim_CAF1 3063 120 + 1 UUGCGAAAUAUCCUCUGGGUGCAAUAAUCCAGUCAUCCCAACCCAGUGCUUUGAAAGAAACAUAAAAUGAUUCUUUACAGCACUGUCCAUUUAAAGCUCCGCCACAAUCUACGGCGCGUC ..((.((((.....((((((......))))))...........(((((((...((((((.(((...))).))))))..)))))))...))))...))..((((.........)))).... ( -29.60) >DroEre_CAF1 12520 120 + 1 UCGCGAAAUAUCCCCUGGGUGCAAUAAUCCAGUCAUCCCAACCCAGUGCUUUGAAAGAAACAUAAAAUGAUUCUUUACAGCACUGUCCAUUUAAAGCUCCGCCACAAUCUACGGCGCGUC ..((.((((.....((((((......))))))...........(((((((...((((((.(((...))).))))))..)))))))...))))...))..((((.........)))).... ( -30.10) >DroYak_CAF1 4313 120 + 1 UCGCGAAAUAUCCCCUGGGUGCAAUAAUCCAGUCAUCCCAACCCAGUGCUUUGAAAGAAACAUAAAAUGAUUCCUUACAGCACUGCCCAUUUAAAGCUCCGCCACAAUCUAUGGCUCGUC ..((((........((((((......))))))...........(((((((.(((..(((.(((...))).))).))).)))))))...............((((.......)))))))). ( -25.80) >consensus UUGCGAAAUAUCCUCUGGGUGCAAUAAUCCAGUCAUCCCAACCCAGUGCUUUGAAAGAAACAUAAAAUGAUUCUUUACAGCACUGUCCAUUUAAAGCUCCGCCACAAUCUACGGCGCGUC ..((..........((((((......))))))...........(((((((...((((((.(((...))).))))))..)))))))..........))..((((.........)))).... (-26.76 = -27.20 + 0.44)

| Location | 1,098,403 – 1,098,523 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 95.42 |
| Mean single sequence MFE | -28.57 |
| Consensus MFE | -23.84 |
| Energy contribution | -24.52 |
| Covariance contribution | 0.68 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.45 |
| Structure conservation index | 0.83 |
| SVM decision value | 0.23 |
| SVM RNA-class probability | 0.642972 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
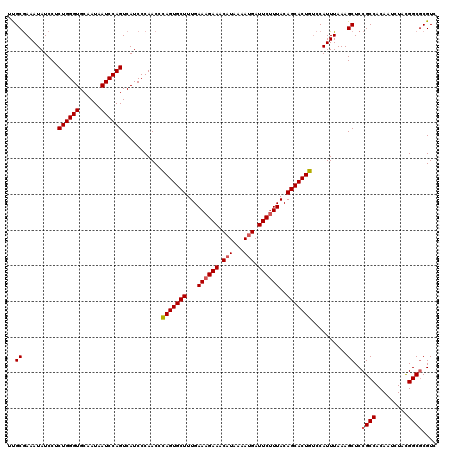
