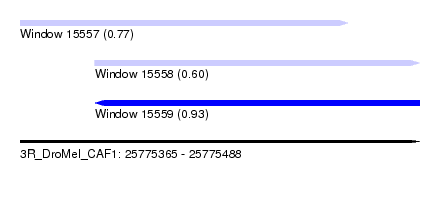

| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 25,775,365 – 25,775,488 |
| Length | 123 |
| Max. P | 0.932987 |

| Location | 25,775,365 – 25,775,466 |
|---|---|
| Length | 101 |
| Sequences | 4 |
| Columns | 109 |
| Reading direction | forward |
| Mean pairwise identity | 78.50 |
| Mean single sequence MFE | -21.48 |
| Consensus MFE | -12.43 |
| Energy contribution | -13.55 |
| Covariance contribution | 1.12 |
| Combinations/Pair | 1.17 |
| Mean z-score | -2.05 |
| Structure conservation index | 0.58 |
| SVM decision value | 0.53 |
| SVM RNA-class probability | 0.770560 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 25775365 101 + 27905053 AAUGUUUAUGCACAAAAUGCACU--------UGUUAUGGGUACCAUAAAUGUGUUAGAAAACUUUUUAAACACGAUUGUUCAAGUUAUGUUUAAAAUUUAUGAGUGUUU .........((((.....((((.--------..((((((...))))))..)))).........(((((((((..(((.....)))..))))))))).......)))).. ( -20.80) >DroSim_CAF1 19182 101 + 1 AAUGUUUAUGAACAAAAUGCACU--------UGUUAUGGUUACCACAAAUGUGUUAACAAACUUUUUAAACACGAUUUUUCAAGUUAUGUUUAAAAUUUAUGAGCACUU ..((((((((((...........--------.(((...((((.(((....))).)))).))).(((((((((..(((.....)))..)))))))))))))))))))... ( -21.00) >DroEre_CAF1 20481 93 + 1 AGUGUUUAUGCACAAAA--------------CGUUAUGGGUGCCAUAAAU--GUUGGCAAACUCCUUAACAAAGAUUUGCCAAGUUUUCUUUAACAUUUACGAGUACUU .((((....))))....--------------......(((((((.(((((--(((((((((...(((....))).))))))(((....))).)))))))).).)))))) ( -21.10) >DroYak_CAF1 17834 107 + 1 GUUGUUUAUGCACAAAAUGCACUUGGACAAAUGUUAUGGGUACCAUAAAU--GUUAGCAAACUUUUUAAACAAGAUUUUCCAAGUUUUGUUUAAAAUUUAUGAGUAACU .(((((((((((.....))))..)))))))...((((((...))))))..--((((.((....((((((((((((((.....))))))))))))))....))..)))). ( -23.00) >consensus AAUGUUUAUGCACAAAAUGCACU________UGUUAUGGGUACCAUAAAU__GUUAGCAAACUUUUUAAACAAGAUUUUCCAAGUUAUGUUUAAAAUUUAUGAGUACUU ((((((((((.......................((((((...))))))...............((((((((((((((.....))))))))))))))..)))))))))). (-12.43 = -13.55 + 1.12)

