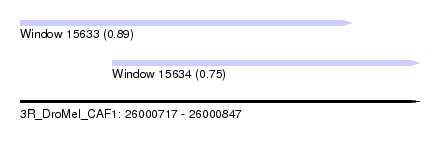
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 26,000,717 – 26,000,847 |
| Length | 130 |
| Max. P | 0.892224 |

| Location | 26,000,717 – 26,000,825 |
|---|---|
| Length | 108 |
| Sequences | 3 |
| Columns | 108 |
| Reading direction | forward |
| Mean pairwise identity | 96.30 |
| Mean single sequence MFE | -23.50 |
| Consensus MFE | -21.85 |
| Energy contribution | -21.63 |
| Covariance contribution | -0.22 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.25 |
| Structure conservation index | 0.93 |
| SVM decision value | 0.97 |
| SVM RNA-class probability | 0.892224 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
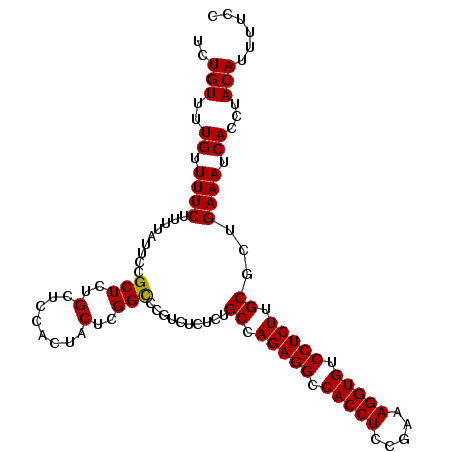
>3R_DroMel_CAF1 26000717 108 + 27905053 UCUGUUUUGUUUUCUUUUAUUUUGCUCUGCUUCACUACUCGGUGCGUCUCUCUGCCAGAGGCCACCUCCGAAAGGUGUCCUCUUGCGCUGAAAUCACCUACAUUUUCC ........................................((((.((.((..(((.(((((.(((((.....))))).))))).)))..)).)))))).......... ( -25.50) >DroSec_CAF1 52753 108 + 1 UCUGUUUUGUUUUCUUUUAUUCCGCUCUGCUCCACUACUCGGCCCGUCUCUCUGCCAGAGGCCACCUCCGAAAGGUGUCCUCUUGCGCUGAAAUCACCUACAUUUUCC ..(((..((.((((.........(((..(........)..)))..........((.(((((.(((((.....))))).))))).))...)))).))...)))...... ( -22.10) >DroSim_CAF1 53186 108 + 1 UCUGUUUUGUUUUCUUUUAUUCCGCUCUGCUCCACUACUCGGCCCGUCUCUCAGCCAGAGGCCACCUCCGAAAGGUGUCCUCUUGCGCUGAAAUCACCUACAUUUUCC .......................(((..(........)..))).......(((((((((((.(((((.....))))).)))).)).)))))................. ( -22.90) >consensus UCUGUUUUGUUUUCUUUUAUUCCGCUCUGCUCCACUACUCGGCCCGUCUCUCUGCCAGAGGCCACCUCCGAAAGGUGUCCUCUUGCGCUGAAAUCACCUACAUUUUCC ..(((..((.((((.........(((..(........)..)))..........((.(((((.(((((.....))))).))))).))...)))).))...)))...... (-21.85 = -21.63 + -0.22)


