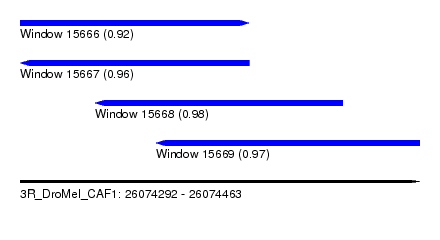

| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 26,074,292 – 26,074,463 |
| Length | 171 |
| Max. P | 0.983961 |

| Location | 26,074,292 – 26,074,390 |
|---|---|
| Length | 98 |
| Sequences | 6 |
| Columns | 103 |
| Reading direction | forward |
| Mean pairwise identity | 79.10 |
| Mean single sequence MFE | -22.53 |
| Consensus MFE | -15.77 |
| Energy contribution | -15.77 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.95 |
| Structure conservation index | 0.70 |
| SVM decision value | 1.11 |
| SVM RNA-class probability | 0.916306 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 26074292 98 + 27905053 CCAUAGAUUA--UUUUCUAAUGAUAAAAAGAAGAGCAAAAAAAA---CUGUGACAGCGGUGGGAUUCGAACCCACGCCCUUUCGGACUGGUGCCUAAAACCAG ((.((((...--...))))..........((((.((.......(---((((....)))))(((.......)))..)).)))).)).(((((.......))))) ( -21.70) >DroSec_CAF1 66025 97 + 1 CCUUAUCAUUCUUUUGUUUAUGUUAAAAAAAA-AAC--AAAAAA---AUGUGACAGCGGUGGGAUUCGAACCCACGCCCUUUCGGACUGGUGCCUAAAACCAG ((....((((.((((((((.(........).)-)))--)))).)---))).((.((.((((((........)).)))))).)))).(((((.......))))) ( -22.90) >DroSim_CAF1 67128 94 + 1 CCUUACCAUU--UUUGGUAAGGUUAAAA-AAA-AAC--AAAAAA---AUGUGACAGCGGUGGGAUUCGAACCCACGCCCUUUCGGACUGGUGCCUAAAACCAG ((((((((..--..))))))))......-...-...--......---.(.(((.((.((((((........)).)))))).))).)(((((.......))))) ( -27.30) >DroEre_CAF1 70489 83 + 1 CCA-------------AUGAUACAAAAA-AAG-AAC--AAAAAA---CUGUGACAGCGGUGGGAUUCGAACCCACGCCCUUUCGGACUGGUGCCUAAAACCAG ...-------------............-...-...--......---(((.((..(((.((((.......)))))))..)).))).(((((.......))))) ( -17.50) >DroYak_CAF1 68629 94 + 1 CUGCAGAUUU--GUUCUUAAUAAUAAAA-AAG-AAC--AAAAAA---CUGUGACAGCGGUGGGAUUCGAACCCACGCCCUUUCGGACUGGUGCCUAAAACCAG ..((((.(((--((((((..........-)))-)))--)))...---))))((.((.((((((........)).)))))).))...(((((.......))))) ( -25.70) >DroMoj_CAF1 93796 87 + 1 AUGAAUA-----UAUUCUAAACAUAAAAA------U--AAAAAAAUC---UGACAGCGGUGGGAUUCGAACCCACGCCCUUUCGGACUGGUGCCUAAAACCAG .......-----.................------.--.......((---(((.((.((((((........)).)))))).)))))(((((.......))))) ( -20.10) >consensus CCAUAGA_UU__UUUGCUAAUGAUAAAA_AAA_AAC__AAAAAA___CUGUGACAGCGGUGGGAUUCGAACCCACGCCCUUUCGGACUGGUGCCUAAAACCAG ..................................................(((.((.((((((........)).)))))).)))..(((((.......))))) (-15.77 = -15.77 + 0.00)



