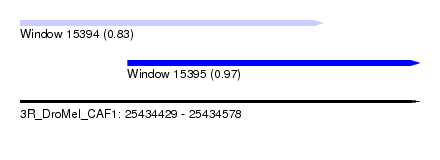
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 25,434,429 – 25,434,578 |
| Length | 149 |
| Max. P | 0.967386 |

| Location | 25,434,429 – 25,434,542 |
|---|---|
| Length | 113 |
| Sequences | 4 |
| Columns | 113 |
| Reading direction | forward |
| Mean pairwise identity | 91.89 |
| Mean single sequence MFE | -35.35 |
| Consensus MFE | -31.50 |
| Energy contribution | -31.75 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.68 |
| Structure conservation index | 0.89 |
| SVM decision value | 0.70 |
| SVM RNA-class probability | 0.828064 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 25434429 113 + 27905053 AUUAGGGGCUGCCUUCAAGGAUCUUUUGCCAAGGACUUUUUGAAAAAGAACUACAUACACGGCAUCGAAUUGCUGGGCACAGAAAGGACCGAGCUGUGCUUUUUGUGCCAAUA ....((..((...(((((((((((((....))))).))))))))..))..)).......(((((......)))))(((((((((((.((......)).))))))))))).... ( -36.20) >DroSec_CAF1 117264 113 + 1 AUUAGGGGCUGCCUUCAAGGAUCUUUUGCCAAGGACUUUUUGCAAAAGAACUACAUACACGGCAUCGAGUUGCUGGGCACAGAGAGGACCGAGCUGUGCUUUCUGUGCCAGUA ...((((....))))...((.((((((((.(((....))).)))))))).))...(((.(((((......)))))(((((((((((.((......)).))))))))))).))) ( -39.60) >DroSim_CAF1 119195 113 + 1 AUUAGGGGCUGCCUUCAAGGAUCUUUUGCCAAGGACUUUCUGCAAAAGAACUACAUACACGGCAUCGAGUUGCUGGGCACAGAGAGGACCGAGCUGUGCUUUCUGUGCCAAUA ...((((....))))...((.((((((((..(((....))))))))))).)).......(((((......)))))(((((((((((.((......)).))))))))))).... ( -39.20) >DroYak_CAF1 127511 113 + 1 AUUAGGGGCUGCCUUCAAGGAUCUUUUGCCAGCGACUUUUUAAAAAAGAAUUACAUACACGGCAUCGAAUUGCUGGGCACCGAAAUGAGCGAACUCUGUUUUCUGUGCCAGUU ......(((..((.....)).......)))...((((......................(((((......)))))(((((.((((.(((....)))...)))).))))))))) ( -26.40) >consensus AUUAGGGGCUGCCUUCAAGGAUCUUUUGCCAAGGACUUUUUGAAAAAGAACUACAUACACGGCAUCGAAUUGCUGGGCACAGAAAGGACCGAGCUGUGCUUUCUGUGCCAAUA ...((((....))))...((.((((((((............)))))))).)).......(((((......)))))(((((((((((.((......)).))))))))))).... (-31.50 = -31.75 + 0.25)

