
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 26,525,619 – 26,525,731 |
| Length | 112 |
| Max. P | 0.937431 |

| Location | 26,525,619 – 26,525,731 |
|---|---|
| Length | 112 |
| Sequences | 6 |
| Columns | 118 |
| Reading direction | forward |
| Mean pairwise identity | 80.27 |
| Mean single sequence MFE | -27.55 |
| Consensus MFE | -19.78 |
| Energy contribution | -20.39 |
| Covariance contribution | 0.61 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.18 |
| Structure conservation index | 0.72 |
| SVM decision value | 1.09 |
| SVM RNA-class probability | 0.913347 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 26525619 112 + 27905053 CU-GGCCAUCCUCUUCCGCCGGCGAUUGCAA-AAUGCUCA-ACAUU-UUAUUGAAAUUCAAUUAAAGCCGUAUUUCAAUAAUUGGUCACGCAUGGUGCGCAUUUCCGUGCC-AG-UCC ((-(((.............(((.(((((((.-.((((...-((...-((((((((((..............))))))))))...))...))))..)))).))).))).)))-))-... ( -27.28) >DroVir_CAF1 144646 101 + 1 ----------AUUUUU-C-GAGGAAAUGCAA-AAUGCUGA-ACAUU-UUAUUGAAAUUCAAUUAAAGUCGUAUUUCAAUAAUUGGACACGCAUGGUGCGCAUUUCCGUGC--UGCUCG ----------......-(-(((.....(((.-.((((((.-.((..-((((((((((.(.((....)).).)))))))))).))..)).))))..)))((((....))))--..)))) ( -26.20) >DroGri_CAF1 255332 106 + 1 -CGAAC---AAA-----G-UAGGAAAUGCAA-AAUGCUGAAACAUU-UUAUUGAAAUUCAAUUAAAGUCGUAUUUCAAUAAUUGGACACGCAUGGUGCCCAUUUCCGUGCACAGUUCG -(((((---...-----(-(((((((((((.-.((((((...((..-((((((((((.(.((....)).).)))))))))).))..)).))))..)))..)))))).)))...))))) ( -29.00) >DroWil_CAF1 122832 111 + 1 ---UGCCAGU-UUUUCGGAUAGGAAAUGCAA-AAUGCUAA-ACAUUUUUAUUGAAAUUCAAUUAAAGUCGUAUUUCAAUAAUUGGUCACGCAUGGUGCGCAUUUCCGUGCUCCG-GAC ---.......-..((((((..((((((((..-.((((...-((....((((((((((.(.((....)).).))))))))))...))...)))).....))))))))....))))-)). ( -30.00) >DroMoj_CAF1 146214 109 + 1 -A-GAC---AAUUUUU-G-GAGGAAAUGCGA-AAUGCUGAAACAUU-UUAUUGAAAUUCAAUUAAAGUCGUAUUUCAAUAAUUGGCCACGCAUGGUGCGCAUUUCCGUGCCGUGCUCG -.-...---.......-(-(.(((((((((.-.((((((...((..-((((((((((.(.((....)).).)))))))))).))..)).))))....)))))))))...))....... ( -30.50) >DroAna_CAF1 111161 108 + 1 C------UUCCUUUUCCGCUACCGAAUGCAAAAAUGCUCA-ACAUU-UUAUUGAAAUUCAAUUAAAGUCGUAUUUCAAUAAUUGGUCACGCAUGGUGCGCAUUUCCGUGCC-AG-UCC .------..........((.(((((..(((....)))...-.....-((((((((((.(.((....)).).)))))))))))))))...)).(((..((......))..))-).-... ( -22.30) >consensus _____C___AAUUUUC_G_UAGGAAAUGCAA_AAUGCUGA_ACAUU_UUAUUGAAAUUCAAUUAAAGUCGUAUUUCAAUAAUUGGUCACGCAUGGUGCGCAUUUCCGUGCC_AG_UCC .....................((((((((....((((.....((...((((((((((..............)))))))))).)).....)))).....))))))))............ (-19.78 = -20.39 + 0.61)

