
| Sequence ID | 4_DroMel_CAF1 |
|---|---|
| Location | 1,070,710 – 1,070,817 |
| Length | 107 |
| Max. P | 0.586635 |

| Location | 1,070,710 – 1,070,817 |
|---|---|
| Length | 107 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 80.24 |
| Mean single sequence MFE | -30.89 |
| Consensus MFE | -21.52 |
| Energy contribution | -21.22 |
| Covariance contribution | -0.30 |
| Combinations/Pair | 1.18 |
| Mean z-score | -1.26 |
| Structure conservation index | 0.70 |
| SVM decision value | 0.11 |
| SVM RNA-class probability | 0.586635 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>4_DroMel_CAF1 1070710 107 + 1281640 AACUCUUCUUUGAUGUCGUAAUUGUCCGAAAAACGCGUACAGGCUGCUGGUGCAGCCAUCGCGAUGGCACACUUAAGAAAAUCUAUUGCGACUAAAGAUCUUCAACU------------- ......((((((..((((((((((((((.....((((....((((((....))))))..)))).))).)))....((.....)))))))))))))))).........------------- ( -30.80) >DroVir_CAF1 2598 101 + 1 AGCUCUUCCUUGAUAUCGUAAUUGUCCGAAAAACGCGUACAGGCUGCUGGAGCAGCCAUCGCGAUGGCACACUUGAGAAAAUCUAUUGUGUGUGUG--UCAAU----------------- .........(((((((.................((((....((((((....))))))..))))...((((((...((.....))...)))))))))--)))).----------------- ( -31.90) >DroGri_CAF1 2089 103 + 1 AGUUCCUCUUUGAUAUCGUAAUUGUCCGAAAAACGCGUACAGGCUGCUGGAGCAGCCAUCGCGAUGGCACACUUGAGAAAAUCUAUCGUGCGUGUGUGUCUAU----------------- ...........((((.......)))).((...((((((((.((((((....)))))).....(((((...............)))))))))))))...))...----------------- ( -29.56) >DroWil_CAF1 3824 118 + 1 AGCUCUUCCUUGAUGUCGUAAUUGUCCGAAAAACGCGUACAGGCUGCUGGAGCAGCCAUCGCGAUGGCGCACUUGAGAAAAUCUAUGACGGCAGUAAAAUUCCAGCAAUGU--UGUGGUA ...((((...((.((((((..((....))....((((....((((((....))))))..))))))))))))...)))).....(((.((((((...............)))--))).))) ( -28.76) >DroMoj_CAF1 3853 103 + 1 AGCUCCUCUUUGAUGUCGUAAUUGUCCGAAAAACGCGUACAGGCUGCUGGAGCAGCCAUCGCGAUAGCACACUUGAGAAAAUCUAUUGUGUGUGUGUAUCAAU----------------- .........(((((((((........)))....((((....((((((....))))))..))))...((((((...((.....))...))))))...)))))).----------------- ( -31.00) >DroPer_CAF1 1726 117 + 1 AGCUCUUCUUUGAUGUCGUAAUUGUCCGAAAAACGCGUACAGGCUGCUGGAGCAGCCAUCGCGAUGGCACACUUAAGAAAAUCUAUUUCGGCAAUAAAUUGUCAGCAAGGCGUUGUA--- .((.(((..((((((...(.((((.((((((..((((....((((((....))))))..))))............((.....)).)))))))))).)..)))))).)))))......--- ( -33.30) >consensus AGCUCUUCUUUGAUGUCGUAAUUGUCCGAAAAACGCGUACAGGCUGCUGGAGCAGCCAUCGCGAUGGCACACUUGAGAAAAUCUAUUGCGGCUGUAAAUCUAC_________________ ..(((.....((.((((((...(((.((.......)).)))((((((....))))))...)))))).)).....)))........................................... (-21.52 = -21.22 + -0.30)

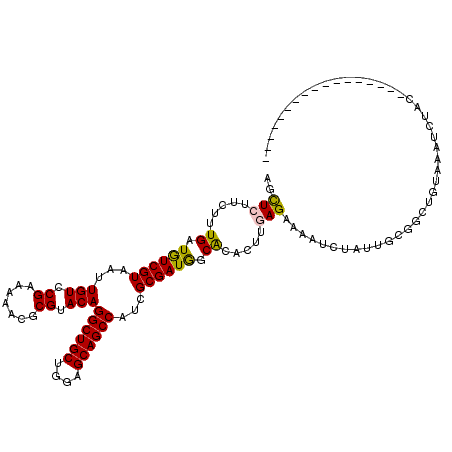
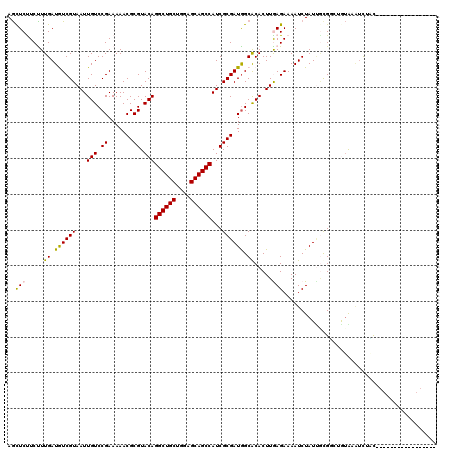
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:37:47 2006