| Location | 26,000,747 – 26,000,847 |
|---|---|
| Length | 100 |
| Sequences | 6 |
| Columns | 106 |
| Reading direction | forward |
| Mean pairwise identity | 77.16 |
| Mean single sequence MFE | -26.18 |
| Consensus MFE | -13.65 |
| Energy contribution | -14.48 |
| Covariance contribution | 0.83 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.94 |
| Structure conservation index | 0.52 |
| SVM decision value | 0.48 |
| SVM RNA-class probability | 0.752942 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
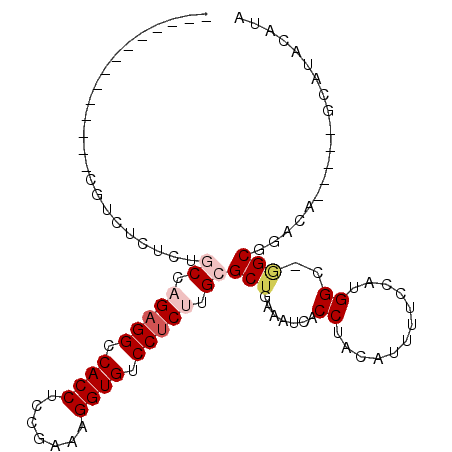
>3R_DroMel_CAF1 26000747 100 + 27905053 UUCACUACUCGGUGCGUCUCUCUGCCAGAGGCCACCUCCGAAAGGUGUCCUCUUGCGCUGAAAUCACCUACAUUUUCCUUGGC-GGCGGACA-----GCAUACAUA ...........(((((((.(.((((((((((.(((((.....))))).)))).......((((..........))))..))))-)).)))).-----))))..... ( -31.10) >DroPse_CAF1 46179 89 + 1 ---------------AUCUUUGCUCUUGAGGCCACCGCCGAAAGGUGUCCUCUUGCGCUGAAAUCACCUACAUU-CCCAUGGC-AGCACACAUUCAAGCAAACAUA ---------------...((((((...((((.((((.......)))).)))).((.((((..((..........-...))..)-))).))......)))))).... ( -22.32) >DroSec_CAF1 52783 100 + 1 UCCACUACUCGGCCCGUCUCUCUGCCAGAGGCCACCUCCGAAAGGUGUCCUCUUGCGCUGAAAUCACCUACAUUUUCCCUGGC-GGCGGACA-----GCAUACAUA ...........(((((((.....((.(((((.(((((.....))))).))))).))(((((((..........))))...)))-)))))...-----))....... ( -28.90) >DroSim_CAF1 53216 100 + 1 UCCACUACUCGGCCCGUCUCUCAGCCAGAGGCCACCUCCGAAAGGUGUCCUCUUGCGCUGAAAUCACCUACAUUUUCCCUGGC-GGCGGACA-----GCAUACAUA ...........(((((((..(((((((((((.(((((.....))))).)))).)).))))).....((............)).-)))))...-----))....... ( -30.00) >DroEre_CAF1 52984 87 + 1 --------------CUGCUCUUUGCCAGCGGCCACCUCCCAAAGGUGUCCUCUUGCGCUGAAAUCACCUACAUUUUCCUUGGCCCGCGGACA-----GCGCACAUA --------------.(((.((...((.(((((((........(((((...((.......))...)))))..........))).))))))..)-----).))).... ( -22.47) >DroPer_CAF1 46280 89 + 1 ---------------AUCUUUGCUCUUGAGGCCACCGCCGAAAGGUGUCCUCUUGCGCUGAAAUCACCUACAUU-CCCAUGGC-AGCACACAUUCAAGCAAACAUA ---------------...((((((...((((.((((.......)))).)))).((.((((..((..........-...))..)-))).))......)))))).... ( -22.32) >consensus ______________CGUCUCUCUGCCAGAGGCCACCUCCGAAAGGUGUCCUCUUGCGCUGAAAUCACCUACAUUUUCCAUGGC_GGCGGACA_____GCAUACAUA .......................((.(((((.((((.......)))).))))).))(((.......((............))..)))................... (-13.65 = -14.48 + 0.83)

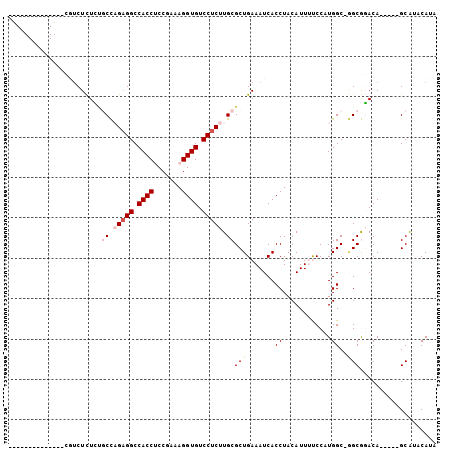
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:42:12 2006