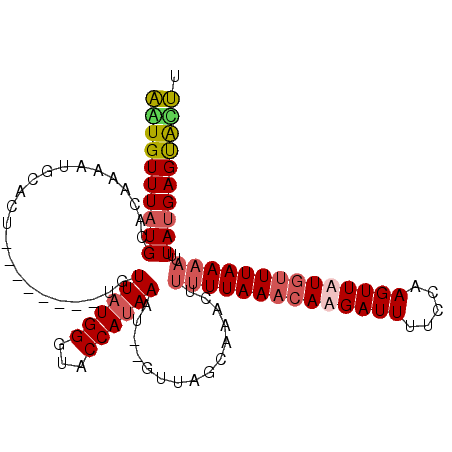
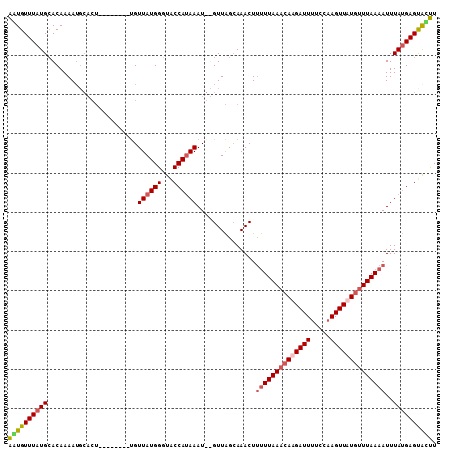
| Location | 25,775,388 – 25,775,488 |
|---|---|
| Length | 100 |
| Sequences | 5 |
| Columns | 111 |
| Reading direction | forward |
| Mean pairwise identity | 78.82 |
| Mean single sequence MFE | -22.02 |
| Consensus MFE | -10.76 |
| Energy contribution | -12.20 |
| Covariance contribution | 1.44 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.93 |
| Structure conservation index | 0.49 |
| SVM decision value | 0.13 |
| SVM RNA-class probability | 0.598556 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 25775388 100 + 27905053 -UGUUAUGGGUACCAUAAAUGUGUUAGAAAACUUUUUAAACACGAUUGUUCAAGUUAUGUUUAAAAUUUAUGAGUGUUUCUUUCAUCGGAAGCCCAUCG----------CU -.((.(((((((((((((((..(((....)))..((((((((..(((.....)))..))))))))))))))).))..((((......)))))))))).)----------). ( -23.20) >DroSec_CAF1 19035 100 + 1 -UGUUAUGGUUACCAUAAAUGUGUUAACAAACUUUUUAAACACGAUUUUUCAAGUUAUGUUUAAAAUUUAUGAGUACUUCUUUCAACAGAAGCCCAUCG----------CU -.((.((((.((((((((((((....)))....(((((((((..(((.....)))..))))))))))))))).)))(((((......))))).)))).)----------). ( -21.30) >DroSim_CAF1 19205 100 + 1 -UGUUAUGGUUACCACAAAUGUGUUAACAAACUUUUUAAACACGAUUUUUCAAGUUAUGUUUAAAAUUUAUGAGCACUUCUUUCAACAGAAGCCCAUCG----------CU -.((.(((((((.(((....))).)))).....(((((((((..(((.....)))..)))))))))..........(((((......)))))..))).)----------). ( -17.80) >DroEre_CAF1 20498 108 + 1 -CGUUAUGGGUGCCAUAAAU--GUUGGCAAACUCCUUAACAAAGAUUUGCCAAGUUUUCUUUAACAUUUACGAGUACUUCGUUCAAAAGCAACCCAUCUCUGUGGCCUGCU -.(..(((((((((.(((((--(((((((((...(((....))).))))))(((....))).)))))))).).)))).)))).)...((((..((((....))))..)))) ( -24.40) >DroYak_CAF1 17864 109 + 1 AUGUUAUGGGUACCAUAAAU--GUUAGCAAACUUUUUAAACAAGAUUUUCCAAGUUUUGUUUAAAAUUUAUGAGUAACUCGUUCACAAGAAACCCAUUGCUGUCGCCUGUU ..((.(((((((((((((((--(((....)))..(((((((((((((.....)))))))))))))))))))).))......(((....))))))))).))........... ( -23.40) >consensus _UGUUAUGGGUACCAUAAAUGUGUUAGCAAACUUUUUAAACACGAUUUUUCAAGUUAUGUUUAAAAUUUAUGAGUACUUCUUUCAACAGAAGCCCAUCG__________CU .....(((((((((((((((..(((....)))..((((((((.((((.....)))).))))))))))))))).))......(((....))))))))).............. (-10.76 = -12.20 + 1.44)