| Location | 26,074,292 – 26,074,390 |
|---|---|
| Length | 98 |
| Sequences | 6 |
| Columns | 103 |
| Reading direction | reverse |
| Mean pairwise identity | 79.10 |
| Mean single sequence MFE | -27.27 |
| Consensus MFE | -20.70 |
| Energy contribution | -20.70 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.40 |
| Structure conservation index | 0.76 |
| SVM decision value | 1.53 |
| SVM RNA-class probability | 0.961735 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 26074292 98 - 27905053 CUGGUUUUAGGCACCAGUCCGAAAGGGCGUGGGUUCGAAUCCCACCGCUGUCACAG---UUUUUUUUGCUCUUCUUUUUAUCAUUAGAAAA--UAAUCUAUGG (((......(((....((((....))))(((((.......))))).)))....)))---......................(((.(((...--...)))))). ( -23.60) >DroSec_CAF1 66025 97 - 1 CUGGUUUUAGGCACCAGUCCGAAAGGGCGUGGGUUCGAAUCCCACCGCUGUCACAU---UUUUUU--GUU-UUUUUUUUAACAUAAACAAAAGAAUGAUAAGG .........((((.(.((((....))))(((((.......))))).).)))).(((---((((((--(((-(............)))))))))))))...... ( -31.20) >DroSim_CAF1 67128 94 - 1 CUGGUUUUAGGCACCAGUCCGAAAGGGCGUGGGUUCGAAUCCCACCGCUGUCACAU---UUUUUU--GUU-UUU-UUUUAACCUUACCAAA--AAUGGUAAGG .........((((.(.((((....))))(((((.......))))).).))))....---......--...-...-......((((((((..--..)))))))) ( -31.10) >DroEre_CAF1 70489 83 - 1 CUGGUUUUAGGCACCAGUCCGAAAGGGCGUGGGUUCGAAUCCCACCGCUGUCACAG---UUUUUU--GUU-CUU-UUUUUGUAUCAU-------------UGG (((......(((....((((....))))(((((.......))))).)))....)))---......--...-...-............-------------... ( -22.10) >DroYak_CAF1 68629 94 - 1 CUGGUUUUAGGCACCAGUCCGAAAGGGCGUGGGUUCGAAUCCCACCGCUGUCACAG---UUUUUU--GUU-CUU-UUUUAUUAUUAAGAAC--AAAUCUGCAG (((......((((.(.((((....))))(((((.......))))).).)))).(((---...(((--(((-(((-..(....)..))))))--))).)))))) ( -31.70) >DroMoj_CAF1 93796 87 - 1 CUGGUUUUAGGCACCAGUCCGAAAGGGCGUGGGUUCGAAUCCCACCGCUGUCA---GAUUUUUUU--A------UUUUUAUGUUUAGAAUA-----UAUUCAU ...((((((((((...((((....))))(((((.......)))))........---.........--.------......)))))))))).-----....... ( -23.90) >consensus CUGGUUUUAGGCACCAGUCCGAAAGGGCGUGGGUUCGAAUCCCACCGCUGUCACAG___UUUUUU__GUU_UUU_UUUUAUCAUUAAAAAA__AA_UAUAAGG .........((((.(.((((....))))(((((.......))))).).))))................................................... (-20.70 = -20.70 + -0.00)


| Location | 26,074,324 – 26,074,430 |
|---|---|
| Length | 106 |
| Sequences | 6 |
| Columns | 116 |
| Reading direction | reverse |
| Mean pairwise identity | 82.05 |
| Mean single sequence MFE | -44.15 |
| Consensus MFE | -43.42 |
| Energy contribution | -43.42 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.27 |
| Structure conservation index | 0.98 |
| SVM decision value | 1.96 |
| SVM RNA-class probability | 0.983961 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 26074324 106 - 27905053 ACGUGUGGGUACAGGCAGCGUGGCCGAGCGGUCUAAGGCGCUGGUUUUAGGCACCAGUCCGAAAGGGCGUGGGUUCGAAUCCCACCGCUGUCACAG----------UUUUUUUUGC .............((((((((((.(((((.(((((((((....))))))))).(((((((....)))).))))))))....))).)))))))....----------.......... ( -43.40) >DroVir_CAF1 96650 108 - 1 CGUGGUUGAACAAGGCAGCGUGGCCGAGCGGUCUAAGGCGCUGGUUUUAGGCACCAGUCCGAAAGGGCGUGGGUUCGAAUCCCACCGCUGUCAAUG------UGUUUUUUUU--AU .((((..(((((.((((((((((.(((((.(((((((((....))))))))).(((((((....)))).))))))))....))).)))))))....------)))))...))--)) ( -45.50) >DroGri_CAF1 82966 107 - 1 CGAAAUGGAAAAAGGCAGCGUGGCCGAGCGGUCUAAGGCGCUGGUUUUAGGCACCAGUCCGAAAGGGCGUGGGUUCGAAUCCCACCGCUGUCAAAG------GUUU-UUUGC--AC .....((.(((((((((((((((.(((((.(((((((((....))))))))).(((((((....)))).))))))))....))).)))))))....------..))-))).)--). ( -43.60) >DroWil_CAF1 89989 113 - 1 CCCAACCACCAACGGCAGCGUGGCCGAGCGGUCUAAGGCGCUGGUUUUAGGCACCAGUCCGAAAGGGCGUGGGUUCGAAUCCCACCGCUGUCAAGUGUAAAAG---UUUUUUUUGU ......(((....((((((((((.(((((.(((((((((....))))))))).(((((((....)))).))))))))....))).)))))))..)))......---.......... ( -43.70) >DroMoj_CAF1 93820 103 - 1 CGAACAUGUGUGAGGCAGCGUGGCCGAGCGGUCUAAGGCGCUGGUUUUAGGCACCAGUCCGAAAGGGCGUGGGUUCGAAUCCCACCGCUGUCA----------GAUUUUUUU--A- .............((((((((((.(((((.(((((((((....))))))))).(((((((....)))).))))))))....))).))))))).----------.........--.- ( -43.40) >DroAna_CAF1 75818 108 - 1 CGAUGAGCACCAAGGCAGCGUGGCCGAGCGGUCUAAGGCGCUGGUUUUAGGCACCAGUCCGAAAGGGCGUGGGUUCGAAUCCCACCGCUGUCACAA------GG--UUUCCUUUUU ....(((.(((..((((((((((.(((((.(((((((((....))))))))).(((((((....)))).))))))))....))).)))))))....------))--))))...... ( -45.30) >consensus CGAAGUGGAAAAAGGCAGCGUGGCCGAGCGGUCUAAGGCGCUGGUUUUAGGCACCAGUCCGAAAGGGCGUGGGUUCGAAUCCCACCGCUGUCAAAG______GG__UUUUUU__AU .............((((((((((.(((((.(((((((((....))))))))).(((((((....)))).))))))))....))).)))))))........................ (-43.42 = -43.42 + -0.00)