>4_DroMel_CAF1 1098403 120 + 1281640 ACCUAGUGCUUUGAAAGAAACAUAAAAAGAUUCUUUACAGCACUGUCCAUUUAAAGCUCCGCCACAAUCUACGGCGCGUCGCCUAACACGCCUUGUUCUGGAUGAUUCUGUAUGAAGUAC .....(((((((.((((((.(.......).))))))((((.(..(((((....(((....(((.........)))((((........)))))))....)))))..).))))..))))))) ( -26.10) >DroSec_CAF1 13120 120 + 1 ACCCAGUGCUUUGAAAGAAACAUAAAAUGAUUCUUUACAGCACUGUCCAUUUAAAGCUCCGCCACAAUCUACGGCGCGUCGCCUGACACGCCUUUUUCUGUAUGAUUCUGUAUGAAGUAC ...(((((((...((((((.(((...))).))))))..)))))))..........(((.((.((.(((((((((((.(((....))).)))).......))).)))).))..)).))).. ( -29.11) >DroSim_CAF1 3103 120 + 1 ACCCAGUGCUUUGAAAGAAACAUAAAAUGAUUCUUUACAGCACUGUCCAUUUAAAGCUCCGCCACAAUCUACGGCGCGUCGCCUGACACGCCUUGUUCUGGAUGAUUCUGUAUGAAGUAC ...(((((((...((((((.(((...))).))))))..)))))))..(((......(((((..((((.....((((.(((....))).))))))))..)))).).......)))...... ( -31.02) >DroEre_CAF1 12560 120 + 1 ACCCAGUGCUUUGAAAGAAACAUAAAAUGAUUCUUUACAGCACUGUCCAUUUAAAGCUCCGCCACAAUCUACGGCGCGUCGCCUAACGCGCCUUGUUUUAGAUGAUUCUGUAUGAAGUAC ...(((((((...((((((.(((...))).))))))..)))))))..((((((((((...(..........)(((((((......)))))))..)))))))))).(((.....))).... ( -35.00) >DroYak_CAF1 4353 120 + 1 ACCCAGUGCUUUGAAAGAAACAUAAAAUGAUUCCUUACAGCACUGCCCAUUUAAAGCUCCGCCACAAUCUAUGGCUCGUCGCCUAACGCGCCUUGUUCUUGAUGAUUCUGUAUGAAGUAC ...(((((((.(((..(((.(((...))).))).))).)))))))..........(((.(((((.......))))((((((...((((.....))))..))))))........).))).. ( -21.60) >consensus ACCCAGUGCUUUGAAAGAAACAUAAAAUGAUUCUUUACAGCACUGUCCAUUUAAAGCUCCGCCACAAUCUACGGCGCGUCGCCUAACACGCCUUGUUCUGGAUGAUUCUGUAUGAAGUAC ...(((((((...((((((.(((...))).))))))..)))))))..((((((.(((...(((.........)))((((........))))...))).)))))).(((.....))).... (-23.84 = -24.52 + 0.68)

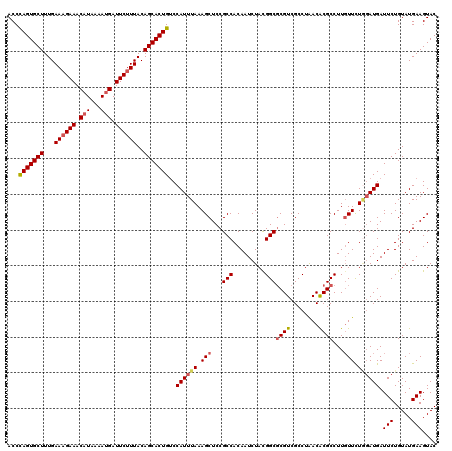
| Location | 1,098,403 – 1,098,523 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 95.42 |
| Mean single sequence MFE | -35.28 |
| Consensus MFE | -33.10 |
| Energy contribution | -33.06 |
| Covariance contribution | -0.04 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.62 |
| Structure conservation index | 0.94 |
| SVM decision value | 1.99 |
| SVM RNA-class probability | 0.984846 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
