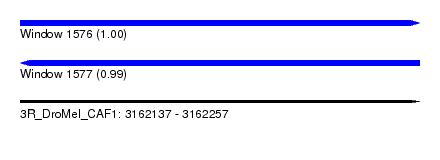
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 3,162,137 – 3,162,257 |
| Length | 120 |
| Max. P | 0.998277 |

| Location | 3,162,137 – 3,162,257 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 96.42 |
| Mean single sequence MFE | -31.01 |
| Consensus MFE | -28.48 |
| Energy contribution | -29.12 |
| Covariance contribution | 0.64 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.14 |
| Structure conservation index | 0.92 |
| SVM decision value | 3.05 |
| SVM RNA-class probability | 0.998277 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
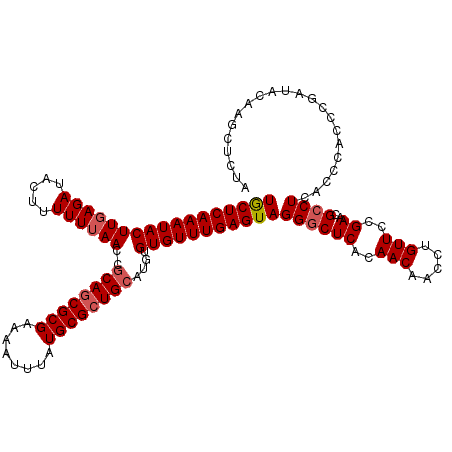
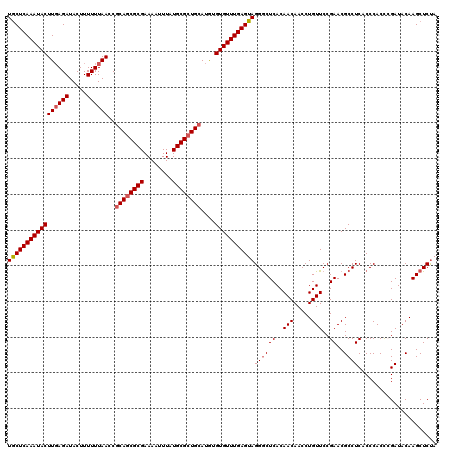
>3R_DroMel_CAF1 3162137 120 + 27905053 UGCUCAAAUACUUGAGAUACUUUUUUAACCGCAGCGCGAAAAUUUAUGCGCUGCAUGUGUGUUUGAGUAGGGCUCACAACAACCUGUUCCGAACGCCUCACCCACCCGAUACAAGCUCUA .((((((((((((((((.....))))))..((((((((........))))))))....))))))))))((((((..........(((..((...............))..))))))))). ( -33.56) >DroSec_CAF1 10015 120 + 1 UGCUCAAAUACUUGAGAUACUUUUUUAACCGCAGCGCGAAAAUUUAUGCGCUGCAUGUGUGUUUGAGUAGGACUCACAACAACCUGUUCCGAACGCCUCACCCACCCGAUACAAGCUCUA (((((((((((((((((.....))))))..((((((((........))))))))....)))))))))))(((...(((......)))))).............................. ( -31.30) >DroSim_CAF1 12183 120 + 1 UACUCAAAUACUUGAGAUACUUUUUUAACCACAGCGCGAAAAUUUAUGCGCUGCAUGUGUGUUUGAGUAGGGCUCACAACAACCUGUUCCGAACGCCUCACCCACCCGAUACAAGCUCUA (((((((((((((((((.....))))))...(((((((........))))))).....)))))))))))(((((..........(((..((...............))..)))))))).. ( -28.86) >DroEre_CAF1 1788 120 + 1 UGCUCAAAUACUUUAGAUACUUUUUUAACCGCAACGCGAAAAUUUAUGCGCUGCAUAUGUGUUUGAGUAGGGCUCACAACAACCUGUUCCGAACGCCUCACCCACUGGAUACAAGCUCUA .((((((((((.(((((......)))))..(((.((((........)))).)))....))))))))))((((((..........(((((((..............)))).))))))))). ( -28.14) >DroYak_CAF1 11597 120 + 1 UGCUCAAAUACUUGAGAUACUUUUUUAACCGCAGCGCGAAAAUUUAUGCGCUGCAUAUGUGUUUGAGUAGGGCUCACAACAACCUGUUCCGAACGCCUCACCCACUAGAUACAAGCUCCA (((((((((((((((((.....))))))..((((((((........))))))))....)))))))))))((((((..(((.....)))..))..))))...................... ( -33.20) >consensus UGCUCAAAUACUUGAGAUACUUUUUUAACCGCAGCGCGAAAAUUUAUGCGCUGCAUGUGUGUUUGAGUAGGGCUCACAACAACCUGUUCCGAACGCCUCACCCACCCGAUACAAGCUCUA (((((((((((((((((.....))))))..((((((((........))))))))....)))))))))))((((((..(((.....)))..))..))))...................... (-28.48 = -29.12 + 0.64)