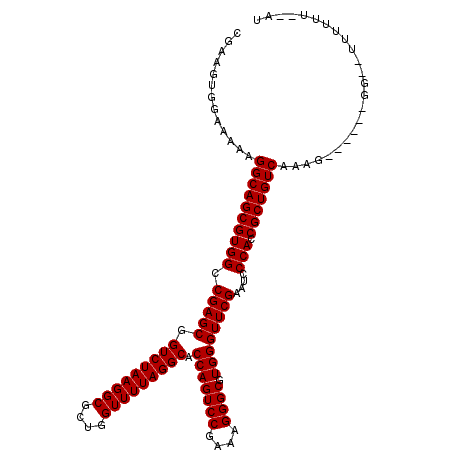

| Location | 26,074,350 – 26,074,463 |
|---|---|
| Length | 113 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 78.64 |
| Mean single sequence MFE | -46.87 |
| Consensus MFE | -31.22 |
| Energy contribution | -31.22 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.53 |
| Structure conservation index | 0.67 |
| SVM decision value | 1.65 |
| SVM RNA-class probability | 0.969997 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 26074350 113 - 27905053 A--AG--GCUC-GC--UAUGUUGAAAUUGCGCAUUUGACAACGUGUGGGUACAGGCAGCGUGGCCGAGCGGUCUAAGGCGCUGGUUUUAGGCACCAGUCCGAAAGGGCGUGGGUUCGAAU .--..--((((-((--(((((((....(((.(((.((....)).))).)))....)))))))).))))).(((((((((....))))))))).(((((((....)))).)))........ ( -46.70) >DroVir_CAF1 96678 119 - 1 GACGGUACUGU-GCUGUGCCUUCAAGUUGCACAUUUGGCACGUGGUUGAACAAGGCAGCGUGGCCGAGCGGUCUAAGGCGCUGGUUUUAGGCACCAGUCCGAAAGGGCGUGGGUUCGAAU ((.(((((...-...))))).))..(((((...((..((.....))..))....))))).....(((((.(((((((((....))))))))).(((((((....)))).))))))))... ( -47.30) >DroPse_CAF1 70616 116 - 1 U--GG--GCUCUGCCGUGCGCUUAAGUAGUGCUUUUGGCACUCAACGGUCAGAGGCAGCGUGGCCGAGCGGUCUAAGGCGCUGGUUUUAGGCACCAGUCCGAAAGGGCGUGGGUUCGAAU .--..--((((.((((((((((.....)))(((((((((........))))))))).))))))).)))).(((((((((....))))))))).(((((((....)))).)))........ ( -52.00) >DroGri_CAF1 82993 117 - 1 A--UGUUGCUU-GCUGUGAAUUCAAAUAGCACAUUUGACACGAAAUGGAAAAAGGCAGCGUGGCCGAGCGGUCUAAGGCGCUGGUUUUAGGCACCAGUCCGAAAGGGCGUGGGUUCGAAU (--((((((((-(((((........))))).(((((......))))).....)))))))))...(((((.(((((((((....))))))))).(((((((....)))).))))))))... ( -46.10) >DroWil_CAF1 90022 112 - 1 A--A---GCACCGA--AGUG-UUAUAUUGAACAUUGGACACCCAACCACCAACGGCAGCGUGGCCGAGCGGUCUAAGGCGCUGGUUUUAGGCACCAGUCCGAAAGGGCGUGGGUUCGAAU .--(---((((...--.)))-))...((((((.((((....))))((((...((((......)))).(..(((((((((....)))))))))..).((((....)))))))))))))).. ( -43.10) >DroAna_CAF1 75846 113 - 1 C--UA--UGGG-GC--UUAGUUCAAGGUGCGCAUUUGACACGAUGAGCACCAAGGCAGCGUGGCCGAGCGGUCUAAGGCGCUGGUUUUAGGCACCAGUCCGAAAGGGCGUGGGUUCGAAU (--((--(((.-((--((((..(..(((((.((((......)))).)))))....((((((((((....))))....)))))))..)))))).)).((((....))))))))........ ( -46.00) >consensus A__AG__GCUC_GC__UGAGUUCAAAUUGCACAUUUGACACGAAAUGGACAAAGGCAGCGUGGCCGAGCGGUCUAAGGCGCUGGUUUUAGGCACCAGUCCGAAAGGGCGUGGGUUCGAAU .....................................................(((......)))((((.(((((((((....))))))))).(((((((....)))).))))))).... (-31.22 = -31.22 + -0.00)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:42:43 2006