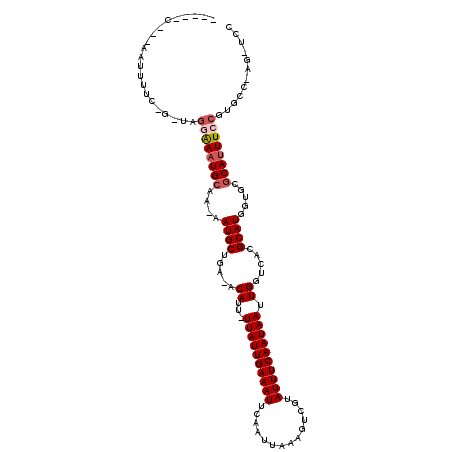

| Location | 26,525,619 – 26,525,731 |
|---|---|
| Length | 112 |
| Sequences | 6 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 80.27 |
| Mean single sequence MFE | -28.62 |
| Consensus MFE | -20.11 |
| Energy contribution | -21.00 |
| Covariance contribution | 0.89 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.47 |
| Structure conservation index | 0.70 |
| SVM decision value | 1.26 |
| SVM RNA-class probability | 0.937431 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 26525619 112 - 27905053 GGA-CU-GGCACGGAAAUGCGCACCAUGCGUGACCAAUUAUUGAAAUACGGCUUUAAUUGAAUUUCAAUAA-AAUGU-UGAGCAUU-UUGCAAUCGCCGGCGGAAGAGGAUGGCC-AG ...-((-(((.(((..((..(((..((((..(((...((((((((((.(((......))).))))))))))-...))-)..)))).-.))).))..))).(....)......)))-)) ( -32.50) >DroVir_CAF1 144646 101 - 1 CGAGCA--GCACGGAAAUGCGCACCAUGCGUGUCCAAUUAUUGAAAUACGACUUUAAUUGAAUUUCAAUAA-AAUGU-UCAGCAUU-UUGCAUUUCCUC-G-AAAAAU---------- ......--....(((((((((....((((.((..((.((((((((((.(((......))).))))))))))-..)).-.)))))).-.)))))))))..-.-......---------- ( -26.40) >DroGri_CAF1 255332 106 - 1 CGAACUGUGCACGGAAAUGGGCACCAUGCGUGUCCAAUUAUUGAAAUACGACUUUAAUUGAAUUUCAAUAA-AAUGUUUCAGCAUU-UUGCAUUUCCUA-C-----UUU---GUUCG- (((((.(((...((((((((((((.....))))))..((((((((((.(((......))).))))))))))-(((((....)))))-....))))))))-)-----...---)))))- ( -28.80) >DroWil_CAF1 122832 111 - 1 GUC-CGGAGCACGGAAAUGCGCACCAUGCGUGACCAAUUAUUGAAAUACGACUUUAAUUGAAUUUCAAUAAAAAUGU-UUAGCAUU-UUGCAUUUCCUAUCCGAAAA-ACUGGCA--- ...-((((....(((((((((....((((..(((...((((((((((.(((......))).))))))))))....))-)..)))).-.)))))))))..))))....-.......--- ( -30.60) >DroMoj_CAF1 146214 109 - 1 CGAGCACGGCACGGAAAUGCGCACCAUGCGUGGCCAAUUAUUGAAAUACGACUUUAAUUGAAUUUCAAUAA-AAUGUUUCAGCAUU-UCGCAUUUCCUC-C-AAAAAUU---GUC-U- .......((...(((((((((....((((..(((...((((((((((.(((......))).))))))))))-...)))...)))).-.))))))))).)-)-.......---...-.- ( -29.00) >DroAna_CAF1 111161 108 - 1 GGA-CU-GGCACGGAAAUGCGCACCAUGCGUGACCAAUUAUUGAAAUACGACUUUAAUUGAAUUUCAAUAA-AAUGU-UGAGCAUUUUUGCAUUCGGUAGCGGAAAAGGAA------G ..(-((-((..(((((((((((.....))..(((...((((((((((.(((......))).))))))))))-...))-)..)))))))))...))))).............------. ( -24.40) >consensus CGA_CU_GGCACGGAAAUGCGCACCAUGCGUGACCAAUUAUUGAAAUACGACUUUAAUUGAAUUUCAAUAA_AAUGU_UCAGCAUU_UUGCAUUUCCUA_C_GAAAAUG___G_____ ............(((((((((....((((.....((.((((((((((.(((......))).))))))))))...)).....))))...)))))))))..................... (-20.11 = -21.00 + 0.89)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:44:54 2006