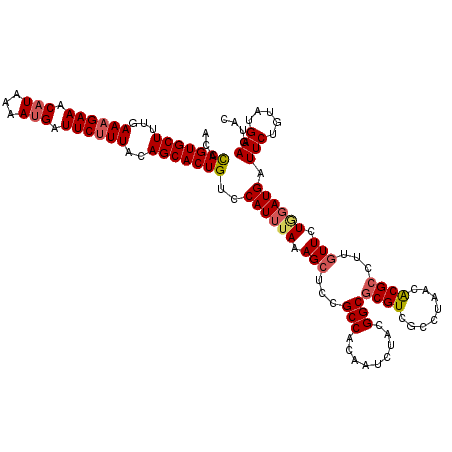
>4_DroMel_CAF1 1098403 120 - 1281640 GUACUUCAUACAGAAUCAUCCAGAACAAGGCGUGUUAGGCGACGCGCCGUAGAUUGUGGCGGAGCUUUAAAUGGACAGUGCUGUAAAGAAUCUUUUUAUGUUUCUUUCAAAGCACUAGGU ....(((((..(((.((..(((.(((..((((((((....))))))))...).)).)))..))..)))..))))).((((((..((((((.(.......).))))))...)))))).... ( -31.50) >DroSec_CAF1 13120 120 - 1 GUACUUCAUACAGAAUCAUACAGAAAAAGGCGUGUCAGGCGACGCGCCGUAGAUUGUGGCGGAGCUUUAAAUGGACAGUGCUGUAAAGAAUCAUUUUAUGUUUCUUUCAAAGCACUGGGU ...((((...((.((((.(((.......((((((((....))))))))))))))).))..))))...........(((((((..((((((.(((...))).))))))...)))))))... ( -38.61) >DroSim_CAF1 3103 120 - 1 GUACUUCAUACAGAAUCAUCCAGAACAAGGCGUGUCAGGCGACGCGCCGUAGAUUGUGGCGGAGCUUUAAAUGGACAGUGCUGUAAAGAAUCAUUUUAUGUUUCUUUCAAAGCACUGGGU ....(((((..(((.((..(((.(((..((((((((....))))))))...).)).)))..))..)))..)))))(((((((..((((((.(((...))).))))))...)))))))... ( -38.10) >DroEre_CAF1 12560 120 - 1 GUACUUCAUACAGAAUCAUCUAAAACAAGGCGCGUUAGGCGACGCGCCGUAGAUUGUGGCGGAGCUUUAAAUGGACAGUGCUGUAAAGAAUCAUUUUAUGUUUCUUUCAAAGCACUGGGU ...((((.((((.....(((((......((((((((....)))))))).)))))))))..))))...........(((((((..((((((.(((...))).))))))...)))))))... ( -37.90) >DroYak_CAF1 4353 120 - 1 GUACUUCAUACAGAAUCAUCAAGAACAAGGCGCGUUAGGCGACGAGCCAUAGAUUGUGGCGGAGCUUUAAAUGGGCAGUGCUGUAAGGAAUCAUUUUAUGUUUCUUUCAAAGCACUGGGU ...((((.(((((..((.....))....(((.((((....)))).))).....)))))..))))...........(((((((..((((((.(((...))).))))))...)))))))... ( -30.30) >consensus GUACUUCAUACAGAAUCAUCCAGAACAAGGCGUGUUAGGCGACGCGCCGUAGAUUGUGGCGGAGCUUUAAAUGGACAGUGCUGUAAAGAAUCAUUUUAUGUUUCUUUCAAAGCACUGGGU ...((((...((.((((.(....)....((((((((....))))))))...)))).))..))))...........(((((((..((((((.(((...))).))))))...)))))))... (-33.10 = -33.06 + -0.04)

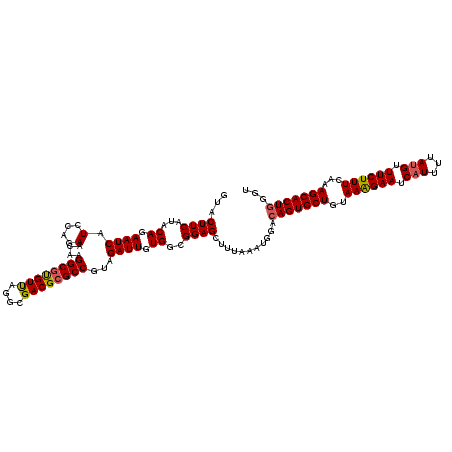
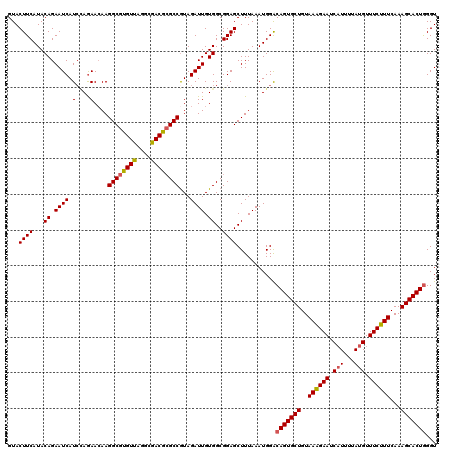
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:38:12 2006