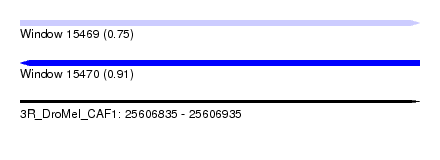

| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 25,606,835 – 25,606,935 |
| Length | 100 |
| Max. P | 0.914924 |

| Location | 25,606,835 – 25,606,935 |
|---|---|
| Length | 100 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | forward |
| Mean pairwise identity | 66.83 |
| Mean single sequence MFE | -32.24 |
| Consensus MFE | -10.64 |
| Energy contribution | -13.24 |
| Covariance contribution | 2.60 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.99 |
| Structure conservation index | 0.33 |
| SVM decision value | 0.48 |
| SVM RNA-class probability | 0.753218 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 25606835 100 + 27905053 CGAUACGUGAUUCUGGCCAGCAAGAGCUGAUCAGACUACUCUAUCA-GCUCUAGCUUCCGAGCUGGAAGAGCUCGGAAUUCGUAUCUGAAACUGGAAGCGG .((((((.((((((((..(((.(((((((((.(((....)))))))-))))).))).))(((((.....))))))))))))))))).....(((....))) ( -41.40) >DroSec_CAF1 33768 92 + 1 CGAUACGUGAUUCUGGCCAGCAAGAGCUGAGCAGACUACUCUACC--GCUCCAGCUUCCGAGCUGGAAGUGCUCGCAAUCUG-------GACUGUAAGCGG ......(((..((((..((((....))))..)))).)))....((--(((((((.((.(((((.......))))).)).)))-------))......)))) ( -30.00) >DroSim_CAF1 34635 99 + 1 UAAUACGUGAUUCUGGCCAGCAAAAGCUGAGCAGACUACUCUACC--GCUCCAGCUUCCGAGCUUGAAGUGCUCGCAAUCCGUAUCUGGAACUGGAAGCGG ......(((..((((..((((....))))..)))).)))....((--(((((((.((((((((.......))))((.....))....)))))))).))))) ( -35.10) >DroEre_CAF1 34753 93 + 1 AGUUACGUGAUUCUGGGCUAUAAGGCAAGUGCAGAGUAAUG--CCAGUCGCUGGCUUCCGUGCUGGAAAUGCCGGC------UAUCUGGAACUGGAAGCUA ((((.(((.((((((.(((........))).)))))).)))--((((..((((((((((.....))))..))))))------...))))))))........ ( -32.10) >DroYak_CAF1 35998 78 + 1 AGAUACGUGAUUAUGGGCUACCAG----------AAU------CCGCUCUUUGGCUUCCGAG-UGGAAGUGUCGGU------UAUCAGGAACUGGAAGCUA .(((((.......(((....))).----------..(------((((((..........)))-)))).)))))(((------(.((((...)))).)))). ( -22.60) >consensus AGAUACGUGAUUCUGGCCAGCAAGAGCUGAGCAGACUACUCUACC__GCUCUAGCUUCCGAGCUGGAAGUGCUCGCAAU__GUAUCUGGAACUGGAAGCGG ...........((((..((((....))))..))))..................((((((((((.......))))...................)))))).. (-10.64 = -13.24 + 2.60)



| Location | 25,606,835 – 25,606,935 |
|---|---|
| Length | 100 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | reverse |
| Mean pairwise identity | 66.83 |
| Mean single sequence MFE | -30.44 |
| Consensus MFE | -11.46 |
| Energy contribution | -14.18 |
| Covariance contribution | 2.72 |
| Combinations/Pair | 1.15 |
| Mean z-score | -2.11 |
| Structure conservation index | 0.38 |
| SVM decision value | 1.10 |
| SVM RNA-class probability | 0.914924 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 25606835 100 - 27905053 CCGCUUCCAGUUUCAGAUACGAAUUCCGAGCUCUUCCAGCUCGGAAGCUAGAGC-UGAUAGAGUAGUCUGAUCAGCUCUUGCUGGCCAGAAUCACGUAUCG ...............((((((...((((((((.....))))))))(((.(((((-(((((((....))).))))))))).)))...........)))))). ( -40.60) >DroSec_CAF1 33768 92 - 1 CCGCUUACAGUC-------CAGAUUGCGAGCACUUCCAGCUCGGAAGCUGGAGC--GGUAGAGUAGUCUGCUCAGCUCUUGCUGGCCAGAAUCACGUAUCG ..(((..((((.-------...))))..)))..((((((((....))))))))(--((((..((..((((..((((....))))..))))...)).))))) ( -34.70) >DroSim_CAF1 34635 99 - 1 CCGCUUCCAGUUCCAGAUACGGAUUGCGAGCACUUCAAGCUCGGAAGCUGGAGC--GGUAGAGUAGUCUGCUCAGCUUUUGCUGGCCAGAAUCACGUAUUA (((((.((((((((......)))((.(((((.......))))).))))))))))--))..((....((((..((((....))))..)))).))........ ( -39.10) >DroEre_CAF1 34753 93 - 1 UAGCUUCCAGUUCCAGAUA------GCCGGCAUUUCCAGCACGGAAGCCAGCGACUGG--CAUUACUCUGCACUUGCCUUAUAGCCCAGAAUCACGUAACU ........(((.((((...------((.(((..((((.....))))))).))..))))--.)))..((((..((........))..))))........... ( -19.90) >DroYak_CAF1 35998 78 - 1 UAGCUUCCAGUUCCUGAUA------ACCGACACUUCCA-CUCGGAAGCCAAAGAGCGG------AUU----------CUGGUAGCCCAUAAUCACGUAUCU ..(((.((((.(((.....------.((((........-.))))..((......))))------)..----------)))).)))................ ( -17.90) >consensus CCGCUUCCAGUUCCAGAUAC__AUUGCGAGCACUUCCAGCUCGGAAGCUAGAGC__GGUAGAGUAGUCUGCUCAGCUCUUGCUGGCCAGAAUCACGUAUCG ..((((((.((..............))((((.......))))))))))..................((((..((((....))))..))))........... (-11.46 = -14.18 + 2.72)

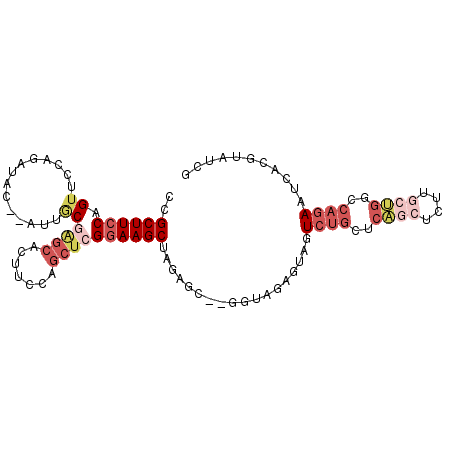

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:39:46 2006