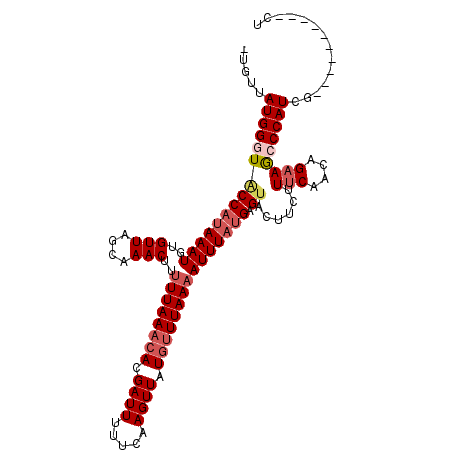
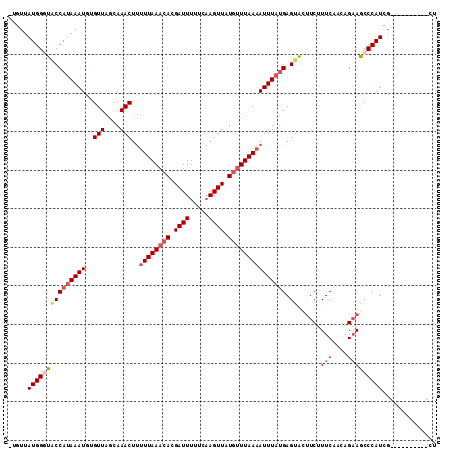
| Location | 25,775,388 – 25,775,488 |
|---|---|
| Length | 100 |
| Sequences | 5 |
| Columns | 111 |
| Reading direction | reverse |
| Mean pairwise identity | 78.82 |
| Mean single sequence MFE | -23.42 |
| Consensus MFE | -13.48 |
| Energy contribution | -12.80 |
| Covariance contribution | -0.68 |
| Combinations/Pair | 1.26 |
| Mean z-score | -2.24 |
| Structure conservation index | 0.58 |
| SVM decision value | 1.23 |
| SVM RNA-class probability | 0.932987 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 25775388 100 - 27905053 AG----------CGAUGGGCUUCCGAUGAAAGAAACACUCAUAAAUUUUAAACAUAACUUGAACAAUCGUGUUUAAAAAGUUUUCUAACACAUUUAUGGUACCCAUAACA- .(----------..(((((...((.((((((((((.(((......((((((((((.............))))))))))))))))))......)))))))..)))))..).- ( -17.12) >DroSec_CAF1 19035 100 - 1 AG----------CGAUGGGCUUCUGUUGAAAGAAGUACUCAUAAAUUUUAAACAUAACUUGAAAAAUCGUGUUUAAAAAGUUUGUUAACACAUUUAUGGUAACCAUAACA- .(----------..((((((((((......))))))((.((((((((((((((((...((....))..)))))))))).(((....)))...))))))))..))))..).- ( -22.40) >DroSim_CAF1 19205 100 - 1 AG----------CGAUGGGCUUCUGUUGAAAGAAGUGCUCAUAAAUUUUAAACAUAACUUGAAAAAUCGUGUUUAAAAAGUUUGUUAACACAUUUGUGGUAACCAUAACA- .(----------..((((((((((......))))))((.((((((((((((((((...((....))..)))))))))).(((....)))...))))))))..))))..).- ( -22.70) >DroEre_CAF1 20498 108 - 1 AGCAGGCCACAGAGAUGGGUUGCUUUUGAACGAAGUACUCGUAAAUGUUAAAGAAAACUUGGCAAAUCUUUGUUAAGGAGUUUGCCAAC--AUUUAUGGCACCCAUAACG- ...........(..((((((((((((.....)))))).(((((((((((.(((....)))((((((.((((.....)))))))))))))--)))))))).))))))..).- ( -29.10) >DroYak_CAF1 17864 109 - 1 AACAGGCGACAGCAAUGGGUUUCUUGUGAACGAGUUACUCAUAAAUUUUAAACAAAACUUGGAAAAUCUUGUUUAAAAAGUUUGCUAAC--AUUUAUGGUACCCAUAACAU ....(((((..((.((((((..((((....))))..))))))...((((((((((...((....))..)))))))))).)))))))...--..((((((...))))))... ( -25.80) >consensus AG__________CGAUGGGCUUCUGUUGAAAGAAGUACUCAUAAAUUUUAAACAUAACUUGAAAAAUCGUGUUUAAAAAGUUUGCUAACACAUUUAUGGUACCCAUAACA_ ..............(((((((((........)))))..(((((((((((((((((...((....))..)))))))))).(((....)))...)))))))...))))..... (-13.48 = -12.80 + -0.68)

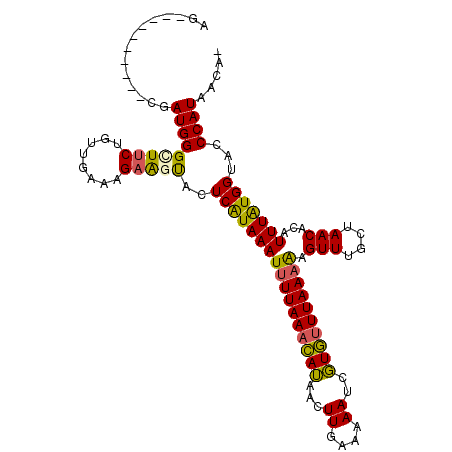
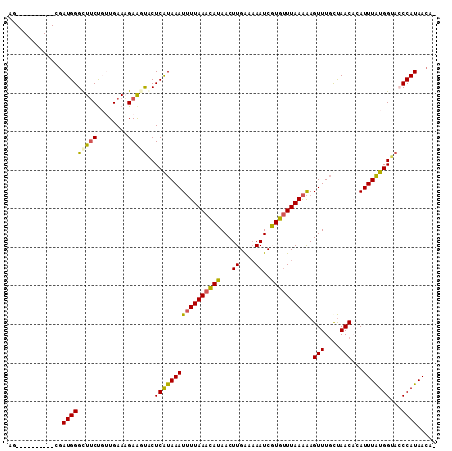
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:41:06 2006