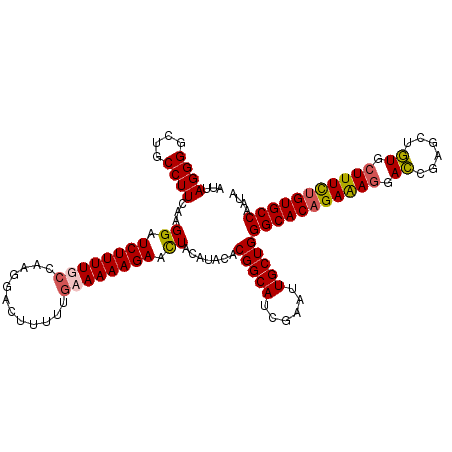
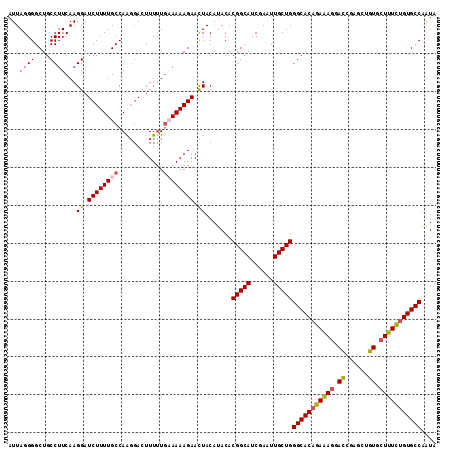
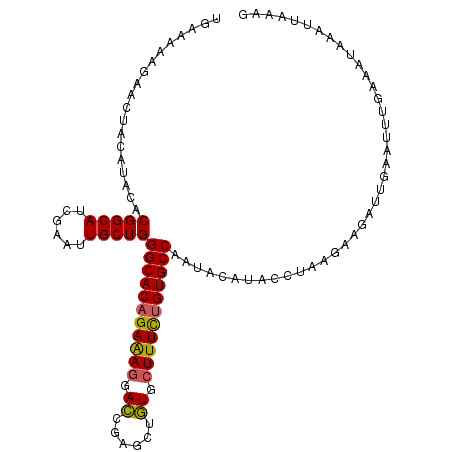
| Location | 25,434,469 – 25,434,578 |
|---|---|
| Length | 109 |
| Sequences | 4 |
| Columns | 109 |
| Reading direction | forward |
| Mean pairwise identity | 90.21 |
| Mean single sequence MFE | -27.38 |
| Consensus MFE | -25.66 |
| Energy contribution | -25.35 |
| Covariance contribution | -0.31 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.99 |
| Structure conservation index | 0.94 |
| SVM decision value | 1.61 |
| SVM RNA-class probability | 0.967386 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 25434469 109 + 27905053 UGAAAAAGAACUACAUACACGGCAUCGAAUUGCUGGGCACAGAAAGGACCGAGCUGUGCUUUUUGUGCCAAUACAUACCUAAGAAGAUUGAAUUUGAAAUAAAUUAAAG ...................(((((......)))))(((((((((((.((......)).)))))))))))........................................ ( -25.70) >DroSec_CAF1 117304 109 + 1 UGCAAAAGAACUACAUACACGGCAUCGAGUUGCUGGGCACAGAGAGGACCGAGCUGUGCUUUCUGUGCCAGUACAUACCUAAGAAGAUUGAAUUUGAAAUAAAUUAAAG ..((((.............(((((......)))))(((((((((((.((......)).))))))))))).......................))))............. ( -29.60) >DroSim_CAF1 119235 109 + 1 UGCAAAAGAACUACAUACACGGCAUCGAGUUGCUGGGCACAGAGAGGACCGAGCUGUGCUUUCUGUGCCAAUACAUACCUAAGAAGAUUGAAUUUGAAAUAAAUUAAAG ..((((.............(((((......)))))(((((((((((.((......)).))))))))))).......................))))............. ( -29.60) >DroYak_CAF1 127551 109 + 1 UAAAAAAGAAUUACAUACACGGCAUCGAAUUGCUGGGCACCGAAAUGAGCGAACUCUGUUUUCUGUGCCAGUUCAUACCUAGAAAGUUGGAAGUUGCAAUAAAUUAAAG .......(((((.......(((((......)))))(((((.((((.(((....)))...)))).))))))))))...((.........))................... ( -24.60) >consensus UGAAAAAGAACUACAUACACGGCAUCGAAUUGCUGGGCACAGAAAGGACCGAGCUGUGCUUUCUGUGCCAAUACAUACCUAAGAAGAUUGAAUUUGAAAUAAAUUAAAG ...................(((((......)))))(((((((((((.((......)).)))))))))))........................................ (-25.66 = -25.35 + -0.31)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:38:40 2006