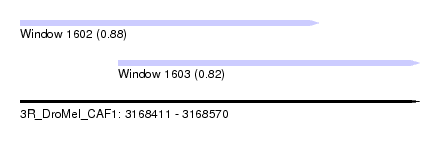

| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 3,168,411 – 3,168,570 |
| Length | 159 |
| Max. P | 0.881569 |

| Location | 3,168,411 – 3,168,530 |
|---|---|
| Length | 119 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 92.47 |
| Mean single sequence MFE | -27.27 |
| Consensus MFE | -21.85 |
| Energy contribution | -22.41 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.16 |
| Mean z-score | -2.34 |
| Structure conservation index | 0.80 |
| SVM decision value | 0.92 |
| SVM RNA-class probability | 0.881569 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 3168411 119 + 27905053 UAAAUUCAAAUUCUUGGCGUUGA-ACUGAUUAAAAAAGCGCUCGAACUUAUUUAGCCAUAUUCGAUGCGAAUCAGCUCUAAUUACCAUUAACAUUCUGGGCUGGCCAGCAUCUGAAUUUC .(((((((..(((..((((((..-............)))))).))).................(((((...(((((((...................)))))))...)))))))))))). ( -26.45) >DroSec_CAF1 16524 119 + 1 UAAAUUAAAUUUCUUUGCGUUGA-AUUGAUUAAAAAAGCGCCCCAACUUAUUUAGCCAUAUUCGAUGCGAAUCAGCUCUAAUUACCAUUAACAUUCUGGGCUGGCCAGCAUCUGAAUUUC ..........(((((((((((((-((((..((((.(((........))).))))..)).))))))))))))(((((((...................))))))).........))).... ( -23.81) >DroSim_CAF1 19250 119 + 1 UAAAUUCAAUUUCUUCGCGUUGA-AUUGAUUAAAAAAGCGCUCCAACUUAUUUAGCCAUAUUCGAUGCGAAUCAGCCCUAAUUACCAUUAACAUUCUGGGCUGGCCAGCAUCUGAAUUUC .(((((((.....((((((((((-((((..((((.(((........))).))))..)).))))))))))))(((((((...................)))))))........))))))). ( -32.61) >DroEre_CAF1 8193 120 + 1 UAAAUGCGAUUUCUUCUCAUUGAGAUUGAUUUAACCAGCGCUCCAACUUAUUUAGCCAUAUUCGAUGCGAUUCAGCCUUAAUUACCAUUAACAUUCUGGGCUGGCCAGCAUCUGAAUUUC (((((.(((((((........))))))))))))....(.(((...........))))..(((((((((...(((((((...................)))))))...))))).))))... ( -26.21) >consensus UAAAUUCAAUUUCUUCGCGUUGA_AUUGAUUAAAAAAGCGCUCCAACUUAUUUAGCCAUAUUCGAUGCGAAUCAGCCCUAAUUACCAUUAACAUUCUGGGCUGGCCAGCAUCUGAAUUUC .(((((((......(((((((((...((..((((.(((........))).))))..))...))))))))).(((((((...................)))))))........))))))). (-21.85 = -22.41 + 0.56)