| Location | 3,162,137 – 3,162,257 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 96.42 |
| Mean single sequence MFE | -36.78 |
| Consensus MFE | -35.36 |
| Energy contribution | -34.72 |
| Covariance contribution | -0.64 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.67 |
| Structure conservation index | 0.96 |
| SVM decision value | 2.26 |
| SVM RNA-class probability | 0.991372 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
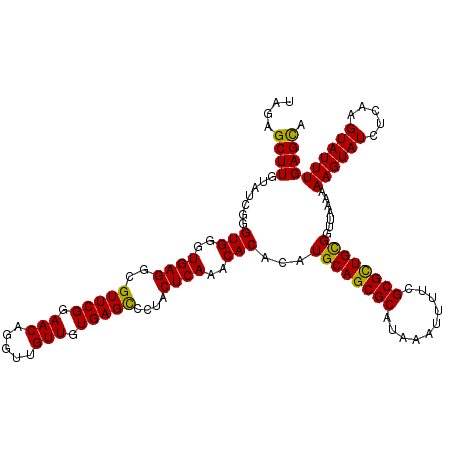
>3R_DroMel_CAF1 3162137 120 - 27905053 UAGAGCUUGUAUCGGGUGGGUGAGGCGUUCGGAACAGGUUGUUGUGAGCCCUACUCAAACACACAUGCAGCGCAUAAAUUUUCGCGCUGCGGUUAAAAAAGUAUCUCAAGUAUUUGAGCA ....((((..((((..(((((.(((.(((((.(((.....))).)))))))))))))..)......(((((((..........)))))))))).....((((((.....)))))))))). ( -38.80) >DroSec_CAF1 10015 120 - 1 UAGAGCUUGUAUCGGGUGGGUGAGGCGUUCGGAACAGGUUGUUGUGAGUCCUACUCAAACACACAUGCAGCGCAUAAAUUUUCGCGCUGCGGUUAAAAAAGUAUCUCAAGUAUUUGAGCA ....((((((((..((((..((((..(((((.(((.....))).)))).)...))))..)))...((((((((..........))))))))...............)..))))..)))). ( -34.30) >DroSim_CAF1 12183 120 - 1 UAGAGCUUGUAUCGGGUGGGUGAGGCGUUCGGAACAGGUUGUUGUGAGCCCUACUCAAACACACAUGCAGCGCAUAAAUUUUCGCGCUGUGGUUAAAAAAGUAUCUCAAGUAUUUGAGUA ....((((..((((..(((((.(((.(((((.(((.....))).)))))))))))))..)......(((((((..........)))))))))).....))))..(((((....))))).. ( -36.40) >DroEre_CAF1 1788 120 - 1 UAGAGCUUGUAUCCAGUGGGUGAGGCGUUCGGAACAGGUUGUUGUGAGCCCUACUCAAACACAUAUGCAGCGCAUAAAUUUUCGCGUUGCGGUUAAAAAAGUAUCUAAAGUAUUUGAGCA ((((((((.((.((.(((..((((..(((((.(((.....))).)))))....))))..)))....(((((((..........))))))))).))...)))).))))............. ( -37.20) >DroYak_CAF1 11597 120 - 1 UGGAGCUUGUAUCUAGUGGGUGAGGCGUUCGGAACAGGUUGUUGUGAGCCCUACUCAAACACAUAUGCAGCGCAUAAAUUUUCGCGCUGCGGUUAAAAAAGUAUCUCAAGUAUUUGAGCA ....((((.......(((..((((..(((((.(((.....))).)))))....))))..)))...((((((((..........)))))))).......((((((.....)))))))))). ( -37.20) >consensus UAGAGCUUGUAUCGGGUGGGUGAGGCGUUCGGAACAGGUUGUUGUGAGCCCUACUCAAACACACAUGCAGCGCAUAAAUUUUCGCGCUGCGGUUAAAAAAGUAUCUCAAGUAUUUGAGCA ....((((.......(((..((((..(((((.(((.....))).)))))....))))..)))...((((((((..........)))))))).......((((((.....)))))))))). (-35.36 = -34.72 + -0.64)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:57:19 2006