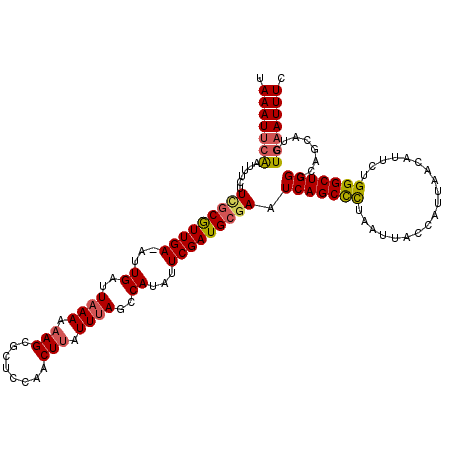
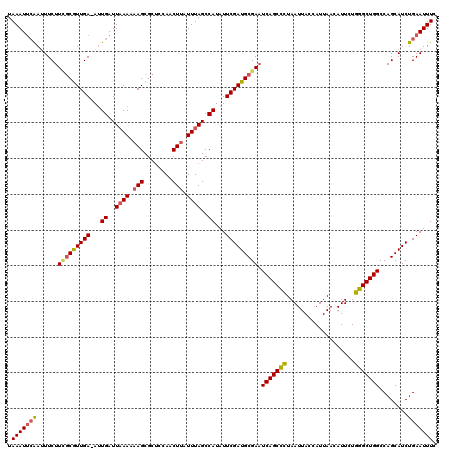
| Location | 3,168,450 – 3,168,570 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 95.42 |
| Mean single sequence MFE | -25.88 |
| Consensus MFE | -25.16 |
| Energy contribution | -24.54 |
| Covariance contribution | -0.62 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.65 |
| Structure conservation index | 0.97 |
| SVM decision value | 0.69 |
| SVM RNA-class probability | 0.824015 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 3168450 120 + 27905053 CUCGAACUUAUUUAGCCAUAUUCGAUGCGAAUCAGCUCUAAUUACCAUUAACAUUCUGGGCUGGCCAGCAUCUGAAUUUCCAAUUCGGCUUGAAACAAGCUUUCUUUGUGCGCAUCUUAA ..............((((((((((((((...(((((((...................)))))))...))))).)))..........((((((...)))))).....)))).))....... ( -27.01) >DroSec_CAF1 16563 120 + 1 CCCCAACUUAUUUAGCCAUAUUCGAUGCGAAUCAGCUCUAAUUACCAUUAACAUUCUGGGCUGGCCAGCAUCUGAAUUUCCAAUUCGGCCUUAAACAAGCUUUCUUUGUGCGCAUUUUAA ......(((.(((((((......(((((...(((((((...................)))))))...))))).((((.....)))))))..)))).)))..................... ( -23.31) >DroSim_CAF1 19289 120 + 1 CUCCAACUUAUUUAGCCAUAUUCGAUGCGAAUCAGCCCUAAUUACCAUUAACAUUCUGGGCUGGCCAGCAUCUGAAUUUCCAAUUCGGCCUGAAACAAGCUUUCUUUGUGCGCAUUUUAA ..........(((((((......(((((...(((((((...................)))))))...))))).((((.....)))))).)))))(((((.....)))))........... ( -27.31) >DroEre_CAF1 8233 120 + 1 CUCCAACUUAUUUAGCCAUAUUCGAUGCGAUUCAGCCUUAAUUACCAUUAACAUUCUGGGCUGGCCAGCAUCUGAAUUUCCGAUUCGGCAAGAAACAAGCUUUCUUUGUGCGCAUUUUAA ..............(((......(((((...(((((((...................)))))))...))))).(((((...))))))))((((((.....)))))).............. ( -25.91) >consensus CUCCAACUUAUUUAGCCAUAUUCGAUGCGAAUCAGCCCUAAUUACCAUUAACAUUCUGGGCUGGCCAGCAUCUGAAUUUCCAAUUCGGCCUGAAACAAGCUUUCUUUGUGCGCAUUUUAA ..............(((......(((((...(((((((...................)))))))...))))).(((((...)))))))).....(((((.....)))))........... (-25.16 = -24.54 + -0.62)

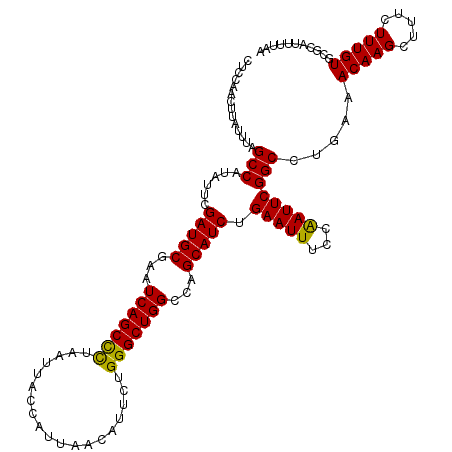
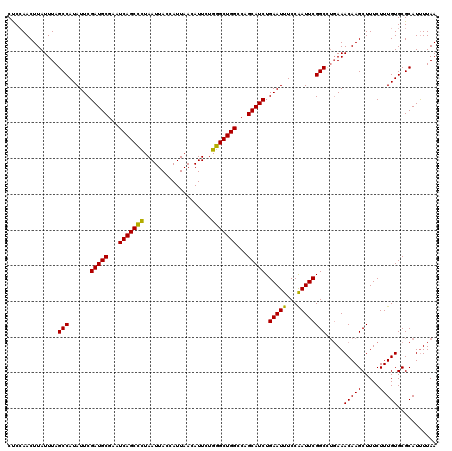
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:57:45 2006